Related Research Articles

DNA ligase is a type of enzyme that facilitates the joining of DNA strands together by catalyzing the formation of a phosphodiester bond. It plays a role in repairing single-strand breaks in duplex DNA in living organisms, but some forms may specifically repair double-strand breaks. Single-strand breaks are repaired by DNA ligase using the complementary strand of the double helix as a template, with DNA ligase creating the final phosphodiester bond to fully repair the DNA.
A restriction enzyme, restriction endonuclease, REase, ENase orrestrictase is an enzyme that cleaves DNA into fragments at or near specific recognition sites within molecules known as restriction sites. Restriction enzymes are one class of the broader endonuclease group of enzymes. Restriction enzymes are commonly classified into five types, which differ in their structure and whether they cut their DNA substrate at their recognition site, or if the recognition and cleavage sites are separate from one another. To cut DNA, all restriction enzymes make two incisions, once through each sugar-phosphate backbone of the DNA double helix.
An inverted repeat is a single stranded sequence of nucleotides followed downstream by its reverse complement. The intervening sequence of nucleotides between the initial sequence and the reverse complement can be any length including zero. For example, 5'---TTACGnnnnnnCGTAA---3' is an inverted repeat sequence. When the intervening length is zero, the composite sequence is a palindromic sequence.

In biochemistry, a nuclease is an enzyme capable of cleaving the phosphodiester bonds between nucleotides of nucleic acids. Nucleases variously effect single and double stranded breaks in their target molecules. In living organisms, they are essential machinery for many aspects of DNA repair. Defects in certain nucleases can cause genetic instability or immunodeficiency. Nucleases are also extensively used in molecular cloning.
A cDNA library is a combination of cloned cDNA fragments inserted into a collection of host cells, which constitute some portion of the transcriptome of the organism and are stored as a "library". cDNA is produced from fully transcribed mRNA found in the nucleus and therefore contains only the expressed genes of an organism. Similarly, tissue-specific cDNA libraries can be produced. In eukaryotic cells the mature mRNA is already spliced, hence the cDNA produced lacks introns and can be readily expressed in a bacterial cell. While information in cDNA libraries is a powerful and useful tool since gene products are easily identified, the libraries lack information about enhancers, introns, and other regulatory elements found in a genomic DNA library.
V(D)J recombination is the mechanism of somatic recombination that occurs only in developing lymphocytes during the early stages of T and B cell maturation. It results in the highly diverse repertoire of antibodies/immunoglobulins and T cell receptors (TCRs) found in B cells and T cells, respectively. The process is a defining feature of the adaptive immune system.

A palindromic sequence is a nucleic acid sequence in a double-stranded DNA or RNA molecule whereby reading in a certain direction on one strand is identical to the sequence in the same direction on the complementary strand. This definition of palindrome thus depends on complementary strands being palindromic of each other.

EcoRV is a type II restriction endonuclease isolated from certain strains of Escherichia coli. It has the alternative name Eco32I.
Fragmentation describes the process of splitting into several pieces or fragments. In cell biology, fragmentation is useful for a cell during both DNA cloning and apoptosis. DNA cloning is important in asexual reproduction or creation of identical DNA molecules, and can be performed spontaneously by the cell or intentionally by laboratory researchers. Apoptosis is the programmed destruction of cells, and the DNA molecules within them, and is a highly regulated process. These two ways in which fragmentation is used in cellular processes describe normal cellular functions and common laboratory procedures performed with cells. However, problems within a cell can sometimes cause fragmentation that results in irregularities such as red blood cell fragmentation and sperm cell DNA fragmentation.
Sequencing by ligation is a DNA sequencing method that uses the enzyme DNA ligase to identify the nucleotide present at a given position in a DNA sequence. Unlike most currently popular DNA sequencing methods, this method does not use a DNA polymerase to create a second strand. Instead, the mismatch sensitivity of a DNA ligase enzyme is used to determine the underlying sequence of the target DNA molecule.
BglII is a type II restriction endonuclease isolated from certain strains of Bacillus globigii.
Topoisomerase-based cloning is a molecular biology technique in which DNA fragments are cloned into specific vectors without the requirement for DNA ligases. Taq polymerase has a nontemplate-dependent terminal transferase activity that adds a single deoxyadenosine (A) to the 3'-end of the PCR products. This characteristic is exploited in "sticky end" TOPO TA cloning. For "blunt end" TOPO cloning, the recipient vector does not have overhangs and blunt-ended DNA fragments can be cloned.
In molecular biology, REBASE is a database of information about restriction enzymes and DNA methyltransferases. REBASE contains an extensive set of references, sites of recognition and cleavage, sequences and structures. It also contains information on the commercial availability of each enzyme. REBASE is one of the longest running biological databases having its roots in a collection of restriction enzymes maintained by Richard J. Roberts since before 1980. Since that time there have been regular descriptions of the resource in the journal Nucleic Acids Research.
DNA ends refer to the properties of the ends of linear DNA molecules, which in molecular biology are described as "sticky" or "blunt" based on the shape of the complementary strands at the terminus. In sticky ends, one strand is longer than the other, such that the longer strand has bases which are left unpaired. In blunt ends, both strands are of equal length – i.e. they end at the same base position, leaving no unpaired bases on either strand.

