
A chromosome is a package of DNA with part or all of the genetic material of an organism. In most chromosomes, the very long thin DNA fibers are coated with nucleosome-forming packaging proteins; in eukaryotic cells the most important of these proteins are the histones. These proteins, aided by chaperone proteins, bind to and condense the DNA molecule to maintain its integrity. These chromosomes display a complex three-dimensional structure, which plays a significant role in transcriptional regulation.
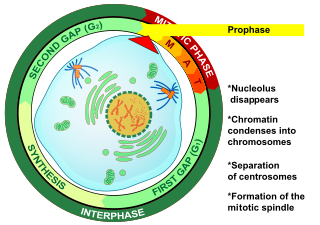
Prophase is the first stage of cell division in both mitosis and meiosis. Beginning after interphase, DNA has already been replicated when the cell enters prophase. The main occurrences in prophase are the condensation of the chromatin reticulum and the disappearance of the nucleolus.

A karyotype is the general appearance of the complete set of chromosomes in the cells of a species or in an individual organism, mainly including their sizes, numbers, and shapes. Karyotyping is the process by which a karyotype is discerned by determining the chromosome complement of an individual, including the number of chromosomes and any abnormalities.
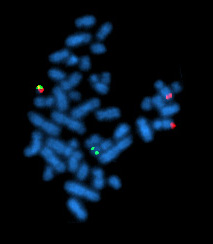
Cytogenetics is essentially a branch of genetics, but is also a part of cell biology/cytology, that is concerned with how the chromosomes relate to cell behaviour, particularly to their behaviour during mitosis and meiosis. Techniques used include karyotyping, analysis of G-banded chromosomes, other cytogenetic banding techniques, as well as molecular cytogenetics such as fluorescence in situ hybridization (FISH) and comparative genomic hybridization (CGH).

Metaphase is a stage of mitosis in the eukaryotic cell cycle in which chromosomes are at their second-most condensed and coiled stage. These chromosomes, carrying genetic information, align in the equator of the cell between the spindle poles at the metaphase plate, before being separated into each of the two daughter nuclei. This alignment marks the beginning of metaphase. Metaphase accounts for approximately 4% of the cell cycle's duration.

Polytene chromosomes are large chromosomes which have thousands of DNA strands. They provide a high level of function in certain tissues such as salivary glands of insects.

Fluorescence in situ hybridization (FISH) is a molecular cytogenetic technique that uses fluorescent probes that bind to only particular parts of a nucleic acid sequence with a high degree of sequence complementarity. It was developed by biomedical researchers in the early 1980s to detect and localize the presence or absence of specific DNA sequences on chromosomes. Fluorescence microscopy can be used to find out where the fluorescent probe is bound to the chromosomes. FISH is often used for finding specific features in DNA for use in genetic counseling, medicine, and species identification. FISH can also be used to detect and localize specific RNA targets in cells, circulating tumor cells, and tissue samples. In this context, it can help define the spatial-temporal patterns of gene expression within cells and tissues.
Torbjörn Oskar Caspersson was a Swedish cytologist and geneticist. He was born in Motala and attended the University of Stockholm, where he studied medicine and biophysics.

In genetics, a locus is a specific, fixed position on a chromosome where a particular gene or genetic marker is located. Each chromosome carries many genes, with each gene occupying a different position or locus; in humans, the total number of protein-coding genes in a complete haploid set of 23 chromosomes is estimated at 19,000–20,000.
Chromosome 13 is one of the 23 pairs of chromosomes in humans. People normally have two copies of this chromosome. Chromosome 13 spans about 113 million base pairs and represents between 3.5 and 4% of the total DNA in cells.

Chromosome 10 is one of the 23 pairs of chromosomes in humans. People normally have two copies of this chromosome. Chromosome 10 spans about 134 million base pairs and represents between 4 and 4.5 percent of the total DNA in cells.

Chromosome 12 is one of the 23 pairs of chromosomes in humans. People normally have two copies of this chromosome. Chromosome 12 spans about 133 million base pairs and represents between 4 and 4.5 percent of the total DNA in cells.
Chromosome 14 is one of the 23 pairs of chromosomes in humans. People normally have two copies of this chromosome. Chromosome 14 spans about 101 million base pairs and represents between 3 and 3.5% of the total DNA in cells.

