Related Research Articles

Tryptophan repressor is a transcription factor involved in controlling amino acid metabolism. It has been best studied in Escherichia coli, where it is a dimeric protein that regulates transcription of the 5 genes in the tryptophan operon. When the amino acid tryptophan is plentiful in the cell, it binds to the protein, which causes a conformational change in the protein. The repressor complex then binds to its operator sequence in the genes it regulates, shutting off the genes.
Charles Yanofsky was an American geneticist on the faculty of Stanford University who contributed to the establishment of the one gene-one enzyme hypothesis and discovered attenuation, a riboswitch mechanism in which messenger RNA changes shape in response to a small molecule and thus alters its binding ability for the regulatory region of a gene or operon.
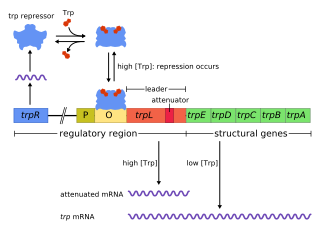
The trp operon is a group of genes that are transcribed together, encoding the enzymes that produce the amino acid tryptophan in bacteria. The trp operon was first characterized in Escherichia coli, and it has since been discovered in many other bacteria. The operon is regulated so that, when tryptophan is present in the environment, the genes for tryptophan synthesis are repressed.
The gene rpoS encodes the sigma factor sigma-38, a 37.8 kD protein in Escherichia coli. Sigma factors are proteins that regulate transcription in bacteria. Sigma factors can be activated in response to different environmental conditions. rpoS is transcribed in late exponential phase, and RpoS is the primary regulator of stationary phase genes. RpoS is a central regulator of the general stress response and operates in both a retroactive and a proactive manner: it not only allows the cell to survive environmental challenges, but it also prepares the cell for subsequent stresses (cross-protection). The transcriptional regulator CsgD is central to biofilm formation, controlling the expression of the curli structural and export proteins, and the diguanylate cyclase, adrA, which indirectly activates cellulose production. The rpoS gene most likely originated in the gammaproteobacteria.

DicF RNA is a non-coding RNA that is an antisense inhibitor of cell division gene ftsZ. DicF is bound by the Hfq protein which enhances its interaction with its targets. Pathogenic E. coli strains possess multiple copies of sRNA DicF in their genomes, while non-pathogenic strains do not. DicF and Hfq are both necessary to reduce FtsZ protein levels, leading to cell filamentation under anaerobic conditions.

DsrA RNA is a non-coding RNA that regulates both transcription, by overcoming transcriptional silencing by the nucleoid-associated H-NS protein, and translation, by promoting efficient translation of the stress sigma factor, RpoS. These two activities of DsrA can be separated by mutation: the first of three stem-loops of the 85 nucleotide RNA is necessary for RpoS translation but not for anti-H-NS action, while the second stem-loop is essential for antisilencing and less critical for RpoS translation. The third stem-loop, which behaves as a transcription terminator, can be substituted by the trp transcription terminator without loss of either DsrA function. The sequence of the first stem-loop of DsrA is complementary with the upstream leader portion of RpoS messenger RNA, suggesting that pairing of DsrA with the RpoS message might be important for translational regulation. The structures of DsrA and DsrA/rpoS complex were studied by NMR. The study concluded that the sRNA contains a dynamic conformational equilibrium for its second stem–loop which might be an important mechanism for DsrA to regulate the translations of its multiple target mRNAs.

Spot 42 (spf) RNA is a regulatory non-coding bacterial small RNA encoded by the spf gene. Spf is found in gammaproteobacteria and the majority of experimental work on Spot42 has been performed in Escherichia coli and recently in Aliivibrio salmonicida. In the cell Spot42 plays essential roles as a regulator in carbohydrate metabolism and uptake, and its expression is activated by glucose, and inhibited by the cAMP-CRP complex.

(p)ppGpp, guanosine pentaphosphate and tetraphosphate, also known as the "magic spot" nucleotides, are alarmones involved in the stringent response in bacteria that cause the inhibition of RNA synthesis when there is a shortage of amino acids. This inhibition by (p)ppGpp decreases translation in the cell, conserving amino acids present. Furthermore, ppGpp and pppGpp cause the up-regulation of many other genes involved in stress response such as the genes for amino acid uptake and biosynthesis.

