HSQC in protein NMR
1H—15N HSQC

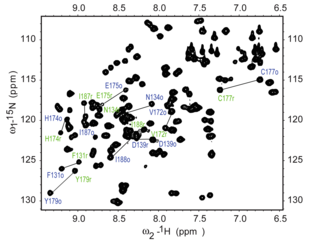
The 15N HSQC experiment is one of the most frequently recorded experiments in protein NMR. The HSQC experiment can be performed using the natural abundance of the 15N isotope, but normally for protein NMR, isotopically labeled proteins are used. Such labelled proteins are usually produced by expressing the protein in cells grown in 15N-labelled media.
Each residue of the protein, with the exception of proline, has an amide proton attached to a nitrogen in the peptide bond. The HSQC provides the correlation between the nitrogen and amide proton, and each amide yields a peak in the HSQC spectra. Each residue (except proline) therefore can produce an observable peak in the spectra, although in practice not all the peaks are always seen due to a number of factors. Normally the N-terminal residue (which has an NH3+ group attached) is not readily observable due to exchange with solvent. [3] In addition to the backbone amide resonances, sidechains with nitrogen-bound protons will also produce peaks.
In a typical HSQC spectrum, the NH2 peaks from the sidechains of asparagine and glutamine appear as doublets on the top right corner, and a smaller peak may appear on top of each peak due to deuterium exchange from the D2O normally added to an NMR sample, giving these sidechain peaks a distinctive appearance. The sidechain amine peaks from tryptophan are usually shifted downfield and appear near the bottom left corner. The backbone amide peaks of glycine normally appear near the top of the spectrum.
The 15N HSQC is normally the first heteronuclear spectrum acquired for the assignment of resonances where each amide peak is assigned to a particular residue in the protein. If the protein is folded, the peaks are usually well-dispersed, and most of the individual peaks can be distinguished. If there is a large cluster of severely overlapped peaks around the middle of the spectrum, that would indicate the presence of significant unstructured elements in the protein. In such cases where there are severe overlap of resonances the assignment of resonances in the spectra can be difficult. The assignment of the HSQC spectrum requires other experiments, ideally using triple resonance experiments with 15N and 13C-labelled proteins, that provide sequential connectivities between residues so that the resonances can be linked to particular residues and sequentially assigned. The assignment of the spectrum is essential for a meaningful interpretation of more advanced NMR experiments such as structure determination and relaxation analysis.
Chemicals labelled with 15N isotope are relatively inexpensive, and the 15N HSQC is a sensitive experiment whereby a spectrum can be acquired in a relatively short time, the 15N HSQC is therefore often used to screen candidates for their suitability for structure determination by NMR, as well as optimization of the sample conditions. The time-consuming process of structure determination is usually not undertaken until a good HSQC spectrum can be obtained. The HSQC experiment is also useful for detecting binding interface in protein-protein interaction, as well the interactions with ligands such as drugs. By comparing the HSQC of the free protein with the one bound to the ligand, changes in the chemical shifts of some peaks may be observed, and these peaks are likely to lie on the binding surface where the binding perturbed their chemical shifts. [4] The 15N HSQC may also be used in relaxation analysis in the studies of molecular dynamics of proteins, the determination of ionization constant, and other studies.
1H—13C HSQC
This experiment provides correlations between a carbon and its attached protons. The constant time (CT) version of 1H—13C HSQC is normally used as it circumvents the issue of splitting of signal due to homonuclear 13C—13C J couplings which reduces spectral resolution. [5] The "constant time" refers to the entire evolution period between the two INEPT steps which is kept constant in this experiment. If this evolution period is set to be the inverse of the J-coupling constant, then the sign of the magnetization of those carbons with an odd number of aliphatic carbon attached will be opposite to those with an even number. For example, if the Cβ of leucine appears as a positive peak (2 aliphatic carbons attached), then the Cγ (3 aliphatic carbons attached) and Cα (1 aliphatic carbons attached) would appear negative.