Chromosome 18 is one of the 23 pairs of chromosomes in humans. People normally have two copies of this chromosome. Chromosome 18 spans about 80 million base pairs and represents about 2.5 percent of the total DNA in cells.

Chromosome 20 is one of the 23 pairs of chromosomes in humans. Chromosome 20 spans around 66 million base pairs and represents between 2 and 2.5 percent of the total DNA in cells. Chromosome 20 was fully sequenced in 2001 and was reported to contain over 59 million base pairs. Since then, due to sequencing improvements and fixes, the length of chromosome 20 has been updated to just over 66 million base pairs.
A dicentric chromosome is an abnormal chromosome with two centromeres. It is formed through the fusion of two chromosome segments, each with a centromere, resulting in the loss of acentric fragments and the formation of dicentric fragments. The formation of dicentric chromosomes has been attributed to genetic processes, such as Robertsonian translocation and paracentric inversion. Dicentric chromosomes have important roles in the mitotic stability of chromosomes and the formation of pseudodicentric chromosomes. Their existence has been linked to certain natural phenomena such as irradiation and have been documented to underlie certain clinical syndromes, notably Kabuki syndrome. The formation of dicentric chromosomes and their implications on centromere function are studied in certain clinical cytogenetics laboratories.

G-banding, G banding or Giemsa banding is a technique used in cytogenetics to produce a visible karyotype by staining condensed chromosomes. It is the most common chromosome banding method. It is useful for identifying genetic diseases through the photographic representation of the entire chromosome complement.
A chromosomal abnormality, chromosomal anomaly, chromosomal aberration, chromosomal mutation, or chromosomal disorder is a missing, extra, or irregular portion of chromosomal DNA. These can occur in the form of numerical abnormalities, where there is an atypical number of chromosomes, or as structural abnormalities, where one or more individual chromosomes are altered. Chromosome mutation was formerly used in a strict sense to mean a change in a chromosomal segment, involving more than one gene. Chromosome anomalies usually occur when there is an error in cell division following meiosis or mitosis. Chromosome abnormalities may be detected or confirmed by comparing an individual's karyotype, or full set of chromosomes, to a typical karyotype for the species via genetic testing.

Molecular cytogenetics combines two disciplines, molecular biology and cytogenetics, and involves the analysis of chromosome structure to help distinguish normal and cancer-causing cells. Human cytogenetics began in 1956 when it was discovered that normal human cells contain 46 chromosomes. However, the first microscopic observations of chromosomes were reported by Arnold, Flemming, and Hansemann in the late 1800s. Their work was ignored for decades until the actual chromosome number in humans was discovered as 46. In 1879, Arnold examined sarcoma and carcinoma cells having very large nuclei. Today, the study of molecular cytogenetics can be useful in diagnosing and treating various malignancies such as hematological malignancies, brain tumors, and other precursors of cancer. The field is overall focused on studying the evolution of chromosomes, more specifically the number, structure, function, and origin of chromosome abnormalities. It includes a series of techniques referred to as fluorescence in situ hybridization, or FISH, in which DNA probes are labeled with different colored fluorescent tags to visualize one or more specific regions of the genome. Introduced in the 1980s, FISH uses probes with complementary base sequences to locate the presence or absence of the specific DNA regions. FISH can either be performed as a direct approach to metaphase chromosomes or interphase nuclei. Alternatively, an indirect approach can be taken in which the entire genome can be assessed for copy number changes using virtual karyotyping. Virtual karyotypes are generated from arrays made of thousands to millions of probes, and computational tools are used to recreate the genome in silico.
The 2000s witnessed an explosion of genome sequencing and mapping in evolutionarily diverse species. While full genome sequencing of mammals is rapidly progressing, the ability to assemble and align orthologous whole chromosomal regions from more than a few species is not yet possible. The intense focus on the building of comparative maps for domestic, laboratory and agricultural (cattle) animals has traditionally been used to understand the underlying basis of disease-related and healthy phenotypes.















