
In molecular biology, DNA replication is the biological process of producing two identical replicas of DNA from one original DNA molecule. DNA replication occurs in all living organisms acting as the most essential part of biological inheritance. This is essential for cell division during growth and repair of damaged tissues, while it also ensures that each of the new cells receives its own copy of the DNA. The cell possesses the distinctive property of division, which makes replication of DNA essential.

A reverse transcriptase (RT) is an enzyme used to convert RNA genome to DNA, a process termed reverse transcription. Reverse transcriptases are used by viruses such as HIV and hepatitis B to replicate their genomes, by retrotransposon mobile genetic elements to proliferate within the host genome, and by eukaryotic cells to extend the telomeres at the ends of their linear chromosomes. Contrary to a widely held belief, the process does not violate the flows of genetic information as described by the classical central dogma, as transfers of information from RNA to DNA are explicitly held possible.
A telomere is a region of repetitive nucleotide sequences associated with specialized proteins at the ends of linear chromosomes. Telomeres are a widespread genetic feature most commonly found in eukaryotes. In most, if not all species possessing them, they protect the terminal regions of chromosomal DNA from progressive degradation and ensure the integrity of linear chromosomes by preventing DNA repair systems from mistaking the very ends of the DNA strand for a double-strand break.

Transcription is the process of copying a segment of DNA into RNA. Some segments of DNA are transcribed into RNA molecules that can encode proteins, called messenger RNA (mRNA). Other segments of DNA are transcribed into RNA molecules called non-coding RNAs (ncRNAs).

A DNA polymerase is a member of a family of enzymes that catalyze the synthesis of DNA molecules from nucleoside triphosphates, the molecular precursors of DNA. These enzymes are essential for DNA replication and usually work in groups to create two identical DNA duplexes from a single original DNA duplex. During this process, DNA polymerase "reads" the existing DNA strands to create two new strands that match the existing ones. These enzymes catalyze the chemical reaction
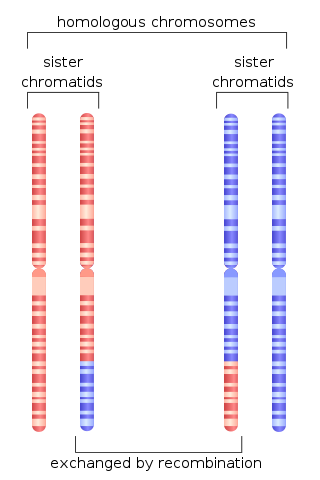
Homologous recombination is a type of genetic recombination in which genetic information is exchanged between two similar or identical molecules of double-stranded or single-stranded nucleic acids.
Subtelomeres are segments of DNA between telomeric caps and chromatin.

Telomeric repeat-binding factor 1 is a protein that in humans is encoded by the TERF1 gene.
In molecular biology, a displacement loop or D-loop is a DNA structure where the two strands of a double-stranded DNA molecule are separated for a stretch and held apart by a third strand of DNA. An R-loop is similar to a D-loop, but in that case the third strand is RNA rather than DNA. The third strand has a base sequence which is complementary to one of the main strands and pairs with it, thus displacing the other complementary main strand in the region. Within that region the structure is thus a form of triple-stranded DNA. A diagram in the paper introducing the term illustrated the D-loop with a shape resembling a capital "D", where the displaced strand formed the loop of the "D".
The MRN complex is a protein complex consisting of Mre11, Rad50 and Nbs1. In eukaryotes, the MRN/X complex plays an important role in the initial processing of double-strand DNA breaks prior to repair by homologous recombination or non-homologous end joining. The MRN complex binds avidly to double-strand breaks both in vitro and in vivo and may serve to tether broken ends prior to repair by non-homologous end joining or to initiate DNA end resection prior to repair by homologous recombination. The MRN complex also participates in activating the checkpoint kinase ATM in response to DNA damage. Production of short single-strand oligonucleotides by Mre11 endonuclease activity has been implicated in ATM activation by the MRN complex.

