
A wide variety of non-coding RNAs have been identified in various species of organisms known to science. However, RNAs have also been identified in "metagenomics" sequences derived from samples of DNA or RNA extracted from the environment, which contain unknown species. Initial work in this area detected homologs of known bacterial RNAs in such metagenome samples. Many of these RNA sequences were distinct from sequences within cultivated bacteria, and provide the potential for additional information on the RNA classes to which they belong.
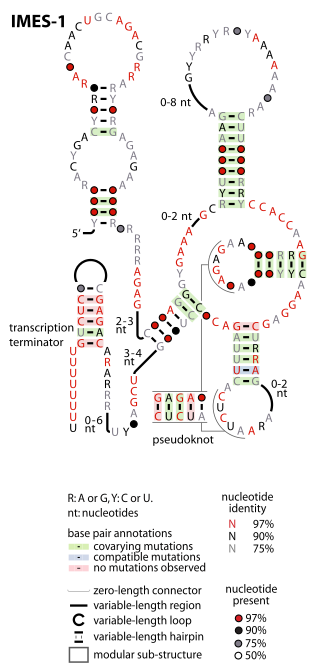
The IMES-1 RNA motif is a conserved RNA structure that was identified in marine environmental sequences by two studies based on metagenomics and bioinformatics, the first analyzing metatranscriptome (RNA) data and the second using metagenome (DNA) data. These RNAs are present in environmental sequences, and as of 2009 are not known to be present in any cultivated species. However, the species that use these RNAs are most closely related to known alphaproteobacteria and gammaproteobacteria. IMES-1 RNAs make up a significant portion of marine RNA transcripts and are exceptionally abundant in that over five times as many IMES-1 RNAs were found as ribosomes in RNAs sampled from the Pacific Ocean. Only two bacterial RNAs are known to be more highly transcribed than ribosomes. IMES-1 RNAs were also detected in abundance in Block Island Sound in the Atlantic Ocean.
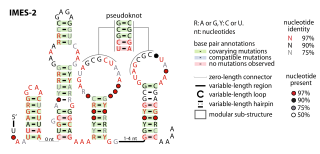
The IMES-2 RNA motif is a conserved RNA structure that was identified by a study based on metagenomics and bioinformatics, and the underlying RNA sequences were identified independently by a similar earlier study. These RNAs are present in environmental sequences, and when discovered were not known to be present in any cultivated species. However, an IMES-2 RNA has been detected in alphaproteobacterium HIMB114, which is classified in the SAR11 clade of marine bacteria. This finding fits with earlier predictions that species that use IMES-2 RNAs are most closely related to alphaproteobacteria. IMES-2 RNAs are exceptionally abundant, as twice as many IMES-2 RNAs were found as ribosomes in RNAs sampled from the Pacific Ocean. Only two bacterial RNAs are known to be more highly transcribed than ribosomes.

The IMES-3 RNA motif is a conserved RNA structure that was identified based on metagenomics and bioinformatics, and the underlying RNA sequences were identified independently by an earlier study. These RNAs are present in environmental sequences, and as of 2009 are not known to be present in any cultivated species. IMES-3 RNAs are abundant in comparison to ribosomes in RNAs sampled from the Pacific Ocean.
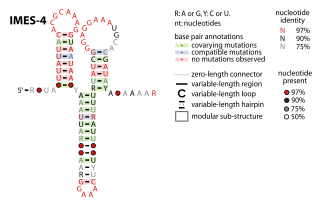
The IMES-4 RNA motif is a conserved RNA structure that was identified in marine environmental sequences by metagenomics and bioinformatics. These RNAs are present in environmental sequences, and as of 2009 are not known to be present in any cultivated species. IMES-4 RNAs are fairly abundant in comparison to ribosomes in RNAs sampled from the Pacific Ocean.

The Acido-1 RNA motif is a conserved RNA structure identified by bioinformatics. It is found only in acidobacteriota, and appears to be a non-coding RNA as it does not have a consistent association with protein-coding genes.

