Related Research Articles

A chemical bond is a lasting attraction between atoms or ions that enables the formation of molecules, crystals, and other structures. The bond may result from the electrostatic force between oppositely charged ions as in ionic bonds, or through the sharing of electrons as in covalent bonds. The strength of chemical bonds varies considerably; there are "strong bonds" or "primary bonds" such as covalent, ionic and metallic bonds, and "weak bonds" or "secondary bonds" such as dipole–dipole interactions, the London dispersion force, and hydrogen bonding.
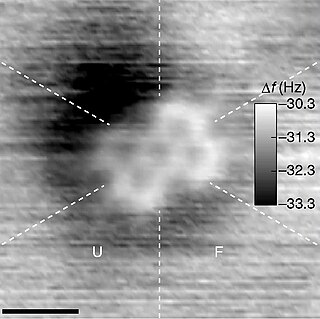
A molecule is a group of two or more atoms held together by attractive forces known as chemical bonds; depending on context, the term may or may not include ions which satisfy this criterion. In quantum physics, organic chemistry, and biochemistry, the distinction from ions is dropped and molecule is often used when referring to polyatomic ions.
Quantum chemistry, also called molecular quantum mechanics, is a branch of physical chemistry focused on the application of quantum mechanics to chemical systems, particularly towards the quantum-mechanical calculation of electronic contributions to physical and chemical properties of molecules, materials, and solutions at the atomic level. These calculations include systematically applied approximations intended to make calculations computationally feasible while still capturing as much information about important contributions to the computed wave functions as well as to observable properties such as structures, spectra, and thermodynamic properties. Quantum chemistry is also concerned with the computation of quantum effects on molecular dynamics and chemical kinetics.

Theoretical chemistry is the branch of chemistry which develops theoretical generalizations that are part of the theoretical arsenal of modern chemistry: for example, the concepts of chemical bonding, chemical reaction, valence, the surface of potential energy, molecular orbitals, orbital interactions, and molecule activation.

Molecular dynamics (MD) is a computer simulation method for analyzing the physical movements of atoms and molecules. The atoms and molecules are allowed to interact for a fixed period of time, giving a view of the dynamic "evolution" of the system. In the most common version, the trajectories of atoms and molecules are determined by numerically solving Newton's equations of motion for a system of interacting particles, where forces between the particles and their potential energies are often calculated using interatomic potentials or molecular mechanical force fields. The method is applied mostly in chemical physics, materials science, and biophysics.

Structural bioinformatics is the branch of bioinformatics that is related to the analysis and prediction of the three-dimensional structure of biological macromolecules such as proteins, RNA, and DNA. It deals with generalizations about macromolecular 3D structures such as comparisons of overall folds and local motifs, principles of molecular folding, evolution, binding interactions, and structure/function relationships, working both from experimentally solved structures and from computational models. The term structural has the same meaning as in structural biology, and structural bioinformatics can be seen as a part of computational structural biology. The main objective of structural bioinformatics is the creation of new methods of analysing and manipulating biological macromolecular data in order to solve problems in biology and generate new knowledge.
Molecular modelling encompasses all methods, theoretical and computational, used to model or mimic the behaviour of molecules. The methods are used in the fields of computational chemistry, drug design, computational biology and materials science to study molecular systems ranging from small chemical systems to large biological molecules and material assemblies. The simplest calculations can be performed by hand, but inevitably computers are required to perform molecular modelling of any reasonably sized system. The common feature of molecular modelling methods is the atomistic level description of the molecular systems. This may include treating atoms as the smallest individual unit, or explicitly modelling protons and neutrons with its quarks, anti-quarks and gluons and electrons with its photons.
Quantitative structure–activity relationship models are regression or classification models used in the chemical and biological sciences and engineering. Like other regression models, QSAR regression models relate a set of "predictor" variables (X) to the potency of the response variable (Y), while classification QSAR models relate the predictor variables to a categorical value of the response variable.
Supramolecular chemistry refers to the branch of chemistry concerning chemical systems composed of a discrete number of molecules. The strength of the forces responsible for spatial organization of the system range from weak intermolecular forces, electrostatic charge, or hydrogen bonding to strong covalent bonding, provided that the electronic coupling strength remains small relative to the energy parameters of the component. While traditional chemistry concentrates on the covalent bond, supramolecular chemistry examines the weaker and reversible non-covalent interactions between molecules. These forces include hydrogen bonding, metal coordination, hydrophobic forces, van der Waals forces, pi–pi interactions and electrostatic effects.
Mathematical chemistry is the area of research engaged in novel applications of mathematics to chemistry; it concerns itself principally with the mathematical modeling of chemical phenomena. Mathematical chemistry has also sometimes been called computer chemistry, but should not be confused with computational chemistry.
In quantum chemistry, the quantum theory of atoms in molecules (QTAIM), sometimes referred to as atoms in molecules (AIM), is a model of molecular and condensed matter electronic systems in which the principal objects of molecular structure - atoms and bonds - are natural expressions of a system's observable electron density distribution function. An electron density distribution of a molecule is a probability distribution that describes the average manner in which the electronic charge is distributed throughout real space in the attractive field exerted by the nuclei. According to QTAIM, molecular structure is revealed by the stationary points of the electron density together with the gradient paths of the electron density that originate and terminate at these points.
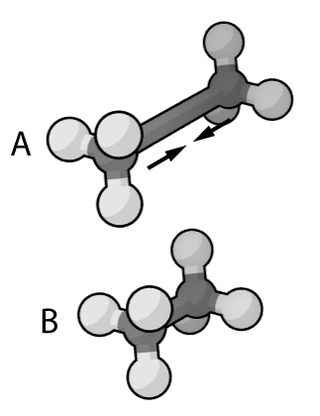
In the context of chemistry and molecular modelling, a force field is a computational method that is used to estimate the forces between atoms within molecules and also between molecules. More precisely, the force field refers to the functional form and parameter sets used to calculate the potential energy of a system of atoms or coarse-grained particles in molecular mechanics, molecular dynamics, or Monte Carlo simulations. The parameters for a chosen energy function may be derived from experiments in physics and chemistry, calculations in quantum mechanics, or both. Force fields are interatomic potentials and utilize the same concept as force fields in classical physics, with the difference that the force field parameters in chemistry describe the energy landscape, from which the acting forces on every particle are derived as a gradient of the potential energy with respect to the particle coordinates.

