
Retroviral integrase (IN) is an enzyme produced by a retrovirus that integrates its genetic information into that of the host cell it infects. Retroviral INs are not to be confused with phage integrases (recombinases) used in biotechnology, such as λ phage integrase, as discussed in site-specific recombination.
Cis-regulatory elements (CREs) or cis-regulatory modules (CRMs) are regions of non-coding DNA which regulate the transcription of neighboring genes. CREs are vital components of genetic regulatory networks, which in turn control morphogenesis, the development of anatomy, and other aspects of embryonic development, studied in evolutionary developmental biology.

The Ssbp, Topoisomerase, Antirestriction, XerDC Integrase RNA motif is a conserved RNA-like structure identified using bioinformatics. STAXI RNAs are located near to genes encoding proteins that interact with DNA or are associated with such proteins. This observation raised the possibility that instances of the STAXI motif function as single-stranded DNA molecules, perhaps during DNA replication or DNA repair. On the other hand, STAXI motifs often contain terminal loops conforming to the stable UNCG tetraloop, but the DNA version of this tetraloop (TNCG) is not especially stable. The STAXI motif consists of a simple pseudoknot structure that is repeated two or more times.

The che1 RNA motif is a conserved RNA structure that was discovered by bioinformatics. che1 motifs are found in the genus Streptomyces, as well as some other organisms that are closely related to this genus.

The COG3610-DE RNA motif is a conserved RNA structure that was discovered by bioinformatics. COG3610-DE motifs are found in the genus Lactobacillales, which is part of the phylum Bacillota.

The COG3943 RNA motif is a conserved RNA structure that was discovered by bioinformatics. COG3943 motifs are found in unknown bacteria whose genomic DNA was isolated from cow rumen. As of 2018, there is no specific, classified organism that is known to contain a COG3943 motif RNA.

The D12-methyl RNA motif is a conserved RNA structure that was discovered by bioinformatics. D12-methyl motifs are found in metagenomic DNA samples, and have not yet been found in a classified organism.

The dinG RNA motif is a conserved RNA structure that was discovered by bioinformatics. dinG motifs are found in Clostridiales.

The DUF2693 RNA motif is a conserved RNA structure that was discovered by bioinformatics. DUF2693 motif RNAs are found in Porphyromonas.
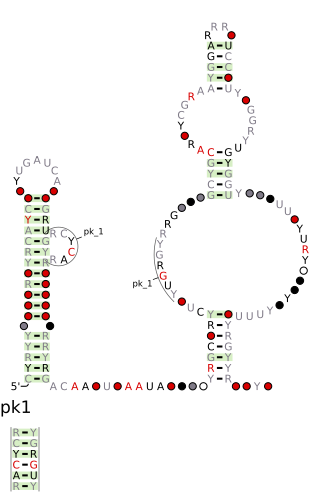
The DUF2800 RNA motif is a conserved RNA structure that was discovered by bioinformatics. DUF2800 motif RNAs are found in Bacillota. DUF2800 RNAs are also predicted in the phyla Actinomycetota and Synergistota, although these RNAs are likely the result of recent horizontal gene transfer or conceivably sequence contamination.

The DUF2815 RNA motif is a conserved RNA structure that was discovered by bioinformatics. As of 2018, the DUF2815 motif has not been identified in any classified organism, but is known through metagenomic sequences isolated from environmental sources.

The DUF805 RNA motif is a conserved RNA structure that was discovered by bioinformatics. The motif is subdivided into the DUF805 motif and the DUF805b motif, which have similar, but distinct secondary structures. Together, these motifs are found in Bacteroidota, Chlorobiota, and Pseudomonadota.
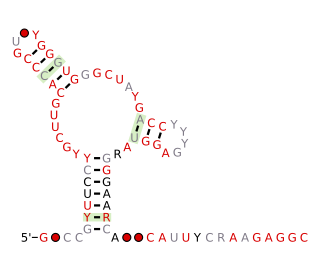
The Fibro-purF RNA motif is a conserved RNA structure that was discovered by bioinformatics. Fibro-purF motif RNAs are found in Fibrobacterota, a group of bacteria that are common in cow rumen. Additionally, the RNAs are found in metagenomic sequences of DNA isolated from cow rumen.
The freshwater-2 RNA motif is a conserved RNA structure that was discovered by bioinformatics. Freshwater-2 motif RNAs are found in metagenomic sequences that are isolated from aquatic and especially freshwater environments. As of 2018, no freshwater-2 RNA has been identified in a classified organism.

The GA-cis RNA motif is a conserved RNA structure that was discovered by bioinformatics. GA-cis motif RNAs are found in one species classified within the phylum Bacillota: specifically, there are 9 predicted copies in Coprocuccus eutactus ATCC 27759.

The gntR-DTE RNA motif is a conserved RNA structure that was discovered by bioinformatics. gntR-DTE motifs are found in some, but not all species within the genus Streptomyces.
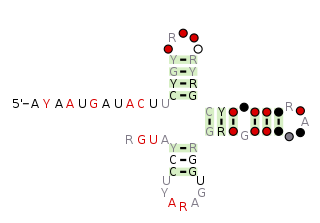
The pemK RNA motif is a conserved RNA structure that was discovered by bioinformatics. pemK motif RNAs are found in organisms within the phylum Bacillota, and is very widespread in this phylum.
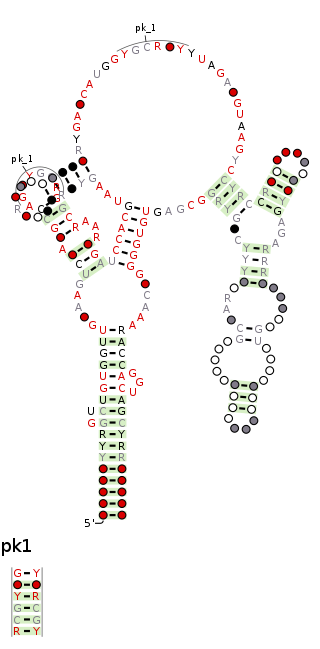
The raiA RNA motif is a conserved RNA structure that was discovered by bioinformatics. raiA motif RNAs are found in Actinomycetota and Bacillota, and have many conserved features—including conserved nucleotide positions, conserved secondary structures and associated protein-coding genes—in both of these phyla. Some conserved features of the raiA RNA motif suggest that they function as cis-regulatory elements, but other aspects of the motif suggest otherwise.

The ssNA-helicase RNA motif is a conserved RNA structure that was discovered by bioinformatics. Although the ssNA-helicase motif was published as an RNA candidate, there is some reason to suspect that it might function as a single-stranded DNA. In terms of secondary structure, RNA and DNA are difficult to distinguish when only sequence information is available.

The uup RNA motif is a conserved RNA structure that was discovered by bioinformatics. uup motif RNAs are found in Bacillota and Gammaproteobacteria.


















