Related Research Articles
Gene silencing is the regulation of gene expression in a cell to prevent the expression of a certain gene. Gene silencing can occur during either transcription or translation and is often used in research. In particular, methods used to silence genes are being increasingly used to produce therapeutics to combat cancer and other diseases, such as infectious diseases and neurodegenerative disorders.
Gene knockdown is an experimental technique by which the expression of one or more of an organism's genes is reduced. The reduction can occur either through genetic modification or by treatment with a reagent such as a short DNA or RNA oligonucleotide that has a sequence complementary to either gene or an mRNA transcript.

In molecular biology, a riboswitch is a regulatory segment of a messenger RNA molecule that binds a small molecule, resulting in a change in production of the proteins encoded by the mRNA. Thus, an mRNA that contains a riboswitch is directly involved in regulating its own activity, in response to the concentrations of its effector molecule. The discovery that modern organisms use RNA to bind small molecules, and discriminate against closely related analogs, expanded the known natural capabilities of RNA beyond its ability to code for proteins, catalyze reactions, or to bind other RNA or protein macromolecules.

Small interfering RNA (siRNA), sometimes known as short interfering RNA or silencing RNA, is a class of double-stranded RNA non-coding RNA molecules, typically 20-27 base pairs in length, similar to miRNA, and operating within the RNA interference (RNAi) pathway. It interferes with the expression of specific genes with complementary nucleotide sequences by degrading mRNA after transcription, preventing translation.
HHV Latency Associated Transcript is a length of RNA which accumulates in cells hosting long-term, or latent, Human Herpes Virus (HHV) infections. The LAT RNA is produced by genetic transcription from a certain region of the viral DNA. LAT regulates the viral genome and interferes with the normal activities of the infected host cell.
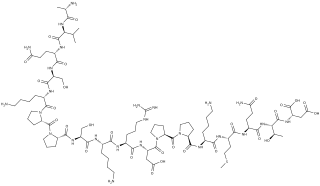
Systemin is a plant peptide hormone involved in the wound response in the family Solanaceae. It was the first plant hormone that was proven to be a peptide having been isolated from tomato leaves in 1991 by a group led by Clarence A. Ryan. Since then, other peptides with similar functions have been identified in tomato and outside of the Solanaceae. Hydroxyproline-rich glycopeptides were found in tobacco in 2001 and AtPEPs were found in Arabidopsis thaliana in 2006. Their precursors are found both in the cytoplasm and cell walls of plant cells, upon insect damage, the precursors are processed to produce one or more mature peptides. The receptor for systemin was first thought to be the same as the brassinolide receptor but this is now uncertain. The signal transduction processes that occur after the peptides bind are similar to the cytokine-mediated inflammatory immune response in animals. Early experiments showed that systemin travelled around the plant after insects had damaged the plant, activating systemic acquired resistance, now it is thought that it increases the production of jasmonic acid causing the same result. The main function of systemins is to coordinate defensive responses against insect herbivores but they also affect plant development. Systemin induces the production of protease inhibitors which protect against insect herbivores, other peptides activate defensins and modify root growth. They have also been shown to affect plants' responses to salt stress and UV radiation. AtPEPs have been shown to affect resistance against oomycetes and may allow A. thaliana to distinguish between different pathogens. In Nicotiana attenuata, some of the peptides have stopped being involved in defensive roles and instead affect flower morphology.
In molecular biology and genetics, the sense of a nucleic acid molecule, particularly of a strand of DNA or RNA, refers to the nature of the roles of the strand and its complement in specifying a sequence of amino acids. Depending on the context, sense may have slightly different meanings. For example, DNA is positive-sense if an RNA version of the same sequence is translated or translatable into protein, negative-sense if not.
Piwi genes were identified as regulatory proteins responsible for stem cell and germ cell differentiation. Piwi is an abbreviation of P-elementInduced WImpy testis in Drosophila. Piwi proteins are highly conserved RNA-binding proteins and are present in both plants and animals. Piwi proteins belong to the Argonaute/Piwi family and have been classified as nuclear proteins. Studies on Drosophila have also indicated that Piwi proteins have slicer activity conferred by the presence of the Piwi domain. In addition, Piwi associates with heterochromatin protein 1, an epigenetic modifier, and piRNA-complementary sequences. These are indications of the role Piwi plays in epigenetic regulation. Piwi proteins are also thought to control the biogenesis of piRNA as many Piwi-like proteins contain slicer activity which would allow Piwi proteins to process precursor piRNA into mature piRNA.