Cation–π interaction is a noncovalent molecular interaction between the face of an electron-rich π system (e.g. benzene, ethylene, acetylene) and an adjacent cation (e.g. Li+, Na+). This interaction is an example of noncovalent bonding between a monopole (cation) and a quadrupole (π system). Bonding energies are significant, with solution-phase values falling within the same order of magnitude as hydrogen bonds and salt bridges. Similar to these other non-covalent bonds, cation–π interactions play an important role in nature, particularly in protein structure, molecular recognition and enzyme catalysis. The effect has also been observed and put to use in synthetic systems.
Physical organic chemistry, a term coined by Louis Hammett in 1940, refers to a discipline of organic chemistry that focuses on the relationship between chemical structures and reactivity, in particular, applying experimental tools of physical chemistry to the study of organic molecules. Specific focal points of study include the rates of organic reactions, the relative chemical stabilities of the starting materials, reactive intermediates, transition states, and products of chemical reactions, and non-covalent aspects of solvation and molecular interactions that influence chemical reactivity. Such studies provide theoretical and practical frameworks to understand how changes in structure in solution or solid-state contexts impact reaction mechanism and rate for each organic reaction of interest.
In the fields of chemical graph theory, molecular topology, and mathematical chemistry, a topological index, also known as a connectivity index, is a type of a molecular descriptor that is calculated based on the molecular graph of a chemical compound. Topological indices are numerical parameters of a graph which characterize its topology and are usually graph invariant. Topological indices are used for example in the development of quantitative structure-activity relationships (QSARs) in which the biological activity or other properties of molecules are correlated with their chemical structure.

In chemistry, isomers are molecules or polyatomic ions with identical molecular formula – that is, same number of atoms of each element – but distinct arrangements of atoms in space. Isomerism refers to the existence or possibility of isomers.

The circuit topology of a folded linear polymer refers to the arrangement of its intra-molecular contacts. Examples of linear polymers with intra-molecular contacts are nucleic acids and proteins. Proteins fold via formation of contacts of various nature, including hydrogen bonds, disulfide bonds, and beta-beta interactions. RNA molecules fold by forming hydrogen bonds between nucleotides, forming nested or non-nested structures. Contacts in the genome are established via protein bridges including CTCF and cohesins and are measured by technologies including Hi-C. Circuit topology categorises the topological arrangement of these physical contacts, that are referred to as hard contacts. Furthermore, chains can fold via knotting. Circuit topology uses a similar language to categorise both "soft" and "hard" contacts, and provides a full description of a folded linear chain. In this framework, a "circuit" refers to a segment of the chain where each contact site within the segment forms connections with other contact sites within the same segment, and thus is not left unpaired. A folded chain can thus be studied based on its constituting circuits.
Matched molecular pair analysis (MMPA) is a method in cheminformatics that compares the properties of two molecules that differ only by a single chemical transformation, such as the substitution of a hydrogen atom by a chlorine one. Such pairs of compounds are known as matched molecular pairs (MMP). Because the structural difference between the two molecules is small, any experimentally observed change in a physical or biological property between the matched molecular pair can more easily be interpreted. The term was first coined by Kenny and Sadowski in the book Chemoinformatics in Drug Discovery.
In chemistry, a chalcogen bond (ChB) is an attractive interaction in the family of σ-hole interactions, along with halogen bonds. Electrostatic, charge-transfer (CT) and dispersion terms have been identified as contributing to this type of interaction. In terms of CT contribution, this family of attractive interactions has been modeled as an electron donor ) interacting with the σ* orbital of a C-X bond of the bond donor. In terms of electrostatic interactions, the molecular electrostatic potential (MEP) maps is often invoked to visualize the electron density of the donor and an electrophilic region on the acceptor, where the potential is depleted, referred to as a σ-hole. ChBs, much like hydrogen and halogen bonds, have been invoked in various non-covalent interactions, such as protein folding, crystal engineering, self-assembly, catalysis, transport, sensing, templation, and drug design.
Topological polymers may refer to a polymeric molecule that possesses unique spatial features, such as linear, branched, or cyclic architectures. It could also refer to polymer networks that exhibit distinct topologies owing to special crosslinkers. When self-assembling or crosslinking in a certain way, polymeric species with simple topological identity could also demonstrate complicated topological structures in a larger spatial scale. Topological structures, along with the chemical composition, determine the macroscopic physical properties of polymeric materials.
References
- ↑ Brown, David; Topology and Chemistry; Structural Chemistry Volume 13, Numbers 3–4, 339–355, doi : 10.1023/A:1015872125545
- Francl, Michelle; Stretching topology Nature Chemistry 1, 334–335 (2009) doi : 10.1038/nchem.302
- Rouvray, D. H.; A rationale for the topological approach to chemistry; Journal of Molecular Structure: THEOCHEM Volume 336, Issues 2–3, 30 June 1995, pages 101–114