Related Research Articles

A haplotype is a group of alleles in an organism that are inherited together from a single parent, and a haplogroup is a group of similar haplotypes that share a common ancestor with a single-nucleotide polymorphism mutation. More specifically, a haplogroup is a combination of alleles at different chromosomal regions that are closely linked and that tend to be inherited together. As a haplogroup consists of similar haplotypes, it is usually possible to predict a haplogroup from haplotypes. Haplogroups pertain to a single line of descent. As such, membership of a haplogroup, by any individual, relies on a relatively small proportion of the genetic material possessed by that individual.
Genetics and archaeogenetics of South Asia is the study of the genetics and archaeogenetics of the ethnic groups of South Asia. It aims at uncovering these groups' genetic history. The geographic position of South Asia makes its biodiversity important for the study of the early dispersal of anatomically modern humans across Asia.
Haplogroup T is a human mitochondrial DNA (mtDNA) haplogroup. It is believed to have originated around 25,100 years ago in the Near East.
Haplogroup U is a human mitochondrial DNA haplogroup (mtDNA). The clade arose from haplogroup R, likely during the early Upper Paleolithic. Its various subclades are found widely distributed across Northern and Eastern Europe, Central, Western and South Asia, as well as North Africa, the Horn of Africa, and the Canary Islands.

Haplogroup R is a widely distributed human mitochondrial DNA (mtDNA) haplogroup. Haplogroup R is associated with the peopling of Eurasia after about 70,000 years ago, and is distributed in modern populations throughout the world outside of sub-Saharan Africa.
Haplogroup I is a human mitochondrial DNA (mtDNA) haplogroup. It is believed to have originated about 21,000 years ago, during the Last Glacial Maximum (LGM) period in West Asia. The haplogroup is unusual in that it is now widely distributed geographically, but is common in only a few small areas of East Africa, West Asia and Europe. It is especially common among the El Molo and Rendille peoples of Kenya, various regions of Iran, the Lemko people of Slovakia, Poland and Ukraine, the island of Krk in Croatia, the department of Finistère in France and some parts of Scotland and Ireland.
Haplogroup D1 or D-M174 is a subclade of Haplogroup D-CTS3946. This male haplogroup is found primarily in East Asia and the Andaman Islands, though it is also found regularly with low frequency in Central Asia and Mainland Southeast Asia. It's also found as far as Europe and Middle east in lower frequencies.

Haplogroup F, also known as F-M89 and previously as Haplogroup FT is a very common Y-chromosome haplogroup. The clade and its subclades constitute over 90% of paternal lineages outside of Africa.

Haplogroup H (Y-DNA), also known as H-L901/M2939 is a Y-chromosome haplogroup.

Haplogroup L-M20 is a human Y-DNA haplogroup, which is defined by SNPs M11, M20, M61 and M185. As a secondary descendant of haplogroup K and a primary branch of haplogroup LT, haplogroup L currently has the alternative phylogenetic name of K1a, and is a sibling of haplogroup T.
In human mitochondrial genetics, Haplogroup G is a human mitochondrial DNA (mtDNA) haplogroup.

Haplogroup N1a is a human mitochondrial DNA (mtDNA) haplogroup.
Haplogroup H is a human mitochondrial DNA (mtDNA) haplogroup. The clade is believed to have originated in Southwest Asia, near present day Syria, around 20,000 to 25,000 years ago. Mitochondrial haplogroup H is today predominantly found in Europe, and is believed to have evolved before the Last Glacial Maximum (LGM). It first expanded in the northern Near East and Southern Caucasus soon, and later migrations from Iberia suggest that the clade reached Europe before the Last Glacial Maximum. The haplogroup has also spread to parts of Africa, Siberia and inner Asia. Today, around 40% of all maternal lineages in Europe belong to haplogroup H.
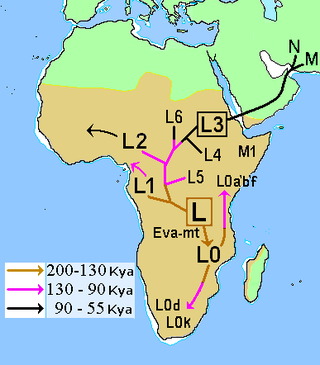
In human mitochondrial genetics, L is the mitochondrial DNA macro-haplogroup that is at the root of the anatomically modern human mtDNA phylogenetic tree. As such, it represents the most ancestral mitochondrial lineage of all currently living modern humans, also dubbed "Mitochondrial Eve".