Double-stranded RNA-specific adenosine deaminase is an enzyme that in humans is encoded by the ADAR gene.
Natural antisense transcripts (NATs) are a group of RNAs encoded within a cell that have transcript complementarity to other RNA transcripts. They have been identified in multiple eukaryotes, including humans, mice, yeast and Arabidopsis thaliana. This class of RNAs includes both protein-coding and non-coding RNAs. Current evidence has suggested a variety of regulatory roles for NATs, such as RNA interference (RNAi), alternative splicing, genomic imprinting, and X-chromosome inactivation. NATs are broadly grouped into two categories based on whether they act in cis or in trans. Trans-NATs are transcribed from a different location than their targets and usually have complementarity to multiple transcripts with some mismatches. MicroRNAs (miRNA) are an example of trans-NATs that can target multiple transcripts with a few mismatches. Cis-natural antisense transcripts (cis-NATs) on the other hand are transcribed from the same genomic locus as their target but from the opposite DNA strand and form perfect pairs.

Long non-coding RNAs are a type of RNA, defined as being transcripts with lengths exceeding 200 nucleotides that are not translated into protein. This somewhat arbitrary limit distinguishes long ncRNAs from small non-coding RNAs such as microRNAs (miRNAs), small interfering RNAs (siRNAs), Piwi-interacting RNAs (piRNAs), small nucleolar RNAs (snoRNAs), and other short RNAs. Long intervening/intergenic noncoding RNAs (lincRNAs) are sequences of lncRNA which do not overlap protein-coding genes.
Trans-acting siRNA are a class of small interfering RNA (siRNA) that repress gene expression through post-transcriptional gene silencing in land plants. Precursor transcripts from TAS loci are polyadenylated and converted to double-stranded RNA, and are then processed into 21-nucleotide-long RNA duplexes with overhangs. These segments are incorporated into a RNA-induced silencing complex (RISC) and direct the sequence-specific cleavage of target mRNA. Ta-siRNAs are classified as siRNA because they arise from double-stranded RNA (dsRNA).
RNA polymerase IV is an enzyme that synthesizes small interfering RNA (siRNA) in plants, which silence gene expression. RNAP IV belongs to a family of enzymes that catalyze the process of transcription known as RNA Polymerases, which synthesize RNA from DNA templates. Discovered via phylogenetic studies of land plants, genes of RNAP IV are thought to have resulted from multistep evolution processes that occurred in RNA Polymerase II phylogenies. Such an evolutionary pathway is supported by the fact that RNAP IV is composed of 12 protein subunits that are either similar or identical to RNA polymerase II, and is specific to plant genomes. Via its synthesis of siRNA, RNAP IV is involved in regulation of heterochromatin formation in a process known as RNA directed DNA Methylation (RdDM).
Cryptic unstable transcripts (CUTs) are a subset of non-coding RNAs (ncRNAs) that are produced from intergenic and intragenic regions. CUTs were first observed in S. cerevisiae yeast models and are found in most eukaryotes. Some basic characteristics of CUTs include a length of around 200–800 base pairs, a 5' cap, poly-adenylated tail, and rapid degradation due to the combined activity of poly-adenylating polymerases and exosome complexes. CUT transcription occurs through RNA Polymerase II and initiates from nucleosome-depleted regions, often in an antisense orientation. To date, CUTs have a relatively uncharacterized function but have been implicated in a number of putative gene regulation and silencing pathways. Thousands of loci leading to the generation of CUTs have been described in the yeast genome. Additionally, stable uncharacterized transcripts, or SUTs, have also been detected in cells and bear many similarities to CUTs but are not degraded through the same pathways.

A genetically modified tomato, or transgenic tomato, is a tomato that has had its genes modified, using genetic engineering. The first trial genetically modified food was a tomato engineered to have a longer shelf life, but never made it to market. The first direct consumption tomato was approved in Japan in 2021. Primary work is focused on developing tomatoes with new traits like increased resistance to pests or environmental stresses. Other projects aim to enrich tomatoes with substances that may offer health benefits or be more nutritious. As well as aiming to produce novel crops, scientists produce genetically modified tomatoes to understand the function of genes naturally present in tomatoes.

