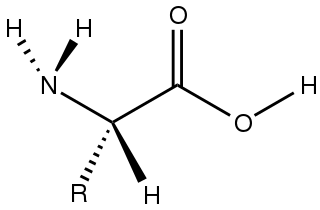
Amino acids are organic compounds that contain both amino and carboxylic acid functional groups. Although over 500 amino acids exist in nature, by far the most important are the 22 α-amino acids incorporated into proteins. Only these 22 appear in the genetic code of life.

Selenocysteine is the 21st proteinogenic amino acid. Selenoproteins contain selenocysteine residues. Selenocysteine is an analogue of the more common cysteine with selenium in place of the sulfur.
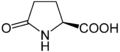
Proline (symbol Pro or P) is an organic acid classed as a proteinogenic amino acid (used in the biosynthesis of proteins), although it does not contain the amino group -NH
2 but is rather a secondary amine. The secondary amine nitrogen is in the protonated form (NH2+) under biological conditions, while the carboxyl group is in the deprotonated −COO− form. The "side chain" from the α carbon connects to the nitrogen forming a pyrrolidine loop, classifying it as a aliphatic amino acid. It is non-essential in humans, meaning the body can synthesize it from the non-essential amino acid L-glutamate. It is encoded by all the codons starting with CC (CCU, CCC, CCA, and CCG).

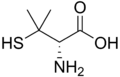
Cysteine is a semiessential proteinogenic amino acid with the formula HOOC−CH(−NH2)−CH2−SH. The thiol side chain in cysteine often participates in enzymatic reactions as a nucleophile. Cysteine is chiral, but interestingly, both D and L-cysteine are found in nature with D-cysteine having been found in developing brain. Cysteine is named after its discovery in urine, which comes from the urinary bladder or cyst, from kystis "bladder".

Methionine is an essential amino acid in humans.

Pyrrolysine is an α-amino acid that is used in the biosynthesis of proteins in some methanogenic archaea and bacteria; it is not present in humans. It contains an α-amino group, a carboxylic acid group. Its pyrroline side-chain is similar to that of lysine in being basic and positively charged at neutral pH.

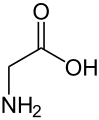
Alanine (symbol Ala or A), or α-alanine, is an α-amino acid that is used in the biosynthesis of proteins. It contains an amine group and a carboxylic acid group, both attached to the central carbon atom which also carries a methyl group side chain. Consequently it is classified as a nonpolar, aliphatic α-amino acid. Under biological conditions, it exists in its zwitterionic form with its amine group protonated (as −NH3+) and its carboxyl group deprotonated (as −CO2−). It is non-essential to humans as it can be synthesized metabolically and does not need to be present in the diet. It is encoded by all codons starting with GC (GCU, GCC, GCA, and GCG).
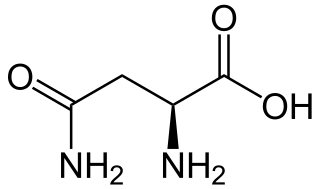
Asparagine is an α-amino acid that is used in the biosynthesis of proteins. It contains an α-amino group, an α-carboxylic acid group, and a side chain carboxamide, classifying it as a polar, aliphatic amino acid. It is non-essential in humans, meaning the body can synthesize it. It is encoded by the codons AAU and AAC.

Proteinogenic amino acids are amino acids that are incorporated biosynthetically into proteins during translation. The word "proteinogenic" means "protein creating". Throughout known life, there are 22 genetically encoded (proteinogenic) amino acids, 20 in the standard genetic code and an additional 2 that can be incorporated by special translation mechanisms.

Dehydroalanine is a dehydroamino acid. It does not exist in its free form, but it occurs naturally as a residue found in peptides of microbial origin. As an amino acid residue, it is unusual because it has an unsaturated backbone.
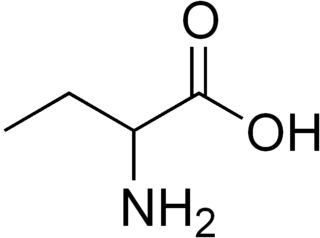
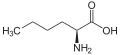
α-Aminobutyric acid (AABA), also known as homoalanine in biochemistry, is a non-proteinogenic alpha amino acid with chemical formula C4H9NO2. The straight two carbon side chain is one carbon longer than alanine, hence the prefix homo-.

