
RNA splicing is a process in molecular biology where a newly-made precursor messenger RNA (pre-mRNA) transcript is transformed into a mature messenger RNA (mRNA). It works by removing all the introns and splicing back together exons. For nuclear-encoded genes, splicing occurs in the nucleus either during or immediately after transcription. For those eukaryotic genes that contain introns, splicing is usually needed to create an mRNA molecule that can be translated into protein. For many eukaryotic introns, splicing occurs in a series of reactions which are catalyzed by the spliceosome, a complex of small nuclear ribonucleoproteins (snRNPs). There exist self-splicing introns, that is, ribozymes that can catalyze their own excision from their parent RNA molecule. The process of transcription, splicing and translation is called gene expression, the central dogma of molecular biology.

Alternative splicing, or alternative RNA splicing, or differential splicing, is an alternative splicing process during gene expression that allows a single gene to code for multiple proteins. In this process, particular exons of a gene may be included within or excluded from the final, processed messenger RNA (mRNA) produced from that gene. This means the exons are joined in different combinations, leading to different (alternative) mRNA strands. Consequently, the proteins translated from alternatively spliced mRNAs will contain differences in their amino acid sequence and, often, in their biological functions.
RNA-binding proteins are proteins that bind to the double or single stranded RNA in cells and participate in forming ribonucleoprotein complexes. RBPs contain various structural motifs, such as RNA recognition motif (RRM), dsRNA binding domain, zinc finger and others. They are cytoplasmic and nuclear proteins. However, since most mature RNA is exported from the nucleus relatively quickly, most RBPs in the nucleus exist as complexes of protein and pre-mRNA called heterogeneous ribonucleoprotein particles (hnRNPs). RBPs have crucial roles in various cellular processes such as: cellular function, transport and localization. They especially play a major role in post-transcriptional control of RNAs, such as: splicing, polyadenylation, mRNA stabilization, mRNA localization and translation. Eukaryotic cells express diverse RBPs with unique RNA-binding activity and protein–protein interaction. According to the Eukaryotic RBP Database (EuRBPDB), there are 2961 genes encoding RBPs in humans. During evolution, the diversity of RBPs greatly increased with the increase in the number of introns. Diversity enabled eukaryotic cells to utilize RNA exons in various arrangements, giving rise to a unique RNP (ribonucleoprotein) for each RNA. Although RBPs have a crucial role in post-transcriptional regulation in gene expression, relatively few RBPs have been studied systematically.It has now become clear that RNA–RBP interactions play important roles in many biological processes among organisms.
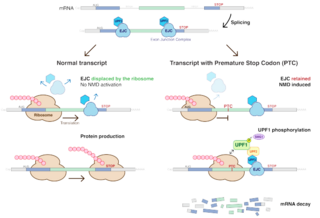
Nonsense-mediated mRNA decay (NMD) is a surveillance pathway that exists in all eukaryotes. Its main function is to reduce errors in gene expression by eliminating mRNA transcripts that contain premature stop codons. Translation of these aberrant mRNAs could, in some cases, lead to deleterious gain-of-function or dominant-negative activity of the resulting proteins.
IKBKAP is a human gene encoding the IKAP protein, which is ubiquitously expressed at varying levels in all tissue types, including brain cells. The IKAP protein is thought to participate as a sub-unit in the assembly of a six-protein putative human holo-Elongator complex, which allows for transcriptional elongation by RNA polymerase II. Further evidence has implicated the IKAP protein as being critical in neuronal development, and directs that decreased expression of IKAP in certain cell types is the molecular basis for the severe, neurodevelopmental disorder familial dysautonomia. Other pathways that have been connected to IKAP protein function in a variety of organisms include tRNA modification, cell motility, and cytosolic stress signalling. Homologs of the IKBKAP gene have been identified in multiple other Eukaryotic model organisms. Notable homologs include Elp1 in yeast, Ikbkap in mice, and D-elp1 in fruit flies. The fruit fly homolog (D-elp1) has RNA-dependent RNA polymerase activity and is involved in RNA interference.
Growth hormone-binding protein (GHBP) is a soluble carrier protein for growth hormone (GH). The full range of functions of GHBP remains to be determined however, current research suggests that the protein is associated with regulation of the GH availability and half-life in the circulatory system, as well as modulating GH receptor function.

Eukaryotic translation termination factor 1 (eRF1), also known as TB3-1, is a protein that in humans is encoded by the ETF1 gene.

RNA-binding protein 8A is a protein that in humans is encoded by the RBM8A gene.
Serine/arginine repetitive matrix protein 1 is a protein that in humans is encoded by the SRRM1 gene.

RNA-binding protein with serine-rich domain 1 is a protein that in humans is encoded by the RNPS1 gene.

Regulator of nonsense transcripts 2 is a protein that in humans is encoded by the UPF2 gene.

Regulator of nonsense transcripts 3B is a protein that in humans is encoded by the UPF3B gene.
Aly/REF export factor, also known as THO complex subunit 4 is a protein that in humans is encoded by the ALYREF gene.
Protein mago nashi homolog is a protein that in humans is encoded by the MAGOH gene.

Eukaryotic initiation factor 4A-III is a protein that in humans is encoded by the EIF4A3 gene.
Protein CASC3 is a protein that in humans is encoded by the CASC3 gene.

Regulator of nonsense transcripts 3A is a protein that in humans is encoded by the UPF3A gene.
mRNA surveillance mechanisms are pathways utilized by organisms to ensure fidelity and quality of messenger RNA (mRNA) molecules. There are a number of surveillance mechanisms present within cells. These mechanisms function at various steps of the mRNA biogenesis pathway to detect and degrade transcripts that have not properly been processed.
An exon junction complex (EJC) is a protein complex which forms on a pre-messenger RNA strand at the junction of two exons which have been joined together during RNA splicing. The EJC has major influences on translation, surveillance and localization of the spliced mRNA. It is first deposited onto mRNA during splicing and is then transported into the cytoplasm. There it plays a major role in post-transcriptional regulation of mRNA. It is believed that exon junction complexes provide a position-specific memory of the splicing event. The EJC consists of a stable heterotetramer core, which serves as a binding platform for other factors necessary for the mRNA pathway. The core of the EJC contains the protein eukaryotic initiation factor 4A-III bound to an adenosine triphosphate (ATP) analog, as well as the additional proteins Magoh and Y14. The binding of these proteins to nuclear speckled domains has been measured recently and it may be regulated by PI3K/AKT/mTOR signaling pathways. In order for the binding of the complex to the mRNA to occur, the eIF4AIII factor is inhibited, stopping the hydrolysis of ATP. This recognizes EJC as an ATP dependent complex. EJC also interacts with a large number of additional proteins; most notably SR proteins. These interactions are suggested to be important for mRNA compaction. The role of EJC in mRNA export is controversial.

Nucleolar protein with MIF4G domain 1 is a protein that in humans is encoded by the NOM1 gene.
















