Related Research Articles

Ribosomes ( ), also called Palade granules, are macromolecular machines, found within all cells, that perform biological protein synthesis. Ribosomes link amino acids together in the order specified by the codons of messenger RNA (mRNA) molecules to form polypeptide chains. Ribosomes consist of two major components: the small and large ribosomal subunits. Each subunit consists of one or more ribosomal RNA (rRNA) molecules and many ribosomal proteins. The ribosomes and associated molecules are also known as the translational apparatus.
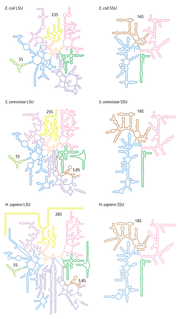
Ribosomal ribonucleic acid (rRNA) is a type of non-coding RNA which is the primary component of ribosomes, essential to all cells. rRNA is a ribozyme which carries out protein synthesis in ribosomes. Ribosomal RNA is transcribed from ribosomal DNA (rDNA) and then bound to ribosomal proteins to form small and large ribosome subunits. rRNA is the physical and mechanical factor of the ribosome that forces transfer RNA (tRNA) and messenger RNA (mRNA) to process and translate the latter into proteins. Ribosomal RNA is the predominant form of RNA found in most cells; it makes up about 80% of cellular RNA despite never being translated into proteins itself. Ribosomes are composed of approximately 60% rRNA and 40% ribosomal proteins by mass.
RNA polymerase 1 is, in higher eukaryotes, the polymerase that only transcribes ribosomal RNA, a type of RNA that accounts for over 50% of the total RNA synthesized in a cell.
Bacterial translation is the process by which messenger RNA is translated into proteins in bacteria.
Eukaryotic translation is the biological process by which messenger RNA is translated into proteins in eukaryotes. It consists of four phases: gene translation, elongation, termination, and recapping.
The Kozak consensus sequence is a nucleic acid motif that functions as the protein translation initiation site in most eukaryotic mRNA transcripts. Regarded as the optimum sequence for initiating translation in eukaryotes, the sequence is an integral aspect of protein regulation and overall cellular health as well as having implications in human disease. It ensures that a protein is correctly translated from the genetic message, mediating ribosome assembly and translation initiation. A wrong start site can result in non-functional proteins. As it has become more studied, expansions of the nucleotide sequence, bases of importance, and notable exceptions have arisen. The sequence was named after the scientist who discovered it, Marilyn Kozak. Kozak discovered the sequence through a detailed analysis of DNA genomic sequences.
Initiation factors are proteins that bind to the small subunit of the ribosome during the initiation of translation, a part of protein biosynthesis.
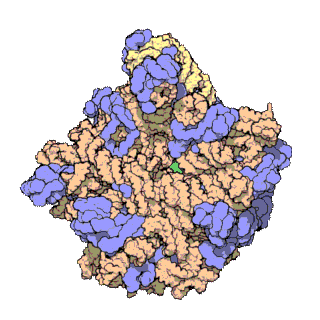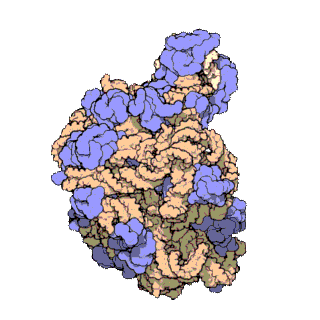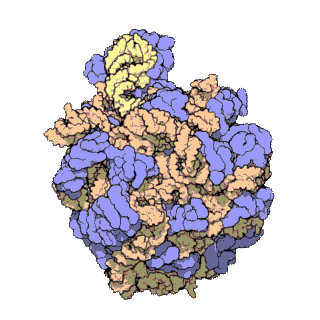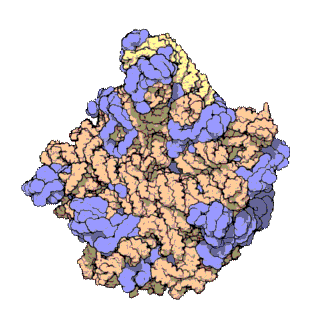
A ribosomal protein is any of the proteins that, in conjunction with rRNA, make up the ribosomal subunits involved in the cellular process of translation. E. coli, other bacteria and Archaea have a 30S small subunit and a 50S large subunit, whereas humans and yeasts have a 40S small subunit and a 60S large subunit. Equivalent subunits are frequently numbered differently between bacteria, Archaea, yeasts and humans.
Eukaryotic initiation factors (eIFs) are proteins or protein complexes involved in the initiation phase of eukaryotic translation. These proteins help stabilize the formation of ribosomal preinitiation complexes around the start codon and are an important input for post-transcription gene regulation. Several initiation factors form a complex with the small 40S ribosomal subunit and Met-tRNAiMet called the 43S preinitiation complex. Additional factors of the eIF4F complex recruit the 43S PIC to the five-prime cap structure of the mRNA, from which the 43S particle scans 5'-->3' along the mRNA to reach an AUG start codon. Recognition of the start codon by the Met-tRNAiMet promotes gated phosphate and eIF1 release to form the 48S preinitiation complex, followed by large 60S ribosomal subunit recruitment to form the 80S ribosome. There exist many more eukaryotic initiation factors than prokaryotic initiation factors, reflecting the greater biological complexity of eukaryotic translation. There are at least twelve eukaryotic initiation factors, composed of many more polypeptides, and these are described below.