The Bacteroidales-1 RNA motif is a conserved RNA structure identified by bioinformatics. It has been identified only in bacteria within the order (biology) Bacteroidales. Its presumed length is marked by a promoter on one end that conforms to an alternate consensus sequence that is common in the phylum Bacteroidota, and its 3′ end is indicated by predicted transcription terminators. It is often located downstream of a gene that encodes the L20 ribosomal subunit, although it is unclear whether there is a functional reason underlying this apparent association.

The Bacteroides-1 RNA motif is a conserved RNA structure identified in bacteria within the genus Bacteroides. The RNAs are typically found downstream of of genes that participate in the synthesis of exopolysaccharides of unknown types. It is possible that Bacteroides-1 RNAs regulate the upstream genes, but since this mode of regulation is unusual in bacteria, it is more likely that the structure functions as a non-coding RNA.
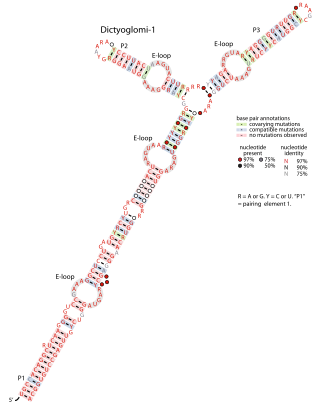
The Dictyoglomi-1 RNA motif is a conserved RNA structure that was discovered via bioinformatics. Only four instances of the RNA were detected, and all are in the bacterial phylum Dictyoglomota, whose members have not been extensively studied. The RNA might have a cis-regulatory role, but the evidence is ambiguous. Because of the few instances of Dictyoglomi-1 RNAs known, it is also unknown whether the RNA structure might extend further in the 5′ or 3′ direction, or in both directions.
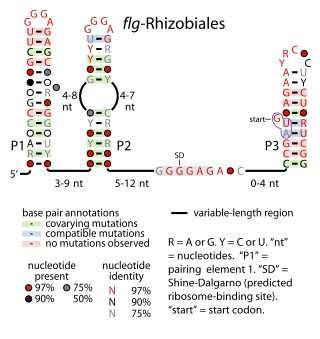
The flg-Rhizobiales RNA motif is an RNA structure that is conserved in certain bacteria. All known flg-Rhizobiales RNAs are located in the presumptive 5' untranslated regions of operons that contain genes whose functions relate to the creation of flagellar basal bodies. The flg-Rhizobiales RNAs are restricted to the Hyphomicrobiales, an order of alphaproteobacteria, although only some Rhizobiales bacterial are predicted to use flg-Rhizobiales RNAs. The exact function of these RNAs is unknown, although it is hypothesized that they have a cis-regulatory function in controlling expression of the downstream operons.

The Gut-1 RNA motif is a conserved RNA structure identified by bioinformatics. These RNAs are present in environmental sequences, and as of 2010 are not known to be present in any species that has been grown under laboratory conditions. Gut-1 RNA is exclusively found in DNA from uncultivated bacteria present in samples from the human gut.

The L17 downstream element RNA motif is a conserved RNA structure identified in bacteria by bioinformatics. All known L17 downstream elements were detected immediately downstream of genes encoding the L17 subunit of the ribosome, and therefore might be in the 3' untranslated regions of these genes. The element is found in a variety of lactic acid bacteria and in the genus Listeria.

The Lacto-rpoB RNA motif is a conserved RNA structure identified by bioinformatics. It has been detected only in lactic acid bacteria, and is always located in the presumed 5' untranslated regions of rpoB genes. These genes encode a subunit of RNA polymerase, and it is hypothesized that Lacto-rpoB RNA participate in the regulation of these genes.

