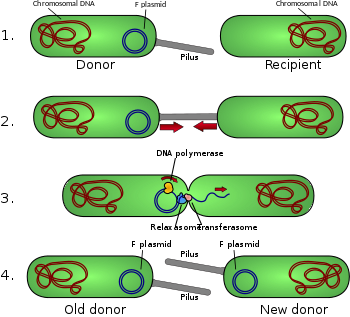
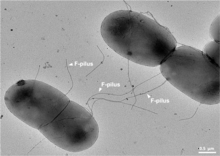
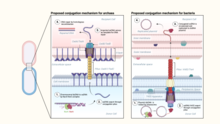
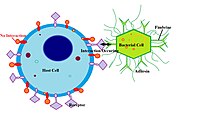
Bacterial conjugation is the transfer of genetic material between bacterial cells by direct cell-to-cell contact or by a bridge-like connection between two cells. This takes place through a pilus. It is a parasexual mode of reproduction in bacteria.

Neisseria gonorrhoeae, also known as gonococcus (singular) or gonococci (plural), is a species of Gram-negative diplococci bacteria isolated by Albert Neisser in 1879. It causes the sexually transmitted genitourinary infection gonorrhea as well as other forms of gonococcal disease including disseminated gonococcemia, septic arthritis, and gonococcal ophthalmia neonatorum.
The evolution of flagella is of great interest to biologists because the three known varieties of flagella – each represent a sophisticated cellular structure that requires the interaction of many different systems.

Neisseria is a large genus of bacteria that colonize the mucosal surfaces of many animals. Of the 11 species that colonize humans, only two are pathogens, N. meningitidis and N. gonorrhoeae.
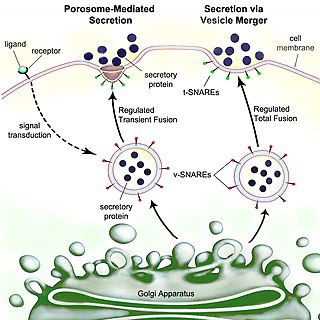
Secretion is the movement of material from one point to another, such as a secreted chemical substance from a cell or gland. In contrast, excretion is the removal of certain substances or waste products from a cell or organism. The classical mechanism of cell secretion is via secretory portals at the plasma membrane called porosomes. Porosomes are permanent cup-shaped lipoprotein structures embedded in the cell membrane, where secretory vesicles transiently dock and fuse to release intra-vesicular contents from the cell.

Agrobacterium tumefaciens is the causal agent of crown gall disease in over 140 species of eudicots. It is a rod-shaped, Gram-negative soil bacterium. Symptoms are caused by the insertion of a small segment of DNA, from a plasmid into the plant cell, which is incorporated at a semi-random location into the plant genome. Plant genomes can be engineered by use of Agrobacterium for the delivery of sequences hosted in T-DNA binary vectors.

An appendage is an external body part, or natural prolongation, that protrudes from an organism's or microorganism's body.
Adhesins are cell-surface components or appendages of bacteria that facilitate adhesion or adherence to other cells or to surfaces, usually in the host they are infecting or living in. Adhesins are a type of virulence factor.
The F-plasmid allows genes to be transferred from one bacterium carrying the factor to another bacterium lacking the factor by conjugation. The F factor was the first plasmid to be discovered. Unlike other plasmids, F factor is constitutive for transfer proteins due to a mutation in the gene finO. The F plasmid belongs to F-like plasmids, a class of conjugative plasmids that control sexual functions of bacteria with a fertility inhibition (Fin) system.
Pilin refers to a class of fibrous proteins that are found in pilus structures in bacteria. These structures can be used for the exchange of genetic material, or as a cell adhesion mechanism. Although not all bacteria have pili or fimbriae, bacterial pathogens often use their fimbriae to attach to host cells. In Gram-negative bacteria, where pili are more common, individual pilin molecules are linked by noncovalent protein-protein interactions, while Gram-positive bacteria often have polymerized LPXTG pilin.
Transfer genes or tra genes, are some genes necessary for non-sexual transfer of genetic material in both gram-positive and gram-negative bacteria. The tra locus includes the pilin gene and regulatory genes, which together form pili on the cell surface, polymeric proteins that can attach themselves to the surface of F-bacteria and initiate the conjugation. The existence of the tra region of a plasmid genome was first discovered in 1979 by David H. Figurski and Donald R. Helinski In the course of their work, Figurski and Helinski also discovered a second key fact about the tra region – that it can act in trans to the mobilization marker which it affects.
In biology, phase variation is a method for dealing with rapidly varying environments without requiring random mutation. It involves the variation of protein expression, frequently in an on-off fashion, within different parts of a bacterial population. As such the phenotype can switch at frequencies that are much higher than classical mutation rates. Phase variation contributes to virulence by generating heterogeneity. Although it has been most commonly studied in the context of immune evasion, it is observed in many other areas as well and is employed by various types of bacteria, including Salmonella species.

