Related Research Articles

The endoplasmic reticulum (ER) is, in essence, the transportation system of the eukaryotic cell, and has many other important functions such as protein folding. It is a type of organelle made up of two subunits – rough endoplasmic reticulum (RER), and smooth endoplasmic reticulum (SER). The endoplasmic reticulum is found in most eukaryotic cells and forms an interconnected network of flattened, membrane-enclosed sacs known as cisternae, and tubular structures in the SER. The membranes of the ER are continuous with the outer nuclear membrane. The endoplasmic reticulum is not found in red blood cells, or spermatozoa.
Inositol trisphosphate or inositol 1,4,5-trisphosphate abbreviated InsP3 or Ins3P or IP3 is an inositol phosphate signaling molecule. It is made by hydrolysis of phosphatidylinositol 4,5-bisphosphate (PIP2), a phospholipid that is located in the plasma membrane, by phospholipase C (PLC). Together with diacylglycerol (DAG), IP3 is a second messenger molecule used in signal transduction in biological cells. While DAG stays inside the membrane, IP3 is soluble and diffuses through the cell, where it binds to its receptor, which is a calcium channel located in the endoplasmic reticulum. When IP3 binds its receptor, calcium is released into the cytosol, thereby activating various calcium regulated intracellular signals.
Calcium release-activated channels (CRAC) are specialized plasma membrane Ca2+ ion channels. When calcium ions (Ca2+) are depleted from the endoplasmic reticulum (a major store of Ca2+) of mammalian cells, the CRAC channel is activated to slowly replenish the level of calcium in the endoplasmic reticulum. The Ca2+ Release-activated Ca2+ (CRAC) Channel (CRAC-C) Family (TC# 1.A.52) is a member of the Cation Diffusion Facilitator (CDF) Superfamily. These proteins typically have between 4 and 6 transmembrane α-helical spanners (TMSs). The 4 TMS CRAC channels arose by loss of 2TMSs from 6TMS CDF carriers, an example of 'reverse' evolution'.
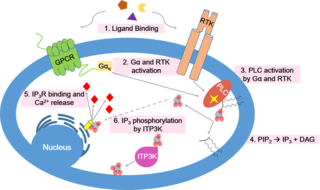
Calcium signaling is the use of calcium ions (Ca2+) to communicate and drive intracellular processes often as a step in signal transduction. Ca2+ is important for cellular signalling, for once it enters the cytosol of the cytoplasm it exerts allosteric regulatory effects on many enzymes and proteins. Ca2+ can act in signal transduction resulting from activation of ion channels or as a second messenger caused by indirect signal transduction pathways such as G protein-coupled receptors.

Ca2+ ATPase is a form of P-ATPase that transfers calcium after a muscle has contracted. The two kinds of calcium ATPase are:
TRPC is a family of transient receptor potential cation channels in animals.
Store-operated channels (SOCs) are ion channels located in the plasma membrane of cells. These channels are most studied in regard to their role in calcium entry into the cytoplasm from extracellular milieu. There are other SOC channels selective to other ions. Calcium SOCs are especially important for the cell because they are the major source of intracellular calcium; and calcium itself is involved in a wide array of vital cellular functions. SOCs are so called because they are activated by intracellular calcium stores depletion by both physiological or pharmacological processes.

Stromal interaction molecule 1 is a protein that in humans is encoded by the STIM1 gene. STIM1 has a single transmembrane domain, and is localized to the endoplasmic reticulum, and to a lesser extent to the plasma membrane.

Calcium release-activated calcium channel protein 1 is a calcium selective ion channel that in humans is encoded by the ORAI1 gene. Orai channels play an important role in the activation of T-lymphocytes. The loss of function mutation of Orai1 causes severe combined immunodeficiency (SCID) in humans The mammalian orai family has two additional homologs, Orai2 and Orai3. Orai proteins share no homology with any other ion channel family of any other known proteins. They have 4 transmembrane domains and form hexamers.

Sarcoplasmic/endoplasmic reticulum calcium ATPase 3 is an enzyme that in humans is encoded by the ATP2A3 gene.

Stromal interaction molecule 2 (STIM2) is a protein that in humans is encoded by the STIM2 gene.
The spine apparatus (SA) is a specialized form of endoplasmic reticulum (ER) that is found in a subpopulation of dendritic spines in central neurons. It was discovered by Edward George Gray in 1959 when he applied electron microscopy to fixed cortical tissue. The SA consists of a series of stacked discs that are connected to each other and to the dendritic system of ER-tubules. The actin binding protein synaptopodin is an essential component of the SA. Mice that lack the gene for synaptopodin do not form a spine apparatus. The SA is believed to play a role in synaptic plasticity, learning and memory, but the exact function of the spine apparatus is still enigmatic.
Membrane contact sites (MCS) are close appositions between two organelles. Ultrastructural studies typically reveal an intermembrane distance in the order of the size of a single protein, as small as 10 nm or wider, with no clear upper limit. These zones of apposition are highly conserved in evolution. These sites are thought to be important to facilitate signalling, and they promote the passage of small molecules, including ions, lipids and reactive oxygen species. MCS are important in the function of the endoplasmic reticulum (ER), since this is the major site of lipid synthesis within cells. The ER makes close contact with many organelles, including mitochondria, Golgi, endosomes, lysosomes, peroxisomes, chloroplasts and the plasma membrane. Both mitochondria and sorting endosomes undergo major rearrangements leading to fission where they contact the ER. Sites of close apposition can also form between most of these organelles most pairwise combinations. First mentions of these contact sites can be found in papers published in the late 1950s mainly visualized using electron microscopy (EM) techniques. Copeland and Dalton described them as “highly specialized tubular form of endoplasmic reticulum in association with the mitochondria and apparently in turn, with the vascular border of the cell”.
LiMETER stands for light-inducible membrane-tethered peripheral endoplasmic reticulum (ER). LiMETER is an optogenetics tool designed to reversibly label cortical ER or the apposition between plasma membrane (PM) and endoplasmic reticulum (ER) membranes.