The 23S rRNA is a 2,904 nucleotide long component of the large subunit (50S) of the bacterial/archean ribosome and makes up the peptidyl transferase center (PTC). The 23S rRNA is divided into six secondary structural domains titled I-VI, with the corresponding 5S rRNA being considered domain VII. The ribosomal peptidyl transferase activity resides in domain V of this rRNA, which is also the most common binding site for antibiotics that inhibit translation, making it a target for ribosomal engineering. A well-known member of this antibiotic class, chloramphenicol, acts by inhibiting peptide bond formation, with recent 3D-structural studies showing two different binding sites depending on the species of ribosome. Numerous mutations in domains of the 23S rRNA with Peptidyl transferase activity have resulted in antibiotic resistance. 23S rRNA genes typically have higher sequence variations, including insertions and/or deletions, compared to other rRNAs.

Eukaryotic translation initiation factor 6 (EIF6), also known as Integrin beta 4 binding protein (ITGB4BP), is a human gene.

60S ribosomal protein L24 is a protein that in humans is encoded by the RPL24 gene.

Ribosome maturation protein SBDS is a protein that in humans is encoded by the SBDS gene. An alternative transcript has been described, but its biological nature has not been determined. This gene has a closely linked pseudogene that is distally located. This gene encodes a member of a highly conserved protein family that exists from archaea to vertebrates and plants.

EF-G is a prokaryotic elongation factor involved in protein translation. As a GTPase, EF-G catalyzes the movement (translocation) of transfer RNA (tRNA) and messenger RNA (mRNA) through the ribosome.
The eukaryotic small ribosomal subunit (40S) is the smaller subunit of the eukaryotic 80S ribosomes, with the other major component being the large ribosomal subunit (60S). The "40S" and "60S" names originate from the convention that ribosomal particles are denoted according to their sedimentation coefficients in Svedberg units. It is structurally and functionally related to the 30S subunit of 70S prokaryotic ribosomes. However, the 40S subunit is much larger than the prokaryotic 30S subunit and contains many additional protein segments, as well as rRNA expansion segments.

