
In immunology, an antigen (Ag) is a molecule, moiety, foreign particulate matter, or an allergen, such as pollen, that can bind to a specific antibody or T-cell receptor. The presence of antigens in the body may trigger an immune response.

Proteomics is the large-scale study of proteins. Proteins are vital parts of living organisms, with many functions such as the formation of structural fibers of muscle tissue, enzymatic digestion of food, or synthesis and replication of DNA. In addition, other kinds of proteins include antibodies that protect an organism from infection, and hormones that send important signals throughout the body.

The enzyme-linked immunosorbent assay (ELISA) is a commonly used analytical biochemistry assay, first described by Eva Engvall and Peter Perlmann in 1971. The assay uses a solid-phase type of enzyme immunoassay (EIA) to detect the presence of a ligand in a liquid sample using antibodies directed against the protein to be measured. ELISA has been used as a diagnostic tool in medicine, plant pathology, and biotechnology, as well as a quality control check in various industries.

A microarray is a multiplex lab-on-a-chip. Its purpose is to simultaneously detect the expression of thousands of biological interactions. It is a two-dimensional array on a solid substrate—usually a glass slide or silicon thin-film cell—that assays (tests) large amounts of biological material using high-throughput screening miniaturized, multiplexed and parallel processing and detection methods. The concept and methodology of microarrays was first introduced and illustrated in antibody microarrays by Tse Wen Chang in 1983 in a scientific publication and a series of patents. The "gene chip" industry started to grow significantly after the 1995 Science Magazine article by the Ron Davis and Pat Brown labs at Stanford University. With the establishment of companies, such as Affymetrix, Agilent, Applied Microarrays, Arrayjet, Illumina, and others, the technology of DNA microarrays has become the most sophisticated and the most widely used, while the use of protein, peptide and carbohydrate microarrays is expanding.

The western blot, or western blotting, is a widely used analytical technique in molecular biology and immunogenetics to detect specific proteins in a sample of tissue homogenate or extract. Besides detecting the proteins, this technique is also utilized to visualize, distinguish, and quantify the different proteins in a complicated protein combination.
An epitope, also known as antigenic determinant, is the part of an antigen that is recognized by the immune system, specifically by antibodies, B cells, or T cells. The part of an antibody that binds to the epitope is called a paratope. Although epitopes are usually non-self proteins, sequences derived from the host that can be recognized are also epitopes.
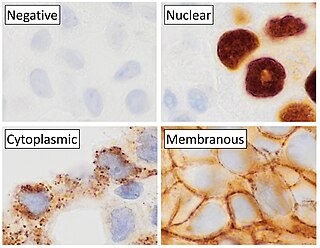
Immunohistochemistry (IHC) is the most common application of immunostaining. It involves the process of selectively identifying antigens (proteins) in cells of a tissue section by exploiting the principle of antibodies binding specifically to antigens in biological tissues. IHC takes its name from the roots "immuno", in reference to antibodies used in the procedure, and "histo", meaning tissue. Albert Coons conceptualized and first implemented the procedure in 1941.
Serology is the scientific study of serum and other body fluids. In practice, the term usually refers to the diagnostic identification of antibodies in the serum. Such antibodies are typically formed in response to an infection, against other foreign proteins, or to one's own proteins. In either case, the procedure is simple.
A protein microarray is a high-throughput method used to track the interactions and activities of proteins, and to determine their function, and determining function on a large scale. Its main advantage lies in the fact that large numbers of proteins can be tracked in parallel. The chip consists of a support surface such as a glass slide, nitrocellulose membrane, bead, or microtitre plate, to which an array of capture proteins is bound. Probe molecules, typically labeled with a fluorescent dye, are added to the array. Any reaction between the probe and the immobilised protein emits a fluorescent signal that is read by a laser scanner. Protein microarrays are rapid, automated, economical, and highly sensitive, consuming small quantities of samples and reagents. The concept and methodology of protein microarrays was first introduced and illustrated in antibody microarrays in 1983 in a scientific publication and a series of patents. The high-throughput technology behind the protein microarray was relatively easy to develop since it is based on the technology developed for DNA microarrays, which have become the most widely used microarrays.

Peptide mass fingerprinting (PMF), also known as protein fingerprinting, is an analytical technique for protein identification in which the unknown protein of interest is first cleaved into smaller peptides, whose absolute masses can be accurately measured with a mass spectrometer such as MALDI-TOF or ESI-TOF. The method was developed in 1993 by several groups independently. The peptide masses are compared to either a database containing known protein sequences or even the genome. This is achieved by using computer programs that translate the known genome of the organism into proteins, then theoretically cut the proteins into peptides, and calculate the absolute masses of the peptides from each protein. They then compare the masses of the peptides of the unknown protein to the theoretical peptide masses of each protein encoded in the genome. The results are statistically analyzed to find the best match.

