Related Research Articles

In genetics, complementary DNA (cDNA) is DNA synthesized from a single-stranded RNA template in a reaction catalyzed by the enzyme reverse transcriptase. cDNA is often used to express a specific protein in a cell that does not normally express that protein, or to sequence or quantify mRNA molecules using DNA based methods. cDNA that codes for a specific protein can be transferred to a recipient cell for expression, often bacterial or yeast expression systems. cDNA is also generated to analyze transcriptomic profiles in bulk tissue, single cells, or single nuclei in assays such as microarrays, qPCR, and RNA-seq.

The polymerase chain reaction (PCR) is a method widely used to make millions to billions of copies of a specific DNA sample rapidly, allowing scientists to amplify a very small sample of DNA sufficiently to enable detailed study. PCR was invented in 1983 by American biochemist Kary Mullis at Cetus Corporation; Mullis and biochemist Michael Smith, who had developed other essential ways of manipulating DNA, were jointly awarded the Nobel Prize in Chemistry in 1993.

A primer is a short single-stranded nucleic acid used by all living organisms in the initiation of DNA synthesis. DNA polymerase enzymes are only capable of adding nucleotides to the 3’-end of an existing nucleic acid, requiring a primer be bound to the template before DNA polymerase can begin a complementary strand. DNA polymerase adds nucleotides after binding to the RNA primer and synthesizes the whole strand. Later, the RNA strands must be removed accurately and replace them with DNA nucleotides forming a gap region known as a nick that is filled in using an enzyme called ligase. The removal process of the RNA primer requires several enzymes, such as Fen1, Lig1, and others that work in coordination with DNA polymerase, to ensure the removal of the RNA nucleotides and the addition of DNA nucleotides. Living organisms use solely RNA primers, while laboratory techniques in biochemistry and molecular biology that require in vitro DNA synthesis usually use DNA primers, since they are more temperature stable. Primers can be designed in laboratory for specific reactions such as polymerase chain reaction (PCR). When designing PCR primers, there are specific measures that must be taken into consideration, like the melting temperature of the primers and the annealing temperature of the reaction itself. Moreover, the DNA binding sequence of the primer in vitro has to be specifically chosen, which is done using a method called basic local alignment search tool (BLAST) that scans the DNA and finds specific and unique regions for the primer to bind.

A retrovirus is a type of virus that inserts a DNA copy of its RNA genome into the DNA of a host cell that it invades, thus changing the genome of that cell. After invading a host cell's cytoplasm, the virus uses its own reverse transcriptase enzyme to produce DNA from its RNA genome, the reverse of the usual pattern, thus retro (backwards). The new DNA is then incorporated into the host cell genome by an integrase enzyme, at which point the retroviral DNA is referred to as a provirus. The host cell then treats the viral DNA as part of its own genome, transcribing and translating the viral genes along with the cell's own genes, producing the proteins required to assemble new copies of the virus. Many retroviruses cause serious diseases in humans, other mammals, and birds.

A reverse transcriptase (RT) is an enzyme used to generate complementary DNA (cDNA) from an RNA template, a process termed reverse transcription. Reverse transcriptases are used by viruses such as HIV and hepatitis B to replicate their genomes, by retrotransposon mobile genetic elements to proliferate within the host genome, and by eukaryotic cells to extend the telomeres at the ends of their linear chromosomes. Contrary to a widely held belief, the process does not violate the flows of genetic information as described by the classical central dogma, as transfers of information from RNA to DNA are explicitly held possible.
DNA primase is an enzyme involved in the replication of DNA and is a type of RNA polymerase. Primase catalyzes the synthesis of a short RNA segment called a primer complementary to a ssDNA template. After this elongation, the RNA piece is removed by a 5' to 3' exonuclease and refilled with DNA.