Genetic studies on the Sinhalese is part of population genetics investigating the origins of the Sinhalese population.
Y-DNA haplogroups in populations of South Asia are haplogroups of the male Y-chromosome found in South Asian populations.
In human mitochondrial genetics, haplogroup M18 is a human mitochondrial DNA (mtDNA) haplogroup. It is an India-specific lineage.
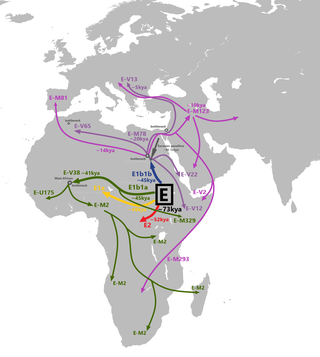
The genetic history of Egypt reflects its geographical location at the crossroads of several major biocultural areas: North Africa, the Sahara, the Middle East, the Mediterranean and sub-Saharan Africa.
The study of the genetics and archaeogenetics of the Gujarati people of India aims at uncovering these people's genetic history. According to the 1000 Genomes Project, "Gujarati" is a general term used to describe people who trace their ancestry to the region of Gujarat, located in the northwestern part of the Indian subcontinent, and who speak the Gujarati language, an Indo-European language. They have some genetic commonalities as well as differences with other ethnic groups of India.
This article explains the genetic makeup and population history of East Asian peoples and their connection to genetically related populations, as well as Oceanians and partly, Central Asians and South Asians, which are collectively referred to as "East Eurasians" in population genomics.
References
- ↑ Mukhtar Ahmed (29 May 2014). Ancient Pakistan - An Archaeological History: Volume I: The Stone Age. Amazon. pp. 245–. ISBN 978-1-4954-9047-7.
- 1 2 3 4 5 Rishishwar, Lavanya; Jordan, I. King (2017). "Implications of human evolution and admixture for mitochondrial replacement therapy". BMC Genomics. 18 (1): 140. doi:10.1186/s12864-017-3539-3. ISSN 1471-2164. PMC 5299762 . PMID 28178941.
- 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 Kivisild, T; Rootsi, S; Metspalu, M; Mastana, S; Kaldma, K; Parik, J; Metspalu, E; Adojaan, M; et al. (2003). "The Genetic Heritage of the Earliest Settlers Persists Both in Indian Tribal and Caste Populations". AJHG. 72 (2): 313–32. doi:10.1086/346068. PMC 379225 . PMID 12536373.
- 1 2 3 4 Ranaweera, Lanka; Kaewsutthi, Supannee; Win Tun, Aung; Boonyarit, Hathaichanoke; Poolsuwan, Samerchai; Lertrit, Patcharee (January 2014). "Mitochondrial DNA history of Sri Lankan ethnic people: their relations within the island and with the Indian subcontinental populations". Journal of Human Genetics. 59 (1): 28–36. doi: 10.1038/jhg.2013.112 . PMID 24196378.
- 1 2 3 4 Ranasinghe, Ruwandi; Tennekoon, Kamani H.; Karunanayake, Eric H.; Lembring, Maria; Allen, Marie (November 2015). "A study of genetic polymorphisms in mitochondrial DNA hypervariable regions I and II of the five major ethnic groups and Vedda population in Sri Lanka". Legal Medicine. 17 (6): 539–546. doi:10.1016/j.legalmed.2015.05.007. PMID 26065620.