Related Research Articles

Polymerase chain reaction (PCR) is a method widely used to rapidly make millions to billions of copies of a specific DNA sample, allowing scientists to take a very small sample of DNA and amplify it to a large enough amount to study in detail. PCR was invented in 1983 by the American biochemist Kary Mullis at Cetus Corporation. It is fundamental to many of the procedures used in genetic testing and research, including analysis of ancient samples of DNA and identification of infectious agents. Using PCR, copies of very small amounts of DNA sequences are exponentially amplified in a series of cycles of temperature changes. PCR is now a common and often indispensable technique used in medical laboratory research for a broad variety of applications including biomedical research and criminal forensics.
Protein engineering is the process of developing useful or valuable proteins. It is a young discipline, with much research taking place into the understanding of protein folding and recognition for protein design principles. It is also a product and services market, with an estimated value of $168 billion by 2017.
Site-directed mutagenesis is a molecular biology method that is used to make specific and intentional changes to the DNA sequence of a gene and any gene products. Also called site-specific mutagenesis or oligonucleotide-directed mutagenesis, it is used for investigating the structure and biological activity of DNA, RNA, and protein molecules, and for protein engineering.
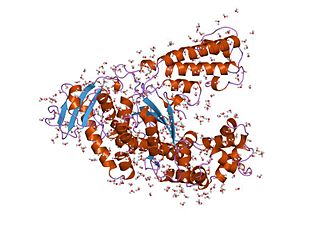
Taq polymerase is a thermostable DNA polymerase I named after the thermophilic eubacterial microorganism Thermus aquaticus, from which it was originally isolated by Chien et al. in 1976. Its name is often abbreviated to Taq or Taq pol. It is frequently used in the polymerase chain reaction (PCR), a method for greatly amplifying the quantity of short segments of DNA.
Pfu DNA polymerase is an enzyme found in the hyperthermophilic archaeon Pyrococcus furiosus, where it functions to copy the organism's DNA during cell division. In the laboratory setting, Pfu is used to amplify DNA in the polymerase chain reaction (PCR), where the enzyme serves the central function of copying a new strand of DNA during each extension step.

Pyrococcus furiosus is an extremophilic species of Archaea. It can be classified as a hyperthermophile because it thrives best under extremely high temperatures—higher than those preferred of a thermophile. It is notable for having an optimum growth temperature of 100 °C, and for being one of the few organisms identified as possessing aldehyde ferredoxin oxidoreductase enzymes containing tungsten, an element rarely found in biological molecules.

Thermostability is the quality of a substance to resist irreversible change in its chemical or physical structure, often by resisting decomposition or polymerization, at a high relative temperature.

Bisulfitesequencing (also known as bisulphite sequencing) is the use of bisulfite treatment of DNA before routine sequencing to determine the pattern of methylation. DNA methylation was the first discovered epigenetic mark, and remains the most studied. In animals it predominantly involves the addition of a methyl group to the carbon-5 position of cytosine residues of the dinucleotide CpG, and is implicated in repression of transcriptional activity.
The polymerase chain reaction (PCR) is a commonly used molecular biology tool for amplifying DNA, and various techniques for PCR optimization which have been developed by molecular biologists to improve PCR performance and minimize failure.

Functional cloning is a molecular cloning technique that relies on prior knowledge of the encoded protein’s sequence or function for gene identification. In this assay, a genomic or cDNA library is screened to identify the genetic sequence of a protein of interest. Expression cDNA libraries may be screened with antibodies specific for the protein of interest or may rely on selection via the protein function. Historically, the amino acid sequence of a protein was used to prepare degenerate oligonucleotides which were then probed against the library to identify the gene encoding the protein of interest. Once candidate clones carrying the gene of interest are identified, they are sequenced and their identity is confirmed. This method of cloning allows researchers to screen entire genomes without prior knowledge of the location of the gene or the genetic sequence.
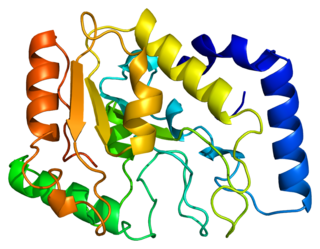
Uracil-DNA glycosylase, also known as UNG or UDG. Its most important function is to prevent mutagenesis by eliminating uracil from DNA molecules by cleaving the N-glycosidic bond and initiating the base-excision repair (BER) pathway.