Ribonuclease H is a family of non-sequence-specific endonuclease enzymes that catalyze the cleavage of RNA in an RNA/DNA substrate via a hydrolytic mechanism. Members of the RNase H family can be found in nearly all organisms, from bacteria to archaea to eukaryotes.
Helicase-dependent amplification (HDA) is a method for in vitro DNA amplification that takes place at a constant temperature.
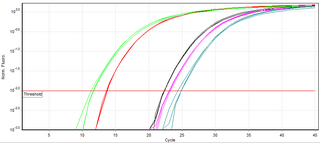
A real-time polymerase chain reaction is a laboratory technique of molecular biology based on the polymerase chain reaction (PCR). It monitors the amplification of a targeted DNA molecule during the PCR, not at its end, as in conventional PCR. Real-time PCR can be used quantitatively and semi-quantitatively.
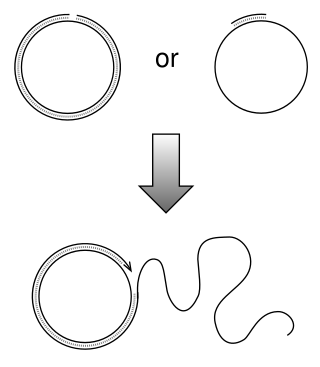

Rolling circle replication (RCR) is a process of unidirectional nucleic acid replication that can rapidly synthesize multiple copies of circular molecules of DNA or RNA, such as plasmids, the genomes of bacteriophages, and the circular RNA genome of viroids. Some eukaryotic viruses also replicate their DNA or RNA via the rolling circle mechanism.
In molecular biology and genetics, the sense of a nucleic acid molecule, particularly of a strand of DNA or RNA, refers to the nature of the roles of the strand and its complement in specifying a sequence of amino acids. Depending on the context, sense may have slightly different meanings. For example, negative-sense strand of DNA is equivalent to the template strand, whereas the positive-sense strand is the non-template strand whose nucleotide sequence is equivalent to the sequence of the mRNA transcript.

Multicopy single-stranded DNA (msDNA) is a type of extrachromosomal satellite DNA that consists of a single-stranded DNA molecule covalently linked via a 2'-5'phosphodiester bond to an internal guanosine of an RNA molecule. The resultant DNA/RNA chimera possesses two stem-loops joined by a branch similar to the branches found in RNA splicing intermediates. The coding region for msDNA, called a "retron", also encodes a type of reverse transcriptase, which is essential for msDNA synthesis.

T7 RNA Polymerase is an RNA polymerase from the T7 bacteriophage that catalyzes the formation of RNA from DNA in the 5'→ 3' direction.
Loop-mediated isothermal amplification (LAMP) is a single-tube technique for the amplification of DNA and a low-cost alternative to detect certain diseases. Reverse transcription loop-mediated isothermal amplification (RT-LAMP) combines LAMP with a reverse transcription step to allow the detection of RNA.
The versatility of polymerase chain reaction (PCR) has led to modifications of the basic protocol being used in a large number of variant techniques designed for various purposes. This article summarizes many of the most common variations currently or formerly used in molecular biology laboratories; familiarity with the fundamental premise by which PCR works and corresponding terms and concepts is necessary for understanding these variant techniques.

T7 DNA polymerase is an enzyme used during the DNA replication of the T7 bacteriophage. During this process, the DNA polymerase “reads” existing DNA strands and creates two new strands that match the existing ones. The T7 DNA polymerase requires a host factor, E. coli thioredoxin, in order to carry out its function. This helps stabilize the binding of the necessary protein to the primer-template to improve processivity by more than 100-fold, which is a feature unique to this enzyme. It is a member of the Family A DNA polymerases, which include E. coli DNA polymerase I and Taq DNA polymerase.