A protein synthesis inhibitor is a compound that stops or slows the growth or proliferation of cells by disrupting the processes that lead directly to the generation of new proteins.
Eukaryotic Initiation Factor 2 (eIF2) is a eukaryotic initiation factor. It is required for most forms of eukaryotic translation initiation. eIF2 mediates the binding of tRNAiMet to the ribosome in a GTP-dependent manner. eIF2 is a heterotrimer consisting of an alpha, a beta, and a gamma subunit.
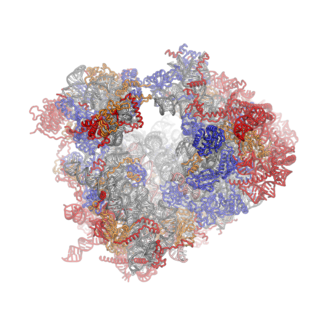
Ribosomes are a large and complex molecular machine that catalyzes the synthesis of proteins, referred to as translation. The ribosome selects aminoacylated transfer RNAs (tRNAs) based on the sequence of a protein-encoding messenger RNA (mRNA) and covalently links the amino acids into a polypeptide chain. Ribosomes from all organisms share a highly conserved catalytic center. However, the ribosomes of eukaryotes are much larger than prokaryotic ribosomes and subject to more complex regulation and biogenesis pathways. Eukaryotic ribosomes are also known as 80S ribosomes, referring to their sedimentation coefficients in Svedberg units, because they sediment faster than the prokaryotic (70S) ribosomes. Eukaryotic ribosomes have two unequal subunits, designated small subunit (40S) and large subunit (60S) according to their sedimentation coefficients. Both subunits contain dozens of ribosomal proteins arranged on a scaffold composed of ribosomal RNA (rRNA). The small subunit monitors the complementarity between tRNA anticodon and mRNA, while the large subunit catalyzes peptide bond formation.

In molecular biology, chaperone DnaJ, also known as Hsp40, is a molecular chaperone protein. It is expressed in a wide variety of organisms from bacteria to humans.

Nucleic acidquaternary structure refers to the interactions between separate nucleic acid molecules, or between nucleic acid molecules and proteins. The concept is analogous to protein quaternary structure, but as the analogy is not perfect, the term is used to refer to a number of different concepts in nucleic acids and is less commonly encountered. Similarly other biomolecules such as proteins, nucleic acids have four levels of structural arrangement: primary, secondary, tertiary, and quaternary structure. Primary structure is the linear sequence of nucleotides, secondary structure involves small local folding motifs, and tertiary structure is the 3D folded shape of nucleic acid molecule. In general, quaternary structure refers to 3D interactions between multiple subunits. In the case of nucleic acids, quaternary structure refers to interactions between multiple nucleic acid molecules or between nucleic acids and proteins. Nucleic acid quaternary structure is important for understanding DNA, RNA, and gene expression because quaternary structure can impact function. For example, when DNA is packed into chromatin, therefore exhibiting a type of quaternary structure, gene transcription will be inhibited.
References
- ↑ Zhang S, Lockshin C, Herbert A, Winter E, Rich A (October 1992). "Zuotin, a putative Z-DNA binding protein in Saccharomyces cerevisiae". The EMBO Journal. 11 (10): 3787–96. doi:10.1002/j.1460-2075.1992.tb05464.x. PMC 556839 . PMID 1396572.
- ↑ Wilhelm ML, Reinbolt J, Gangloff J, Dirheimer G, Wilhelm FX (August 1994). "Transfer RNA binding protein in the nucleus of Saccharomyces cerevisiae". FEBS Letters. 349 (2): 260–4. doi: 10.1016/0014-5793(94)00683-0 . PMID 8050578.
- ↑ Leidig C, Bange G, Kopp J, Amlacher S, Aravind A, Wickles S, et al. (January 2013). "Structural characterization of a eukaryotic chaperone--the ribosome-associated complex". Nature Structural & Molecular Biology. 20 (1): 23–8. doi:10.1038/nsmb.2447. PMID 23202586. S2CID 22950001.
- ↑ Lee K, Sharma R, Shrestha OK, Bingman CA, Craig EA (November 2016). "Dual interaction of the Hsp70 J-protein cochaperone Zuotin with the 40S and 60S ribosomal subunits". Nature Structural & Molecular Biology. 23 (11): 1003–1010. doi:10.1038/nsmb.3299. PMC 5097012 . PMID 27669034.