The enzyme anthranilate synthase catalyzes the chemical reaction
The gal operon is a prokaryotic operon, which encodes enzymes necessary for galactose metabolism. Repression of gene expression for this operon works via binding of repressor molecules to two operators. These repressors dimerize, creating a loop in the DNA. The loop as well as hindrance from the external operator prevent RNA polymerase from binding to the promoter, and thus prevent transcription. Additionally, since the metabolism of galactose in the cell is involved in both anabolic and catabolic pathways, a novel regulatory system using two promoters for differential repression has been identified and characterized within the context of the gal operon.
Roberto Kolter is Professor of Microbiology, Emeritus at Harvard Medical School, an author, and past president of the American Society for Microbiology. Kolter has been a professor at Harvard Medical School since 1983 and was Co-director of Harvard's Microbial Sciences Initiative from 2003-2018. During the 35-year term of the Kolter laboratory from 1983 to 2018, more than 130 graduate student and postdoctoral trainees explored an eclectic mix of topics gravitating around the study of microbes. Kolter is a fellow of the American Association for the Advancement of Science and of the American Academy of Microbiology.
The fim switch in Escherichia coli is the mechanism by which the fim gene cluster, encoding Type I Pili, is transcriptionally controlled.
The HP1 Holin Family is a member of the Holin Superfamily II. Proteins in this family are typically found to contain two transmembrane segments (TMSs) and range between 70 and 80 amino acyl residues (aas) in length. A representative list of proteins belonging to the HP1 holin family can be found in the Transporter Classification Database.
The locus of enterocyte effacement-encoded regulator (Ler) is a regulatory protein that controls bacterial pathogenicity of enteropathogenic Escherichia coli (EPEC) and enterohemorrhagic Escherichia coli (EHEC). More specifically, Ler regulates the locus of enterocyte effacement (LEE) pathogenicity island genes, which are responsible for creating intestinal attachment and effacing lesions and subsequent diarrhea: LEE1, LEE2, and LEE3. LEE1, 2, and 3 carry the information necessary for a type III secretion system. The transcript encoding the Ler protein is the open reading frame 1 on the LEE1 operon.
CsgD is a transcription and response regulator protein referenced to as the master modulator of bacterial biofilm development. In E. coli cells, CsgD is tasked with aiding the transition from planktonic cell motility to the stationary phase of biofilm formation, in response to environmental growth factors. A transcription analysis assay illustrated a heightened decrease in CsgD's DNA-binding capacity when phosphorylated at A.A. D59 of the protein's primary sequence. Therefore, in the protein's active form (unphosphorylated), CsgD is capable of carrying out its normal functions of regulating curli proteins (fimbria) and producing ECM polysaccharides (cellulose). Following a promoter-lacZ fusion assay of CsgD binding to specific target sites on E. coli's genome, two classes of binding targets were identified: group I genes and group II genes. The group I genes, akin to fliE and yhbT, exhibit repressed transcription following their interaction with CsgD, whilst group II genes, including yccT and adrA, illustrated active functionality. Other group I operons that illustrate repressed transcription include fliE and fliEFGH, for motile flagellum formation. Other group II genes, imperative to the transition towards stationary biofilm development, include csgBA, encoding for curli fimbriae, and adrA, encoding for the synthesis of cyclic diguanylate. In this context, c-di-GMP functions as a bacterial secondary messenger, enhancing the production of extracellular cellulose and impeding flagellum production and rotation.

The Phosphate (Pho) regulon is a regulatory mechanism used for the conservation and management of inorganic phosphate within the cell. It was first discovered in Escherichia coli as an operating system for the bacterial strain, and was later identified in other species. The Pho system is composed of various components including extracellular enzymes and transporters that are capable of phosphate assimilation in addition to extracting inorganic phosphate from organic sources. This is an essential process since phosphate plays an important role in cellular membranes, genetic expression, and metabolism within the cell. Under low nutrient availability, the Pho regulon helps the cell survive and thrive despite a depletion of phosphate within the environment. When this occurs, phosphate starvation-inducible (psi) genes activate other proteins that aid in the transport of inorganic phosphate.