Hepatitis B virus DNA polymerase is a hepatitis B viral protein. It is a DNA polymerase that can use either DNA or RNA templates and a ribonuclease H that cuts RNA in the duplex. Both functions are supplied by the reverse transcriptase (RT) domain.
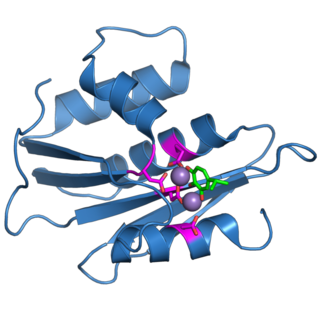
The retroviral ribonuclease H is a catalytic domain of the retroviral reverse transcriptase (RT) enzyme. The RT enzyme is used to generate complementary DNA (cDNA) from the retroviral RNA genome. This process is called reverse transcription. To complete this complex process, the retroviral RT enzymes need to adopt a multifunctional nature. They therefore possess 3 of the following biochemical activities: RNA-dependent DNA polymerase, ribonuclease H, and DNA-dependent DNA polymerase activities. Like all RNase H enzymes, the retroviral RNase H domain cleaves DNA/RNA duplexes and will not degrade DNA or unhybridized RNA.
Recombinase polymerase amplification (RPA) is a single tube, isothermal alternative to the polymerase chain reaction (PCR). By adding a reverse transcriptase enzyme to an RPA reaction it can detect RNA as well as DNA, without the need for a separate step to produce cDNA,. Because it is isothermal, RPA can use much simpler equipment than PCR, which requires a thermal cycler. Operating best at temperatures of 37–42 °C and still working, albeit more slowly, at room temperature means RPA reactions can in theory be run quickly simply by holding a tube. This makes RPA an excellent candidate for developing low-cost, rapid, point-of-care molecular tests. An international quality assessment of molecular detection of Rift Valley fever virus performed as well as the best RT-PCR tests, detecting less concentrated samples missed by some PCR tests and an RT-LAMP test. RPA was developed and launched by TwistDx Ltd., a biotechnology company based in Cambridge, UK.