Ligation is the joining of two nucleic acid fragments through the action of an enzyme. It is an essential laboratory procedure in the molecular cloning of DNA, whereby DNA fragments are joined to create recombinant DNA molecules (such as when a foreign DNA fragment is inserted into a plasmid). The ends of DNA fragments are joined by the formation of phosphodiester bonds between the 3'-hydroxyl of one DNA terminus with the 5'-phosphoryl of another. RNA may also be ligated similarly. A co-factor is generally involved in the reaction, and this is usually ATP or NAD+. Eukaryotic cells ligases belong to ATP type, and NAD+ - dependent are found in bacteria (e.g. E. coli).

Cas12a is a subtype of Cas12 proteins and an RNA-guided endonuclease that forms part of the CRISPR system in some bacteria and archaea. It originates as part of a bacterial immune mechanism, where it serves to destroy the genetic material of viruses and thus protect the cell and colony from viral infection. Cas12a and other CRISPR associated endonucleases use an RNA to target nucleic acid in a specific and programmable matter. In the organisms from which it originates, this guide RNA is a copy of a piece of foreign nucleic acid that previously infected the cell.
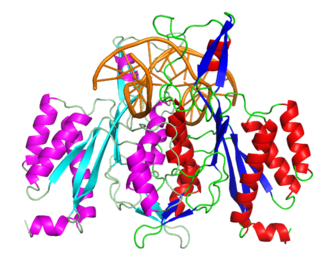
EcoRI is a restriction endonuclease enzyme isolated from species E. coli. It is a restriction enzyme that cleaves DNA double helices into fragments at specific sites, and is also a part of the restriction modification system. The Eco part of the enzyme's name originates from the species from which it was isolated - "E" denotes generic name which is "Escherichia" and "co" denotes species name, "coli" - while the R represents the particular strain, in this case RY13, and the I denotes that it was the first enzyme isolated from this strain.
This glossary of cellular and molecular biology is a list of definitions of terms and concepts commonly used in the study of cell biology, molecular biology, and related disciplines, including molecular genetics, biochemistry, and microbiology. It is split across two articles:
This glossary of cellular and molecular biology is a list of definitions of terms and concepts commonly used in the study of cell biology, molecular biology, and related disciplines, including genetics, biochemistry, and microbiology. It is split across two articles:
References
- ↑ Russell, Peter J. (2006). iGenetics: A Mendelian Approach . Benjamin Cummings. ISBN 978-0805346664.
- 1 2 3 Lehninger, Albert L.; Nelson, David L.; Cox, Michael M. (2008). Principles of Biochemistry (5th ed.). New York, NY: W.H. Freeman and Company. p. 305–306. ISBN 978-0-7167-7108-1.
- ↑ Mousavi-Khattat, Mohammad; Rafati, Adele; Gill, Pooria (5 February 2015). "Fabrication of DNA nanotubes using origami-based nanostructures with sticky ends". Journal of Nanostructure in Chemistry. 5 (2): 177–183. doi: 10.1007/s40097-015-0148-z .
- ↑ Gao, Song; Zhang, Jiannan; Miao, Tianjin; Ma, Di; Su, Ying; An, Yingfeng; Zhang, Qingrui (28 March 2015). "A Simple and Convenient Sticky/Blunt-End Ligation Method for Fusion Gene Construction". Biochemical Genetics. 53 (1–3): 42–48. doi:10.1007/s10528-015-9669-x. PMID 25820211. S2CID 16709792.
- ↑ Roberts, Richard J.; Vincze, Tamas; Posfai, Janos; Macelis, Dana (2009-10-21). "REBASE—a database for DNA restriction and modification: enzymes, genes and genomes". Nucleic Acids Research. 38 (suppl_1): D234–D236. doi:10.1093/nar/gkp874. ISSN 0305-1048. PMC 2808884 . PMID 19846593.
- ↑ Roberts, Richard J.; Vincze, Tamas; Posfai, Janos; Macelis, Dana (2014-11-05). "REBASE—a database for DNA restriction and modification: enzymes, genes and genomes". Nucleic Acids Research. 43 (D1): D298–D299. doi:10.1093/nar/gku1046. ISSN 1362-4962. PMC 4383893 . PMID 25378308.
- ↑ Koulouras, Grigorios; Frith, Martin C (2021-04-06). "Significant non-existence of sequences in genomes and proteomes". Nucleic Acids Research. 49 (6): 3139–3155. doi: 10.1093/nar/gkab139 . ISSN 0305-1048. PMC 8034619 . PMID 33693858.