The history of the polymerase chain reaction (PCR) has variously been described as a classic "Eureka!" moment, or as an example of cooperative teamwork between disparate researchers. Following is a list of events before, during, and after its development:
The versatility of polymerase chain reaction (PCR) has led to a large number of variants of PCR.
TOPO cloning is a molecular biology technique in which DNA fragments are cloned into specific vectors without the requirement for DNA ligases. Taq polymerase has a nontemplate-dependent terminal transferase activity that adds a single deoxyadenosine (A) to the 3'-end of the PCR products. This characteristic is exploited in "sticky end" TOPO TA cloning. For "blunt end" TOPO cloning, the recipient vector does not have overhangs and blunt-ended DNA fragments can be cloned.
The ligase chain reaction (LCR) is a method of DNA amplification. The ligase chain reaction (LCR) is an amplification process that differs from PCR in that it involves a thermostable ligase to join two probes or other molecules together which can then be amplified by standard polymerase chain reaction (PCR) cycling. Each cycle results in a doubling of the target nucleic acid molecule. A key advantage of LCR is greater specificity as compared to PCR. Thus, LCR requires two completely different enzymes to operate properly: ligase, to join probe molecules together, and a thermostable polymerase to amplify those molecules involved in successful ligation. The probes involved in the ligation are designed such that the 5′ end of one probe is directly adjacent to the 3′ end of the other probe, thereby providing the requisite 3′-OH and 5′-PO4 group substrates for the ligase.

T7 DNA polymerase is an enzyme used during the DNA replication of the T7 bacteriophage. During this process, the DNA polymerase “reads” existing DNA strands and creates two new strands that match the existing ones. The T7 DNA polymerase requires a host factor, E. coli thioredoxin, in order to carry out its function. This helps stabilize the binding of the necessary protein to the primer-template to improve processivity by more than 100-fold, which is a feature unique to this enzyme. It is a member of the Family A DNA polymerases, which include E. coli DNA polymerase I and Taq DNA polymerase.
TA cloning is a subcloning technique that avoids the use of restriction enzymes and is easier and quicker than traditional subcloning. The technique relies on the ability of adenine (A) and thymine (T) on different DNA fragments to hybridize and, in the presence of ligase, become ligated together. PCR products are usually amplified using Taq DNA polymerase which preferentially adds an adenine to the 3' end of the product. Such PCR amplified inserts are cloned into linearized vectors that have complementary 3' thymine overhangs.
Gibson assembly is a molecular cloning method which allows for the joining of multiple DNA fragments in a single, isothermal reaction. It is named after its creator, Daniel G. Gibson, who was the chief technology officer and co-founder of the synthetic biology company Codex DNA.
Φ29 DNA polymerase is an enzyme from the bacteriophage Φ29. It is being increasingly used in molecular biology for multiple displacement DNA amplification procedures, and has a number of features that make it particularly suitable for this application. It was discovered and characterised by Spanish scientist Margarita Salas.
Sequence saturation mutagenesis (SeSaM) is a chemo-enzymatic random mutagenesis method applied for the directed evolution of proteins and enzymes. It is one of the most common saturation mutagenesis techniques. In four PCR-based reaction steps, phosphorothioate nucleotides are inserted in the gene sequence, cleaved and the resulting fragments elongated by universal or degenerate nucleotides. These nucleotides are then replaced by standard nucleotides, allowing for a broad distribution of nucleic acid mutations spread over the gene sequence with a preference to transversions and with a unique focus on consecutive point mutations, both difficult to generate by other mutagenesis techniques. The technique was developed by Professor Ulrich Schwaneberg at Jacobs University Bremen and RWTH Aachen University.
References
- ↑ Ghasemi A, Salmanian AH, Sadeghifard N, Salarian AA, Gholi MK (September 2011). "Cloning, expression and purification of Pwo polymerase from Pyrococcus woesei". Iranian Journal of Microbiology. 3 (3): 118–22. PMC 3279813 . PMID 22347593.
- ↑ Lasken RS, Schuster DM, Rashtchian A (July 1996). "Archaebacterial DNA polymerases tightly bind uracil-containing DNA". The Journal of Biological Chemistry. 271 (30): 17692–6. doi: 10.1074/jbc.271.30.17692 . PMID 8663453.
- ↑ Rashtchian A, Thornton CG, Heidecker G (November 1992). "A novel method for site-directed mutagenesis using PCR and uracil DNA glycosylase". PCR Methods and Applications. 2 (2): 124–30. doi: 10.1101/gr.2.2.124 . PMID 1477668.