
Catechol-O-methyltransferase is one of several enzymes that degrade catecholamines, catecholestrogens, and various drugs and substances having a catechol structure. In humans, catechol-O-methyltransferase protein is encoded by the COMT gene. Two isoforms of COMT are produced: the soluble short form (S-COMT) and the membrane bound long form (MB-COMT). As the regulation of catecholamines is impaired in a number of medical conditions, several pharmaceutical drugs target COMT to alter its activity and therefore the availability of catecholamines. COMT was first discovered by the biochemist Julius Axelrod in 1957.

Methoxsalen sold under the brand name Oxsoralen among others, is a medication used to treat psoriasis, eczema, vitiligo, and some cutaneous lymphomas in conjunction with exposing the skin to ultraviolet (UVA) light from lamps or sunlight. Methoxsalen modifies the way skin cells receive the UVA radiation, allegedly clearing up the disease. Levels of individual patient PUVA exposure were originally determined using the Fitzpatrick scale. The scale was developed after patients demonstrated symptoms of phototoxicity after oral ingestion of methoxsalen followed by PUVA therapy. Chemically, methoxsalen is a derivative of psoralen and belongs to a class of organic natural molecules known as furanocoumarins. They consist of coumarin annulated with furan. It can also be injected and used topically.

Thiopurine methyltransferase or thiopurine S-methyltransferase (TPMT) is an enzyme that in humans is encoded by the TPMT gene. A pseudogene for this locus is located on chromosome 18q.

Methyltransferases are a large group of enzymes that all methylate their substrates but can be split into several subclasses based on their structural features. The most common class of methyltransferases is class I, all of which contain a Rossmann fold for binding S-Adenosyl methionine (SAM). Class II methyltransferases contain a SET domain, which are exemplified by SET domain histone methyltransferases, and class III methyltransferases, which are membrane associated. Methyltransferases can also be grouped as different types utilizing different substrates in methyl transfer reactions. These types include protein methyltransferases, DNA/RNA methyltransferases, natural product methyltransferases, and non-SAM dependent methyltransferases. SAM is the classical methyl donor for methyltransferases, however, examples of other methyl donors are seen in nature. The general mechanism for methyl transfer is a SN2-like nucleophilic attack where the methionine sulfur serves as the leaving group and the methyl group attached to it acts as the electrophile that transfers the methyl group to the enzyme substrate. SAM is converted to S-Adenosyl homocysteine (SAH) during this process. The breaking of the SAM-methyl bond and the formation of the substrate-methyl bond happen nearly simultaneously. These enzymatic reactions are found in many pathways and are implicated in genetic diseases, cancer, and metabolic diseases. Another type of methyl transfer is the radical S-Adenosyl methionine (SAM) which is the methylation of unactivated carbon atoms in primary metabolites, proteins, lipids, and RNA.

Phenylethanolamine N-methyltransferase (PNMT) is an enzyme found primarily in the adrenal medulla that converts norepinephrine (noradrenaline) to epinephrine (adrenaline). It is also expressed in small groups of neurons in the human brain and in selected populations of cardiomyocytes.

N-Acetylserotonin O-methyltransferase, also known as ASMT, is an enzyme which catalyzes the final reaction in melatonin biosynthesis: converting Normelatonin to melatonin. This reaction is embedded in the more general tryptophan metabolism pathway. The enzyme also catalyzes a second reaction in tryptophan metabolism: the conversion of 5-hydroxy-indoleacetate to 5-methoxy-indoleacetate. The other enzyme which catalyzes this reaction is n-acetylserotonin-o-methyltransferase-like-protein.
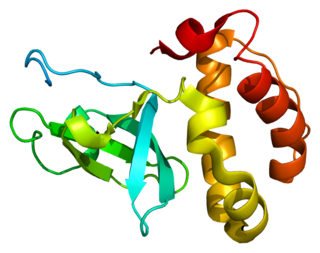
DNA (cytosine-5)-methyltransferase 3 beta, is an enzyme that in humans in encoded by the DNMT3B gene. Mutation in this gene are associated with immunodeficiency, centromere instability and facial anomalies syndrome.
In enzymology, a sterigmatocystin 8-O-methyltransferase is an enzyme that catalyzes the chemical reaction

Methylated-DNA--protein-cysteine methyltransferase(MGMT), also known as O6-alkylguanine DNA alkyltransferaseAGT, is a protein that in humans is encoded by the MGMT gene. MGMT is crucial for genome stability. It repairs the naturally occurring mutagenic DNA lesion O6-methylguanine back to guanine and prevents mismatch and errors during DNA replication and transcription. Accordingly, loss of MGMT increases the carcinogenic risk in mice after exposure to alkylating agents. The two bacterial isozymes are Ada and Ogt.

Histone-lysine N-methyltransferase 2D (KMT2D), also known as MLL4 and sometimes MLL2 in humans and Mll4 in mice, is a major mammalian histone H3 lysine 4 (H3K4) mono-methyltransferase. It is part of a family of six Set1-like H3K4 methyltransferases that also contains KMT2A, KMT2B, KMT2C, KMT2F, and KMT2G.
The O-methylated flavonoids or methoxyflavonoids are flavonoids with methylations on hydroxyl groups. O-methylation has an effect on the solubility of flavonoids.
An O-methyltransferase (OMT) is a type of methyltransferase enzyme transferring a methyl group on a molecule.

Syringetin is an O-methylated flavonol, a type of flavonoid. It is found in red grape, in Lysimachia congestiflora and in Vaccinium uliginosum. It is one of the phenolic compounds present in wine.
5-hydroxyfuranocoumarin 5-O-methyltransferase is an enzyme with systematic name S-adenosyl-L-methionine:5-hydroxyfurocoumarin 5-O-methyltransferase. This enzyme catalyses the following chemical reaction
8-hydroxyfuranocoumarin 8-O-methyltransferase is an enzyme with systematic name S-adenosyl-L-methionine:8-hydroxyfurocoumarin 8-O-methyltransferase. This enzyme catalyses the following chemical reaction
S-adenosyl-L-methionine:xanthotoxol O-methyltransferase may refer to:
23S rRNA (adenosine1067-2'-O)-methyltransferase is an enzyme with systematic name S-adenosyl-L-methionine:23S rRNA (adenosine1067-2'-O)-methyltransferase. This enzyme catalyses the following chemical reaction
Mycinamicin VI 2''-O-methyltransferase is an enzyme with systematic name S-adenosyl-L-methionine:mycinamicin VI 2''-O-methyltransferase. This enzyme catalyses the following chemical reaction:

3-Deazaneplanocin A is a drug which acts as both a S-adenosylhomocysteine synthesis inhibitor and also a histone methyltransferase EZH2 inhibitor. Studies have shown that it has effects in vitro against a variety of different tumor cell lines.











