Related Research Articles
Molecular biology is the branch of biology that seeks to understand the molecular basis of biological activity in and between cells, including molecular synthesis, modification, mechanisms, and interactions. The study of chemical and physical structure of biological macromolecules is known as molecular biology.
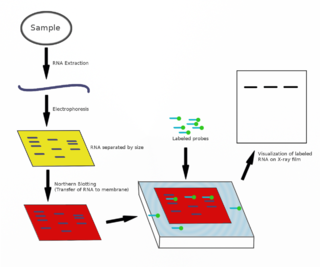
The northern blot, or RNA blot, is a technique used in molecular biology research to study gene expression by detection of RNA in a sample.

A Southern blot is a method used in molecular biology for detection of a specific DNA sequence in DNA samples. Southern blotting combines transfer of electrophoresis-separated DNA fragments to a filter membrane and subsequent fragment detection by probe hybridization.
Oligonucleotides are short DNA or RNA molecules, oligomers, that have a wide range of applications in genetic testing, research, and forensics. Commonly made in the laboratory by solid-phase chemical synthesis, these small bits of nucleic acids can be manufactured as single-stranded molecules with any user-specified sequence, and so are vital for artificial gene synthesis, polymerase chain reaction (PCR), DNA sequencing, molecular cloning and as molecular probes. In nature, oligonucleotides are usually found as small RNA molecules that function in the regulation of gene expression, or are degradation intermediates derived from the breakdown of larger nucleic acid molecules.

A DNA microarray is a collection of microscopic DNA spots attached to a solid surface. Scientists use DNA microarrays to measure the expression levels of large numbers of genes simultaneously or to genotype multiple regions of a genome. Each DNA spot contains picomoles of a specific DNA sequence, known as probes. These can be a short section of a gene or other DNA element that are used to hybridize a cDNA or cRNA sample under high-stringency conditions. Probe-target hybridization is usually detected and quantified by detection of fluorophore-, silver-, or chemiluminescence-labeled targets to determine relative abundance of nucleic acid sequences in the target. The original nucleic acid arrays were macro arrays approximately 9 cm × 12 cm and the first computerized image based analysis was published in 1981. It was invented by Patrick O. Brown. An example of its application is in SNPs arrays for polymorphisms in cardiovascular diseases, cancer, pathogens and GWAS analysis. Also for identification of structural variations and measurement of gene expression.
A lysis buffer is a buffer solution used for the purpose of breaking open cells for use in molecular biology experiments that analyze the labile macromolecules of the cells. Most lysis buffers contain buffering salts and ionic salts to regulate the pH and osmolarity of the lysate. Sometimes detergents are added to break up membrane structures. For lysis buffers targeted at protein extraction, protease inhibitors are often included, and in difficult cases may be almost required. Lysis buffers can be used on both animal and plant tissue cells.
A blot, in molecular biology and genetics, is a method of transferring proteins, DNA or RNA onto a carrier. In many instances, this is done after a gel electrophoresis, transferring the molecules from the gel onto the blotting membrane, and other times adding the samples directly onto the membrane. After the blotting, the transferred proteins, DNA or RNA are then visualized by colorant staining, autoradiographic visualization of radiolabelled molecules, or specific labelling of some proteins or nucleic acids. The latter is done with antibodies or hybridization probes that bind only to some molecules of the blot and have an enzyme joined to them. After proper washing, this enzymatic activity is visualized by incubation with proper reactive, rendering either a colored deposit on the blot or a chemiluminescent reaction which is registered by photographic film.
Comparative genomic hybridization(CGH) is a molecular cytogenetic method for analysing copy number variations (CNVs) relative to ploidy level in the DNA of a test sample compared to a reference sample, without the need for culturing cells. The aim of this technique is to quickly and efficiently compare two genomic DNA samples arising from two sources, which are most often closely related, because it is suspected that they contain differences in terms of either gains or losses of either whole chromosomes or subchromosomal regions. This technique was originally developed for the evaluation of the differences between the chromosomal complements of solid tumor and normal tissue, and has an improved resolution of 5–10 megabases compared to the more traditional cytogenetic analysis techniques of giemsa banding and fluorescence in situ hybridization (FISH) which are limited by the resolution of the microscope utilized.
This is a list of topics in molecular biology. See also index of biochemistry articles.
In molecular biology, a hybridization probe(HP) is a fragment of DNA or RNA of usually 15–10000 nucleotide long which can be radioactively or fluorescently labeled. HP can be used to detect the presence of nucleotide sequences in analyzed RNA or DNA that are complementary to the sequence in the probe. The labeled probe is first denatured into single stranded DNA (ssDNA) and then hybridized to the target ssDNA or RNA immobilized on a membrane or in situ.