The manA RNA motif refers to a conserved RNA structure that was identified by bioinformatics. Instances of the manA RNA motif were detected in bacteria in the genus Photobacterium and phages that infect certain kinds of cyanobacteria. However, most predicted manA RNA sequences are derived from DNA collected from uncultivated marine bacteria. Almost all manA RNAs are positioned such that they might be in the 5' untranslated regions of protein-coding genes, and therefore it was hypothesized that manA RNAs function as cis-regulatory elements. Given the relative complexity of their secondary structure, and their hypothesized cis-regulatory role, they might be riboswitches.
The wcaG RNA motif is an RNA structure conserved in some bacteria that was detected by bioinformatics. wcaG RNAs are found in certain phages that infect cyanobacteria. Most known wcaG RNAs were found in sequences of DNA extracted from uncultivated marine bacteria. wcaG RNAs might function as cis-regulatory elements, in view of their consistent location in the possible 5' untranslated regions of genes. It was suggested the wcaG RNAs might further function as riboswitches.

The Rhizobiales-2 RNA motif is a set of RNAs found in certain bacteria that are presumed to be homologous because they conserve a common primary and secondary structure. The motif was discovered using bioinformatics, and is found only within bacteria that belong to the order Hyphomicrobiales, in turn a kind of alphaproteobacteria. Because Rhizobiales-2 RNAs are not consistently located in proximity to genes of a consistent class or function, these RNAs are presumed to function as non-coding RNAs.
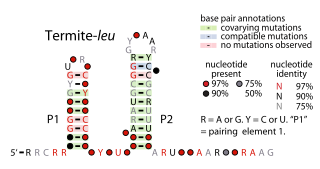
The Termite-leu RNA motif is a conserved RNA structure discovered by bioinformatics. It is found only in DNA sequences extracted from uncultivated bacteria living in termite hindguts, and has not yet been detected in any known cultivated organism. In many cases, Termite-leu RNAs are found in the likely 5′ untranslated regions of multive genes related to the synthesis of the amino acid leucine. However, in several cases it is not found in this type of location. Therefore, it was considered ambiguous as to whether Termite-leu RNAs constitute cis-regulatory elements.

The Whalefall-1 RNA motif refers to a conserved RNA structure that was discovered using bioinformatics. Structurally, the motif consists of two stem-loops, the second of which is often terminated by a CUUG tetraloop, which is an energetically favorable RNA sequence. Whalefall-1 RNAs are found only in DNA extracted from uncultivated bacteria found on whale fall, i.e., a whale carcass. As of 2010, Whalefall-1 RNAs have not been detected in any known, cultivated species of bacteria, and are thus one of several RNAs present in environmental samples.
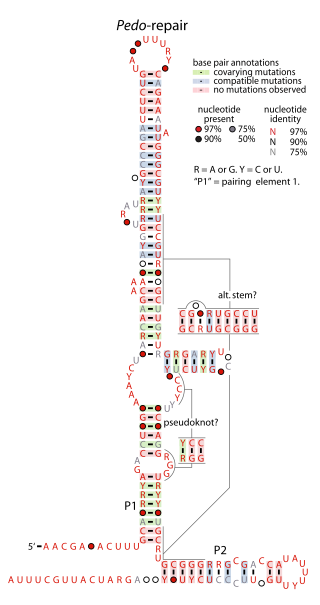
The Pedo-repair RNA motif is a conserved RNA structure identified by using bioinformatics. It has been detected in only one species of bacteria: Pedobacter sp. BAL39, within the phylum Bacteroidota. The motif might be in the 5′ untranslated regions of operons containing genes predicted to be involved in DNA repair or related to restriction enzymes.
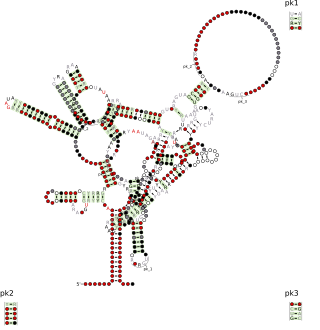
The Rumen-Originating, Ornate, Large (ROOL) RNA motif was originally discovered by bioinformatics by analyzing metagenomic sequences from cow rumen. ROOL RNAs are found in a variety of bacterial species and apparently do not code for proteins. The RNA has a complex RNA secondary structure and its average size of 581 nucleotides is unusually large for bacterial non-coding RNAs. This large size and structural complexity for a bacterial RNA is consistent with properties of large ribozymes.



