In immunology, epitope mapping is the process of experimentally identifying the binding site, or epitope, of an antibody on its target antigen. Identification and characterization of antibody binding sites aid in the discovery and development of new therapeutics, vaccines, and diagnostics. Epitope characterization can also help elucidate the binding mechanism of an antibody and can strengthen intellectual property (patent) protection. Experimental epitope mapping data can be incorporated into robust algorithms to facilitate in silico prediction of B-cell epitopes based on sequence and/or structural data.

Immunoelectrophoresis is a general name for a number of biochemical methods for separation and characterization of proteins based on electrophoresis and reaction with antibodies. All variants of immunoelectrophoresis require immunoglobulins, also known as antibodies, reacting with the proteins to be separated or characterized. The methods were developed and used extensively during the second half of the 20th century. In somewhat chronological order: Immunoelectrophoretic analysis, crossed immunoelectrophoresis, rocket-immunoelectrophoresis, fused rocket immunoelectrophoresis ad modum Svendsen and Harboe, affinity immunoelectrophoresis ad modum Bøg-Hansen.
Protein methods are the techniques used to study proteins. There are experimental methods for studying proteins. Computational methods typically use computer programs to analyze proteins. However, many experimental methods require computational analysis of the raw data.
In academia, computational immunology is a field of science that encompasses high-throughput genomic and bioinformatics approaches to immunology. The field's main aim is to convert immunological data into computational problems, solve these problems using mathematical and computational approaches and then convert these results into immunologically meaningful interpretations.

An antibody microarray is a specific form of protein microarray. In this technology, a collection of captured antibodies are spotted and fixed on a solid surface such as glass, plastic, membrane, or silicon chip, and the interaction between the antibody and its target antigen is detected. Antibody microarrays are often used for detecting protein expression from various biofluids including serum, plasma and cell or tissue lysates. Antibody arrays may be used for both basic research and medical and diagnostic applications.

Protein mass spectrometry refers to the application of mass spectrometry to the study of proteins. Mass spectrometry is an important method for the accurate mass determination and characterization of proteins, and a variety of methods and instrumentations have been developed for its many uses. Its applications include the identification of proteins and their post-translational modifications, the elucidation of protein complexes, their subunits and functional interactions, as well as the global measurement of proteins in proteomics. It can also be used to localize proteins to the various organelles, and determine the interactions between different proteins as well as with membrane lipids.
Immunosignaturing is a medical diagnostic test which uses arrays of random-sequence peptides to associate antibodies in a blood sample with a disease.
Immunomics is the study of immune system regulation and response to pathogens using genome-wide approaches. With the rise of genomic and proteomic technologies, scientists have been able to visualize biological networks and infer interrelationships between genes and/or proteins; recently, these technologies have been used to help better understand how the immune system functions and how it is regulated. Two thirds of the genome is active in one or more immune cell types and less than 1% of genes are uniquely expressed in a given type of cell. Therefore, it is critical that the expression patterns of these immune cell types be deciphered in the context of a network, and not as an individual, so that their roles be correctly characterized and related to one another. Defects of the immune system such as autoimmune diseases, immunodeficiency, and malignancies can benefit from genomic insights on pathological processes. For example, analyzing the systematic variation of gene expression can relate these patterns with specific diseases and gene networks important for immune functions.
Secretomics is a type of proteomics which involves the analysis of the secretome—all the secreted proteins of a cell, tissue or organism. Secreted proteins are involved in a variety of physiological processes, including cell signaling and matrix remodeling, but are also integral to invasion and metastasis of malignant cells. Secretomics has thus been especially important in the discovery of biomarkers for cancer and understanding molecular basis of pathogenesis. The analysis of the insoluble fraction of the secretome has been termed matrisomics.

A peptide microarray is a collection of peptides displayed on a solid surface, usually a glass or plastic chip. Peptide chips are used by scientists in biology, medicine and pharmacology to study binding properties and functionality and kinetics of protein-protein interactions in general. In basic research, peptide microarrays are often used to profile an enzyme, to map an antibody epitope or to find key residues for protein binding. Practical applications are seromarker discovery, profiling of changing humoral immune responses of individual patients during disease progression, monitoring of therapeutic interventions, patient stratification and development of diagnostic tools and vaccines.