The southwestern blot, is a lab technique that involves identifying as well as characterizing DNA-binding proteins by their ability to bind to specific oligonucleotide probes. Determination of molecular weight of proteins binding to DNA is also made possible by the technique. The name originates from a combination of ideas underlying Southern blotting and Western blotting techniques of which they detect DNA and protein respectively. Similar to other types of blotting, proteins are separated by SDS-PAGE and are subsequently transferred to nitrocellulose membranes. Thereafter southwestern blotting begins to vary with regards to procedure as since the first blotting’s, many more have been proposed and discovered with goals of enhancing results. Former protocols were hampered by the need for large amounts of proteins and their susceptibility to degradation while being isolated.

Sir Edwin Mellor Southern is an English Lasker Award-winning molecular biologist, Emeritus Professor of Biochemistry at the University of Oxford and a fellow of Trinity College, Oxford. He is most widely known for the invention of the Southern blot, published in 1975 and now a common laboratory procedure.

The following outline is provided as an overview of and topical guide to biochemistry:
An RNA spike-in is an RNA transcript of known sequence and quantity used to calibrate measurements in RNA hybridization assays, such as DNA microarray experiments, RT-qPCR, and RNA-Seq.
A zoo blot or garden blot is a type of Southern blot that demonstrates the similarity between specific, usually protein-coding, DNA sequences of different species. A zoo blot compares animal species while a garden blot compares plant species. The purpose of the zoo blot is to detect the conservation of the gene(s) of interest throughout the evolution of different species.
The following outline is provided as an overview of and topical guide to genetics:
Nucleic acid methods are the techniques used to study nucleic acids: DNA and RNA.
The reverse northern blot is a method by which gene expression patterns may be analyzed by comparing isolated RNA molecules from a tester sample to samples in a control cDNA library. It is a variant of the northern blot in which the nucleic acid immobilized on a membrane is a collection of isolated DNA fragments rather than RNA, and the probe is RNA extracted from a tissue and radioactively labelled. A reverse northern blot can be used to profile expression levels of particular sets of RNA sequences in a tissue or to determine presence of a particular RNA sequence in a sample. Although DNA Microarrays and newer next-generation techniques have generally supplanted reverse northern blotting, it is still utilized today and provides a relatively cheap and easy means of defining expression of large sets of genes.
The northwestern blot, also known as the northwestern assay, is a hybrid analytical technique of the western blot and the northern blot, and is used in molecular biology to detect interactions between RNA and proteins. A related technique, the western blot, is used to detect a protein of interest that involves transferring proteins that are separated by gel electrophoresis onto a nitrocellulose membrane. A colored precipitate clusters along the band on the membrane containing a particular target protein. A northern blot is a similar analytical technique that, instead of detecting a protein of interest, is used to study gene expression by detection of RNA on a similar membrane. The northwestern blot combines the two techniques, and specifically involves the identification of labeled RNA that interact with proteins that are immobilized on a similar nitrocellulose membrane.
In molecular biology, hybridization is a phenomenon in which single-stranded deoxyribonucleic acid (DNA) or ribonucleic acid (RNA) molecules anneal to complementary DNA or RNA. Though a double-stranded DNA sequence is generally stable under physiological conditions, changing these conditions in the laboratory will cause the molecules to separate into single strands. These strands are complementary to each other but may also be complementary to other sequences present in their surroundings. Lowering the surrounding temperature allows the single-stranded molecules to anneal or “hybridize” to each other.
References
- Material safety data sheet for 20x SSC buffer, Mediatech, Inc.
- Molecular Cytogenetics and biology Buffers, University of Leicester