Pyridoxal phosphate (PLP, pyridoxal 5'-phosphate, P5P), the active form of vitamin B6, is a coenzyme in a variety of enzymatic reactions. The International Union of Biochemistry and Molecular Biology has catalogued more than 140 PLP-dependent activities, corresponding to ~4% of all classified activities. The versatility of PLP arises from its ability to covalently bind the substrate, and then to act as an electrophilic catalyst, thereby stabilizing different types of carbanionic reaction intermediates.

Beta-peptides (β-peptides) are peptides derived from β-amino acids, in which the amino group is attached to the β-carbon (i.e. the carbon two atoms away from the carboxylate group). The parent β-amino acid is β-alanine (H2NCH2CH2CO2H), a common natural substance, but most examples feature substituents in place of one or more C-H bonds. β-peptides usually do not occur in nature. β-peptide-based antibiotics are being explored as ways of evading antibiotic resistance. Early studies in this field were published in 1996 by the group of Dieter Seebach and that of Samuel Gellman.

A catalytic triad is a set of three coordinated amino acids that can be found in the active site of some enzymes. Catalytic triads are most commonly found in hydrolase and transferase enzymes. An acid-base-nucleophile triad is a common motif for generating a nucleophilic residue for covalent catalysis. The residues form a charge-relay network to polarise and activate the nucleophile, which attacks the substrate, forming a covalent intermediate which is then hydrolysed to release the product and regenerate free enzyme. The nucleophile is most commonly a serine or cysteine amino acid, but occasionally threonine or even selenocysteine. The 3D structure of the enzyme brings together the triad residues in a precise orientation, even though they may be far apart in the sequence.

Amino acid biosynthesis is the set of biochemical processes by which the amino acids are produced. The substrates for these processes are various compounds in the organism's diet or growth media. Not all organisms are able to synthesize all amino acids. For example, humans can synthesize 11 of the 20 standard amino acids. These 11 are called the non-essential amino acids).

Cystathionine beta-lyase, also commonly referred to as CBL or β-cystathionase, is an enzyme that primarily catalyzes the following α,β-elimination reaction
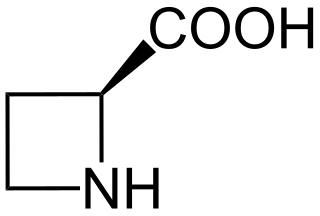
Azetidine-2-carboxylic acid (abbreviated Aze or Azc) is a plant non-protein amino acid homologue of proline with the molecular formula C4H7NO2. Aze is a heterocyclic, 4 membered ring with nitrogen as its heteroatom (an azetidine), and a carboxylic acid group substituted on one of the ring carbon atoms. The main difference between Aze and proline is the ring of Aze has four members and the ring of proline has five. Aze has the ability to act as an analog of proline and can be incorporated into proteins in place of proline.

In biochemistry, a dehydroamino acid or α,β-dehydroamino acid is an amino acids, usually with a C=C double bond in its side chain. Dehydroamino acids are not coded by DNA, but arise via post-translational modification.
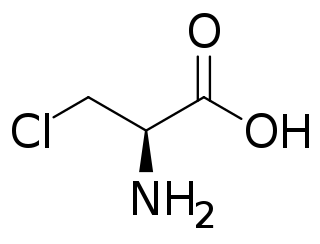
Chloroalanine (3-chloroalanine) is an unnatural amino acid with the formula ClCH2CH(NH2)CO2H. It is a white, water-soluble solid. The compound is usually derived from chlorination of serine. The compound is used in the synthesis of other amino acids by replacement of the chloride. Protected forms of the related iodoalanine are also known.
In organic chemistry, secondary amino acids are amino acids which do not contain the amino group −NH2 but is rather a secondary amine. Secondary amino acids can be classified to cyclic acids, such as proline, and acyclic N-substituted amino acids.