Bacterial motility is the ability of bacteria to move independently using metabolic energy. Most motility mechanisms that evolved among bacteria also evolved in parallel among the archaea. Most rod-shaped bacteria can move using their own power, which allows colonization of new environments and discovery of new resources for survival. Bacterial movement depends not only on the characteristics of the medium, but also on the use of different appendages to propel. Swarming and swimming movements are both powered by rotating flagella. Whereas swarming is a multicellular 2D movement over a surface and requires the presence of surfactants, swimming is movement of individual cells in liquid environments.
Sortase refers to a group of prokaryotic enzymes that modify surface proteins by recognizing and cleaving a carboxyl-terminal sorting signal. For most substrates of sortase enzymes, the recognition signal consists of the motif LPXTG (Leu-Pro-any-Thr-Gly), then a highly hydrophobic transmembrane sequence, followed by a cluster of basic residues such as arginine. Cleavage occurs between the Thr and Gly, with transient attachment through the Thr residue to the active site Cys residue, followed by transpeptidation that attaches the protein covalently to cell wall components. Sortases occur in almost all Gram-positive bacteria and the occasional Gram-negative bacterium or Archaea, where cell wall LPXTG-mediated decoration has not been reported. Although sortase A, the "housekeeping" sortase, typically acts on many protein targets, other forms of sortase recognize variant forms of the cleavage motif, or catalyze the assembly of pilins into pili.
Plasmid-mediated resistance is the transfer of antibiotic resistance genes which are carried on plasmids. Plasmids possess mechanisms that ensure their independent replication as well as those that regulate their replication number and guarantee stable inheritance during cell division. By the conjugation process, they can stimulate lateral transfer between bacteria from various genera and kingdoms. Numerous plasmids contain addiction-inducing systems that are typically based on toxin-antitoxin factors and capable of killing daughter cells that don't inherit the plasmid during cell division. Plasmids often carry multiple antibiotic resistance genes, contributing to the spread of multidrug-resistance (MDR). Antibiotic resistance mediated by MDR plasmids severely limits the treatment options for the infections caused by Gram-negative bacteria, especially family Enterobacteriaceae. The global spread of MDR plasmids has been enhanced by selective pressure from antimicrobial medications used in medical facilities and when raising animals for food.
Gabriel Waksman FMedSci, FRS, is Courtauld professor of biochemistry and molecular biology at University College London (UCL), and professor of structural and molecular biology at Birkbeck College, University of London. He is the director of the Institute of Structural and Molecular Biology (ISMB) at UCL and Birkbeck, head of the Department of Structural and Molecular Biology at UCL, and head of the Department of Biological Sciences at Birkbeck.

Twitching motility is a form of crawling bacterial motility used to move over surfaces. Twitching is mediated by the activity of hair-like filaments called type IV pili which extend from the cell's exterior, bind to surrounding solid substrates, and retract, pulling the cell forwards in a manner similar to the action of a grappling hook. The name twitching motility is derived from the characteristic jerky and irregular motions of individual cells when viewed under the microscope. It has been observed in many bacterial species, but is most well studied in Pseudomonas aeruginosa, Neisseria gonorrhoeae and Myxococcus xanthus. Active movement mediated by the twitching system has been shown to be an important component of the pathogenic mechanisms of several species.
The Curli protein is a type of amyloid fiber produced by certain strains of enterobacteria. They are extracellular fibers located on bacteria such as E. coli and Salmonella spp. These fibers serve to promote cell community behavior through biofilm formation in the extracellular matrix. Amyloids are associated with several human neurodegenerative diseases such as Alzheimer's disease, Huntington's disease, Parkinson's disease, and prion diseases. The study of curli may help to understand human diseases thought to arise from improper amyloid fiber formation. The curli pili are generally assembled through the extracellular nucleation precipitation pathway.
P fimbriae are chaperone-usher type fimbrial appendages found on the surface of many Escherichia coli bacteria. The P fimbriae is considered to be one of the most important virulence factor in uropathogenic E. coli and plays an important role in upper urinary tract infections. P fimbriae mediate adherence to host cells, a key event in the pathogenesis of urinary tract infections.
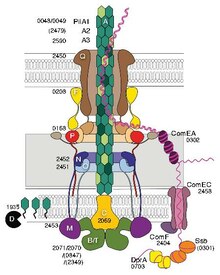
The bacterial type IV secretion system, also known as the type IV secretion system or the T4SS, is a secretion protein complex found in gram negative bacteria, gram positive bacteria, and archaea. It is able to transport proteins and DNA across the cell membrane. The type IV secretion system is just one of many bacterial secretion systems. Type IV secretion systems are related to conjugation machinery which generally involve a single-step secretion system and the use of a pilus. Type IV secretion systems are used for conjugation, DNA exchange with the extracellular space, and for delivering proteins to target cells. The type IV secretion system is divided into type IVA and type IVB based on genetic ancestry.