Mitochondria-associated membranes (MAM) represent a region of the endoplasmic reticulum (ER) which is reversibly tethered to mitochondria. These membranes are involved in import of certain lipids from the ER to mitochondria and in regulation of calcium homeostasis, mitochondrial function, autophagy and apoptosis. They also play a role in development of neurodegenerative diseases and glucose homeostasis.

ORMDL sphingolipid biosynthesis regulator 3 is a protein that in humans is encoded by the ORMDL3 gene. This gene is associated with asthma in childhood. Transgenic mice which overexpress human ORMDL3 have increased levels of IgE. This correlated with increased numbers of macrophages, neutrophils, eosinophils, CD4+ and enhanced Th2 cytokine levels in the lung tissue.
Anjana Rao is a cellular and molecular biologist of Indian ethnicity, working in the US. She uses immune cells as well as other types of cells to understand intracellular signaling and gene expression. Her research focuses on how signaling pathways control gene expression.
GRAM domain-containing 2A protein is a protein encoded by the GRAMD2A gene. Like GRAMD2B, the protein consists of a GRAM domain and a transmembrane domain that anchors it to the endoplasmic reticulum.
Nir1 or Membrane-associated phosphatidylinositol transfer protein 3 (PITPNM3) is a mammalian protein that localizes to endoplasmic reticulum (ER) and plasma membrane (PM) membrane contact sites (MCS) and aids the transfer of phosphatidylinositol between these two membranes, potentially by recruiting additional proteins to the ER-PM MCS.
Patrick G. Hogan is a cellular and molecular biologist who studies how cellular signaling leads to gene expression. He obtained his bachelor’s degree from Harvard University and a PhD in neurobiology from Harvard Medical School. In 2010, he moved to the La Jolla Institute for Immunology in San Diego as a Professor in the Division of Signaling and Gene Expression. He is a Founder and Member of the Scientific Advisory Board, CalciMedica Inc, La Jolla, CA.
References
- 1 2 Jing, Ji; He, Lian; Sun, Aomin; Quintana, Ariel; Ding, Yuehe; Ma, Guolin; Tan, Peng; Liang, Xiaowen; Zheng, Xiaolu (2015-01-01). "Proteomic mapping of ER–PM junctions identifies STIMATE as a regulator of Ca2+ influx". Nature Cell Biology. 17 (10): 1339–47. doi:10.1038/ncb3234. PMC 4589512 . PMID 26322679.
- ↑ Carrasco, Silvia; Meyer, Tobias (2011-01-01). "STIM Proteins and the Endoplasmic Reticulum-Plasma Membrane Junctions". Annual Review of Biochemistry. 80 (1): 973–1000. doi:10.1146/annurev-biochem-061609-165311. PMC 3897197 . PMID 21548779.
- ↑ Zhou, Yubin; Srinivasan, Prasanna; Razavi, Shiva; Seymour, Sam; Meraner, Paul; Gudlur, Aparna; Stathopulos, Peter B; Ikura, Mitsuhiko; Rao, Anjana (2013-01-01). "Initial activation of STIM1, the regulator of store-operated calcium entry". Nature Structural & Molecular Biology. 20 (8): 973–981. doi:10.1038/nsmb.2625. PMC 3784406 . PMID 23851458.
- ↑ Ma, Guolin; Wei, Ming; He, Lian; Liu, Chongxu; Wu, Bo; Zhang, Shenyuan L.; Jing, Ji; Liang, Xiaowen; Senes, Alessandro (2015-07-17). "Inside-out Ca2+ signalling prompted by STIM1 conformational switch". Nature Communications. 6: 7826. Bibcode:2015NatCo...6.7826M. doi:10.1038/ncomms8826. PMC 4509486 . PMID 26184105.
- ↑ Müller, Martin R.; Rao, Anjana (2010-01-01). "NFAT, immunity and cancer: a transcription factor comes of age". Nature Reviews Immunology. 10 (9): 645–656. doi:10.1038/nri2818. PMID 20725108. S2CID 29256284.
- ↑ Oh-hora, Masatsugu; Rao, Anjana (2008-01-01). "Calcium signaling in lymphocytes". Current Opinion in Immunology. 20 (3): 250–258. doi:10.1016/j.coi.2008.04.004. PMC 2574011 . PMID 18515054.