Fanconi anemia, complementation group M, also known as FANCM is a human gene. It is an emerging target in cancer therapy, in particular cancers with specific genetic deficiencies.
Microhomology-mediated end joining (MMEJ), also known as alternative nonhomologous end-joining (Alt-NHEJ) is one of the pathways for repairing double-strand breaks in DNA. As reviewed by McVey and Lee, the foremost distinguishing property of MMEJ is the use of microhomologous sequences during the alignment of broken ends before joining, thereby resulting in deletions flanking the original break. MMEJ is frequently associated with chromosome abnormalities such as deletions, translocations, inversions and other complex rearrangements.
Telomere-binding proteins function to bind telomeric DNA in various species. In particular, telomere-binding protein refers to TTAGGG repeat binding factor-1 (TERF1) and TTAGGG repeat binding factor-2 (TERF2). Telomere sequences in humans are composed of TTAGGG sequences which provide protection and replication of chromosome ends to prevent degradation. Telomere-binding proteins can generate a T-loop to protect chromosome ends. TRFs are double-stranded proteins which are known to induce bending, looping, and pairing of DNA which aids in the formation of T-loops. They directly bind to TTAGGG repeat sequence in the DNA. There are also subtelomeric regions present for regulation. However, in humans, there are six subunits forming a complex known as shelterin.
Shelterin is a protein complex known to protect telomeres in many eukaryotes from DNA repair mechanisms, as well as to regulate telomerase activity. In mammals and other vertebrates, telomeric DNA consists of repeating double-stranded 5'-TTAGGG-3' (G-strand) sequences along with the 3'-AATCCC-5' (C-strand) complement, ending with a 50-400 nucleotide 3' (G-strand) overhang. Much of the final double-stranded portion of the telomere forms a T-loop (Telomere-loop) that is invaded by the 3' (G-strand) overhang to form a small D-loop (Displacement-loop).

Synthesis-dependent strand annealing (SDSA) is a major mechanism of homology-directed repair of DNA double-strand breaks (DSBs). Although many of the features of SDSA were first suggested in 1976, the double-Holliday junction model proposed in 1983 was favored by many researchers. In 1994, studies of double-strand gap repair in Drosophila were found to be incompatible with the double-Holliday junction model, leading researchers to propose a model they called synthesis-dependent strand annealing. Subsequent studies of meiotic recombination in S. cerevisiae found that non-crossover products appear earlier than double-Holliday junctions or crossover products, challenging the previous notion that both crossover and non-crossover products are produced by double-Holliday junctions and leading the authors to propose that non-crossover products are generated through SDSA.

SLX4 interacting protein is a protein that in humans is encoded by the SLX4IP gene.

Telomeric repeat–containing RNA (TERRA) is a long non-coding RNA transcribed from telomeres - repetitive nucleotide regions found on the ends of chromosomes that function to protect DNA from deterioration or fusion with neighboring chromosomes. TERRA has been shown to be ubiquitously expressed in almost all cell types containing linear chromosomes - including humans, mice, and yeasts. While the exact function of TERRA is still an active area of research, it is generally believed to play a role in regulating telomerase activity as well as maintaining the heterochromatic state at the ends of chromosomes. TERRA interaction with other associated telomeric proteins has also been shown to help regulate telomere integrity in a length-dependent manner.
Telomeres, the caps on the ends of eukaryotic chromosomes, play critical roles in cellular aging and cancer. An important facet to how telomeres function in these roles is their involvement in cell cycle regulation.

DNA end resection, also called 5′–3′ degradation, is a biochemical process where the blunt end of a section of double-stranded DNA (dsDNA) is modified by cutting away some nucleotides from the 5' end to produce a 3' single-stranded sequence. The presence of a section of single-stranded DNA (ssDNA) allows the broken end of the DNA to line up accurately with a matching sequence, so that it can be accurately repaired.

Jan Karlseder an Austrian molecular biologist, is the Chief Science Officer and a Senior Vice President at the Salk Institute for Biological Studies. He is also a professor in the Molecular and Cellular Biology Laboratory, the Director of the Paul F. Glenn Center for Biology of Aging Research and the holder of the Donald and Darlene Shiley Chair at the Salk Institute for Biological Studies.