RNA interference (RNAi) is a biological process in which RNA molecules are involved in sequence-specific suppression of gene expression by double-stranded RNA, through translation or transcriptional repression. Historically, RNAi was known by other names, including co-suppression, post-transcriptional gene silencing (PTGS), and quelling. The detailed study of each of these seemingly different processes elucidated that the identity of these phenomena were all actually RNAi. Andrew Fire and Craig C. Mello shared the 2006 Nobel Prize in Physiology or Medicine for their work on RNA interference in the nematode worm Caenorhabditis elegans, which they published in 1998. Since the discovery of RNAi and its regulatory potentials, it has become evident that RNAi has immense potential in suppression of desired genes. RNAi is now known as precise, efficient, stable and better than antisense therapy for gene suppression. Antisense RNA produced intracellularly by an expression vector may be developed and find utility as novel therapeutic agents.
An endogenous viral element (EVE) is a DNA sequence derived from a virus, and present within the germline of a non-viral organism. EVEs may be entire viral genomes (proviruses), or fragments of viral genomes. They arise when a viral DNA sequence becomes integrated into the genome of a germ cell that goes on to produce a viable organism. The newly established EVE can be inherited from one generation to the next as an allele in the host species, and may even reach fixation.

DNA-directed RNA interference (ddRNAi) is a gene-silencing technique that utilizes DNA constructs to activate an animal cell's endogenous RNA interference (RNAi) pathways. DNA constructs are designed to express self-complementary double-stranded RNAs, typically short-hairpin RNAs (shRNA), that once processed bring about silencing of a target gene or genes. Any RNA, including endogenous mRNAs or viral RNAs, can be silenced by designing constructs to express double-stranded RNA complementary to the desired mRNA target.
Jian-Kang Zhu is a plant scientist, researcher and academic. He is a Senior Principal Investigator in the Shanghai Center for Plant Stress Biology, Chinese Academy of Sciences (CAS). He is also the Academic Director of CAS Center of Excellence in Plant Sciences.

RNA-directed DNA methylation (RdDM) is a biological process in which non-coding RNA molecules direct the addition of DNA methylation to specific DNA sequences. The RdDM pathway is unique to plants, although other mechanisms of RNA-directed chromatin modification have also been described in fungi and animals. To date, the RdDM pathway is best characterized within angiosperms, and particularly within the model plant Arabidopsis thaliana. However, conserved RdDM pathway components and associated small RNAs (sRNAs) have also been found in other groups of plants, such as gymnosperms and ferns. The RdDM pathway closely resembles other sRNA pathways, particularly the highly conserved RNAi pathway found in fungi, plants, and animals. Both the RdDM and RNAi pathways produce sRNAs and involve conserved Argonaute, Dicer and RNA-dependent RNA polymerase proteins.
References
- ↑ Moldovan, D.; Spriggs, A.; Dennis, E. S.; Wilson, I. W. (2010). "The hunt for hypoxia responsive natural antisense short interfering RNAs". Plant Signaling & Behavior. 5 (3): 247–251. doi:10.4161/psb.5.3.10548. PMC 2881268 . PMID 20009576.
- ↑ Katiyar-Agarwal, S.; Morgan, R.; Dahlbeck, D.; Borsani, O.; Villegas, A.; Zhu, J. -K.; Staskawicz, B. J.; Jin, H. (2006). "A pathogen-inducible endogenous siRNA in plant immunity". Proceedings of the National Academy of Sciences. 103 (47): 18002–18007. doi:10.1073/pnas.0608258103. PMC 1693862 . PMID 17071740.
- ↑ Borsani, O.; Zhu, J.; Verslues, P. E.; Sunkar, R.; Zhu, J. K. (2005). "Endogenous siRNAs Derived from a Pair of Natural cis-Antisense Transcripts Regulate Salt Tolerance in Arabidopsis". Cell. 123 (7): 1279–1291. doi:10.1016/j.cell.2005.11.035. PMC 3137516 . PMID 16377568.
- ↑ Held, M. A.; Penning, B.; Brandt, A. S.; Kessans, S. A.; Yong, W.; Scofield, S. R.; Carpita, N. C. (2008). "Small-interfering RNAs from natural antisense transcripts derived from a cellulose synthase gene modulate cell wall biosynthesis in barley". Proceedings of the National Academy of Sciences. 105 (51): 20534–20539. doi:10.1073/pnas.0809408105. PMC 2603254 . PMID 19075248.
- ↑ Zhang, Xiaoming; Xia, Jing (22 March 2012). "Genome-wide analysis of plant nat-siRNAs reveals insights into their distribution, biogenesis and function". Genome Biology. 13 (20): R20. doi:10.1186/gb-2012-13-3-r20. PMC 3439971 . PMID 22439910.