Reverse transcription loop-mediated isothermal amplification (RT-LAMP) is a one step nucleic acid amplification method to multiply specific sequences of RNA. It is used to diagnose infectious disease caused by RNA viruses.
References
- ↑ Deiman, Birgit; van Aarle, Pierre; Sillekens, Peter (2002). "Characteristics and Applications of Nucleic Acid Sequence-Based Amplification (NASBA)". Molecular Biotechnology. 20 (2): 163–180. doi:10.1385/mb:20:2:163. ISSN 1073-6085. PMID 11876473. S2CID 28712952.
- ↑ Reed, Adam J.; Connelly, Ryan P.; Williams, Allison; Tran, Maithi; Shim, Byoung-Shik; Choe, Hyeryun; Gerasimova, Yulia V. (March 2019). "Label-free pathogen detection by a deoxyribozyme cascade with visual signal readout". Sensors and Actuators B: Chemical. 282: 945–951. doi:10.1016/j.snb.2018.11.147. ISSN 0925-4005. PMC 6713451 . PMID 31462856.
- ↑ Lamb, Laura E.; Bartolone, Sarah N.; Tree, Maya O.; Conway, Michael J.; Rossignol, Julien; Smith, Christopher P.; Chancellor, Michael B. (December 2018). "Rapid Detection of Zika Virus in Urine Samples and Infected Mosquitos by Reverse Transcription-Loop-Mediated Isothermal Amplification". Scientific Reports. 8 (1): 3803. Bibcode:2018NatSR...8.3803L. doi:10.1038/s41598-018-22102-5. ISSN 2045-2322. PMC 5830622 . PMID 29491389.
- ↑ Schachter, Julius (1997), "Evaluation of Diagnostic Tests — Special Problems Introduced by DNA Amplification Procedures", Nucleic Acid Amplification Technologies Application to Disease Diagnosis, Boston, MA: Birkhäuser Boston, pp. 165–169, doi:10.1007/978-1-4612-2454-9_12, ISBN 978-1-4612-7543-5 , retrieved 2020-11-15
- ↑ Biolabs, New England. "Isothermal Amplification | NEB". www.neb.com. Retrieved 2020-11-15.
- ↑ Biolabs, New England. "Isothermal Amplification | NEB". www.neb.com. Retrieved 2020-11-15.
- ↑ Biolabs, New England. "Isothermal Amplification | NEB". www.neb.com. Retrieved 2020-11-15.
- ↑ Malek, L.; Sooknanan, R.; Compton, J. (1994). Nucleic acid sequence-based amplification (NASBA). Methods in Molecular Biology. Vol. 28. pp. 253–260. doi:10.1385/0-89603-254-x:253. ISSN 1064-3745. PMID 7509695. S2CID 30720773.
- ↑ Malek, L.; Sooknanan, R.; Compton, J. (1994). Nucleic acid sequence-based amplification (NASBA). Methods in Molecular Biology. Vol. 28. pp. 253–260. doi:10.1385/0-89603-254-x:253. ISSN 1064-3745. PMID 7509695. S2CID 30720773.
- ↑ Vasileva Wand, Nadina I.; Bonney, Laura C.; Watson, Robert J.; Graham, Victoria; Hewson, Roger (August 2018). "Point-of-care diagnostic assay for the detection of Zika virus using the recombinase polymerase amplification method". The Journal of General Virology. 99 (8): 1012–1026. doi:10.1099/jgv.0.001083. ISSN 1465-2099. PMC 6171711 . PMID 29897329.
- ↑ "PDB101: Molecule of the Month: RNA Polymerase". RCSB: PDB-101. Retrieved 2020-11-15.
- ↑ Compton, J (1991). "Nucleic acid sequence-based amplification". Nature. 350 (6313): 91–2. Bibcode:1991Natur.350...91C. doi:10.1038/350091a0. PMID 1706072. S2CID 4304204.
- ↑ Kievits, T; Van Gemen, B; Van Strijp, D; Schukkink, R; Dircks, M; Adriaanse, H; Malek, L; Sooknanan, R; Lens, P (1991). "NASBA isothermal enzymatic in vitro nucleic acid amplification optimized for the diagnosis of HIV-1 infection". Journal of Virological Methods. 35 (3): 273–86. doi:10.1016/0166-0934(91)90069-c. PMID 1726172.
- ↑ Schneider, P; Wolters, L; Schoone, G; Schallig, H; Sillekens, P; Hermsen, R; Sauerwein, R (2005). "Real-time nucleic acid sequence-based amplification is more convenient than real-time PCR for quantification of Plasmodium falciparum". Journal of Clinical Microbiology. 43 (1): 402–5. doi:10.1128/JCM.43.1.402-405.2005. PMC 540116 . PMID 15635001.
- ↑ Arnaud-Barbe, Nadege; Cheynet Sauvion, Valerie; Oriol, Guy; Mandrand, Bernard; Mallet, Francois (1998). "Transcription of RNA templates by T7 RNA polymerase". Nucleic Acids Research. 26 (15): 3550–3554. doi:10.1093/nar/26.15.3550. PMC 147742 . PMID 9671817.
- ↑ Collins, RA; Ko, LS; So, KL; Ellis, T; Lau, LT; Yu, AC (2002). "Detection of highly pathogenic and low pathogenic avian influenza subtype H5 (Eurasian lineage) using NASBA". Journal of Virological Methods. 103 (2): 213–25. doi:10.1016/S0166-0934(02)00034-4. PMID 12008015.
- ↑ Collins, RA; Ko, LS; Fung, KY; Lau, LT; Xing, J; Yu, AC (2002). "A method to detect major serotypes of foot-and-mouth disease virus". Biochemical and Biophysical Research Communications. 297 (2): 267–74. CiteSeerX 10.1.1.328.625 . doi:10.1016/S0006-291X(02)02178-2. PMID 12237113.
- ↑ Keightley, MC; Sillekens, P; Schippers, W; Rinaldo, C; George, KS (2005). "Real-time NASBA detection of SARS-associated coronavirus and comparison with real-time reverse transcription-PCR". Journal of Medical Virology. 77 (4): 602–8. doi: 10.1002/jmv.20498 . PMC 7167117 . PMID 16254971.
- ↑ Böhmer, A; Schildgen, V; Lüsebrink, J; Ziegler, S; Tillmann, RL; Kleines, M; Schildgen, O (2009). "Novel application for isothermal nucleic acid sequence-based amplification (NASBA)". Journal of Virological Methods. 158 (1–2): 199–201. doi:10.1016/j.jviromet.2009.02.010. PMID 19428591.
- ↑ Mugasa, CM; Laurent, T; Schoone, GJ; Kager, PA; Lubega, GW; Schallig, HD (2009). "Nucleic acid sequence-based amplification with oligochromatography for detection of Trypanosoma brucei in clinical samples". Journal of Clinical Microbiology. 47 (3): 630–5. doi:10.1128/JCM.01430-08. PMC 2650916 . PMID 19116352.
- ↑ Wu, Qianxin; Suo, Chenqu; Brown, Tom; Wang, Tengyao; Teichmann, Sarah A.; Bassett, Andrew R. (February 2021). "INSIGHT: A population-scale COVID-19 testing strategy combining point-of-care diagnosis with centralized high-throughput sequencing". Science Advances. 7 (7): eabe5054. doi:10.1126/sciadv.abe5054. ISSN 2375-2548. PMC 7880595 . PMID 33579697.