Manjula Reddy is an Indian bacterial geneticist. She is the chief scientist at the Centre for Cellular and Molecular Biology in Hyderabad, India. In 2019, she won the Infosys Prize in Life Sciences for her work on bacterial cell wall structure and synthesis. She is a Fellow of the Telangana Academy of Sciences and the Indian Academy of Sciences.
Terri Grodzicker is an American molecular geneticist and virologist who is currently the Dean of Academic Affairs at Cold Spring Harbor Laboratory. She also serves as the editor-in-chief of Genes & Development. Her research is in the field of human Adenoviridae. During her career at Cold Spring Harbor Laboratory she has edited more than a dozen books, and co-authored papers and books with other renown members of the laboratory such as Joseph Sambrook and Bruce Stillman.
Transcription-translation coupling is a mechanism of gene expression regulation in which synthesis of an mRNA (transcription) is affected by its concurrent decoding (translation). In prokaryotes, mRNAs are translated while they are transcribed. This allows communication between RNA polymerase, the multisubunit enzyme that catalyzes transcription, and the ribosome, which catalyzes translation. Coupling involves both direct physical interactions between RNA polymerase and the ribosome, as well as ribosome-induced changes to the structure and accessibility of the intervening mRNA that affect transcription.
Colanic acid is an exopolysaccharide synthesized by the bacteria Enterobacteriaceae. It is produced as a chemical defense to protect the cell surface, and it assists in the formation of biofilms.
References
- 1 2 3 4 5 "Catherine L. (Kearney) Squires". Winters Express. 2021-09-11. Retrieved 2021-10-29.
- ↑ Squires, Catherine Kearney (1967). Studies on a mutant of E. coli exhibiting a cold-sensitive phenotype for lactose fermentation (Thesis). Davis, Calif. OCLC 81257999.
- ↑ Squires, Catherine Louise (1972). Biochemical and genetic study of CRM in the L-arabinose operon (Thesis).
- 1 2 3 Henkin, Tina M. (2021-10-04). "Catherine L. Squires, 1941-2021: Scientist, Academic Leader, Mentor". Journal of Bacteriology. 203 (24): JB.00472–21. doi:10.1128/JB.00472-21. ISSN 0021-9193. PMC 8604069 . S2CID 238356903.
- ↑ Squires, Catherine K.; Ingraham, John L. (1969). "Mutant of Escherichia coli Exhibiting a Cold-sensitive Phenotype for Growth on Lactose". Journal of Bacteriology. 97 (2): 488–494. doi:10.1128/jb.97.2.488-494.1969. ISSN 0021-9193. PMC 249716 . PMID 4886277.
- ↑ Marr, Allen G.; Ingraham, John L.; Squires, Craig L. (1964). "EFFECT OF THE TEMPERATURE OF GROWTH OF ESCHERICHIA COLI ON THE FORMATION OF β-GALACTOSIDASE". Journal of Bacteriology. 87 (2): 356–362. doi:10.1128/jb.87.2.356-362.1964. ISSN 0021-9193. PMC 277016 . PMID 14151057.
- ↑ Squires, Catherine L.; Rose, John K.; Yanofsky, Charles; Yang, Huey-Lang; Zubay, Geoffrey (1973). "Tryptophanyl-tRNA and Tryptophanyl-tRNA Synthetase are not Required for in vitro Repression of the Tryptophan Operon". Nature New Biology. 245 (144): 131–133. doi:10.1038/newbio245131a0. ISSN 0090-0028. PMID 4582891.
- ↑ Squires, Catherine L.; Lee, Frank D.; Yanofsky, Charles (1975-02-15). "Interaction of the trp repressor and RNA polymerase with the trp operon". Journal of Molecular Biology. 92 (1): 93–111. doi:10.1016/0022-2836(75)90093-5. ISSN 0022-2836. PMID 1097702.
- ↑ Squires, C.; Krainer, A.; Barry, G.; Shen, W.-F.; Squires, C.L. (1981). "Nudeotide sequence at the end of the gene for the RNA polymerase β subunit (rpoC)". Nucleic Acids Research. 9 (24): 6827–6840. doi:10.1093/nar/9.24.6827. ISSN 0305-1048. PMC 327645 . PMID 6278450.
- ↑ Barry, G.; Squires, C.; Squires, C. L. (1980-06-01). "Attenuation and processing of RNA from the rplJL--rpoBC transcription unit of Escherichia coli". Proceedings of the National Academy of Sciences. 77 (6): 3331–3335. Bibcode:1980PNAS...77.3331B. doi: 10.1073/pnas.77.6.3331 . ISSN 0027-8424. PMC 349609 . PMID 6158044.
- ↑ Squires, C; Squires, C L (1992). "The Clp proteins: proteolysis regulators or molecular chaperones?". Journal of Bacteriology. 174 (4): 1081–1085. doi:10.1128/jb.174.4.1081-1085.1992. ISSN 0021-9193. PMC 206400 . PMID 1735703.
- ↑ Asai, Tsuneaki; Condon, Ciarán; Voulgaris, Justina; Zaporojets, Dmitry; Shen, Binghua; Al-Omar, Michaal; Squires, Craig; Squires, Catherine L. (1999). "Construction and Initial Characterization of Escherichia coli Strains with Few or No Intact Chromosomal rRNA Operons". Journal of Bacteriology. 181 (12): 3803–3809. doi:10.1128/JB.181.12.3803-3809.1999. ISSN 0021-9193. PMC 93859 . PMID 10368156.
- ↑ Asai, T.; Zaporojets, D.; Squires, C.; Squires, C. L. (1999-03-02). "An Escherichia coli strain with all chromosomal rRNA operons inactivated: Complete exchange of rRNA genes between bacteria". Proceedings of the National Academy of Sciences. 96 (5): 1971–1976. Bibcode:1999PNAS...96.1971A. doi: 10.1073/pnas.96.5.1971 . ISSN 0027-8424. PMC 26721 . PMID 10051579.
- ↑ Hilts, Philip J. (1992-04-09). "HEAD OF GENE MAP THREATENS TO QUIT". The New York Times. ISSN 0362-4331 . Retrieved 2021-10-29.
- ↑ "Columbia University Record". Vol. 19, no. 27. May 6, 1994. Archived from the original on 15 December 2011. Retrieved 2021-10-29.
{{cite magazine}}: Cite magazine requires|magazine=(help) - ↑ "Historic Fellows | American Association for the Advancement of Science". www.aaas.org. Retrieved 2021-10-29.
- ↑ "The 1990s". Tufts University School of Medicine. 2018-06-20. Archived from the original on 2018-10-24. Retrieved 2021-10-29.