The gene rpoS encodes the sigma factor sigma-38, a 37.8 kD protein in Escherichia coli. Sigma factors are proteins that regulate transcription in bacteria. Sigma factors can be activated in response to different environmental conditions. rpoS is transcribed in late exponential phase, and RpoS is the primary regulator of stationary phase genes. RpoS is a central regulator of the general stress response and operates in both a retroactive and a proactive manner: it not only allows the cell to survive environmental challenges, but it also prepares the cell for subsequent stresses (cross-protection). The transcriptional regulator CsgD is central to biofilm formation, controlling the expression of the curli structural and export proteins, and the diguanylate cyclase, adrA, which indirectly activates cellulose production. The rpoS gene most likely originated in the gammaproteobacteria.

DsrA RNA is a non-coding RNA that regulates both transcription, by overcoming transcriptional silencing by the nucleoid-associated H-NS protein, and translation, by promoting efficient translation of the stress sigma factor, RpoS. These two activities of DsrA can be separated by mutation: the first of three stem-loops of the 85 nucleotide RNA is necessary for RpoS translation but not for anti-H-NS action, while the second stem-loop is essential for antisilencing and less critical for RpoS translation. The third stem-loop, which behaves as a transcription terminator, can be substituted by the trp transcription terminator without loss of either DsrA function. The sequence of the first stem-loop of DsrA is complementary with the upstream leader portion of RpoS messenger RNA, suggesting that pairing of DsrA with the RpoS message might be important for translational regulation. The structures of DsrA and DsrA/rpoS complex were studied by NMR. The study concluded that the sRNA contains a dynamic conformational equilibrium for its second stem–loop which might be an important mechanism for DsrA to regulate the translations of its multiple target mRNAs.

GadY RNA is a non-coding RNA. The GadY gene is located on between and on the opposite strand to the GadX and GadW genes. GadY can form base pairs with the 3' UTR of its target mRNA gadX, this pairing is thought to confer increased stability to the transcript, allowing accumulation of gadX and therefore increased expression of downstream acid resistance genes. The GadY gene produces three overlapping transcripts that differ in length. The long form is 105 nucleotides in length and two processed versions are 59 and 90 nucleotides in length. It has been shown that all three forms of GadY bind to the Hfq protein.

The gcvB RNA gene encodes a small non-coding RNA involved in the regulation of a number of amino acid transport systems as well as amino acid biosynthetic genes. The GcvB gene is found in enteric bacteria such as Escherichia coli. GcvB regulates genes by acting as an antisense binding partner of the mRNAs for each regulated gene. This binding is dependent on binding to a protein called Hfq. Transcription of the GcvB RNA is activated by the adjacent GcvA gene and repressed by the GcvR gene. A deletion of GcvB RNA from Y. pestis changed colony shape as well as reducing growth. It has been shown by gene deletion that GcvB is a regulator of acid resistance in E. coli. GcvB enhances the ability of the bacterium to survive low pH by upregulating the levels of the alternate sigma factor RpoS. A polymeric form of GcvB has recently been identified. Interaction of GcvB with small RNA SroC triggers the degradation of GcvB by RNase E, lifting the GcvB-mediated mRNA repression of its target genes.

The OmrA-B RNA gene family is a pair of homologous OmpR-regulated small non-coding RNA that was discovered in E. coli during two large-scale screens. OmrA-B is highly abundant in stationary phase, but low levels could be detected in exponentially growing cells as well. RygB is adjacent to RygA a closely related RNA. These RNAs bind to the Hfq protein and regulate gene expression by antisense binding. They negatively regulate the expression of several genes encoding outer membrane proteins, including cirA, CsgD, fecA, fepA and ompT by binding in the vicinity of the Shine-Dalgarno sequence, suggesting the control of these targets is dependent on Hfq protein and RNase E. Taken together, these data suggest that OmrA-B participates in the regulation of outer membrane composition, responding to environmental conditions.
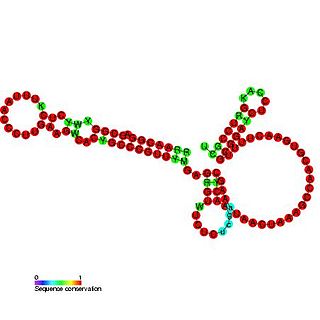
OxyS RNA is a small non-coding RNA which is induced in response to oxidative stress in Escherichia coli. This RNA acts as a global regulator to activate or repress the expression of as many as 40 genes, by an antisense mechanism, including the fhlA-encoded transcriptional activator and the rpoS-encoded sigma(s) subunit of RNA polymerase. OxyS is bound by the Hfq protein, that increases the OxyS RNA interaction with its target messages. Binding to Hfq alters the conformation of OxyS. The 109 nucleotide RNA is thought to be composed of three stem-loops.

The repression of heat shock gene expression (ROSE) element is an RNA element found in the 5' UTR of some heat shock protein's mRNAs. The ROSE element is an RNA thermometer that negatively regulates heat shock gene expression. The secondary structure is thought to be altered by temperature, thus it is an RNA thermometer. This structure blocks access to the ribosome binding site at normal temperatures. During heat shock however, the structure changes freeing the ribosome binding site and allowing expression to occur.

The Hfq protein encoded by the hfq gene was discovered in 1968 as an Escherichia coli host factor that was essential for replication of the bacteriophage Qβ. It is now clear that Hfq is an abundant bacterial RNA binding protein which has many important physiological roles that are usually mediated by interacting with Hfq binding sRNA.
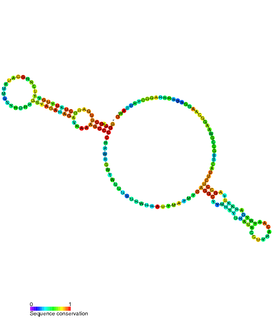
The hok/sok system is a postsegregational killing mechanism employed by the R1 plasmid in Escherichia coli. It was the first type I toxin-antitoxin pair to be identified through characterisation of a plasmid-stabilising locus. It is a type I system because the toxin is neutralised by a complementary RNA, rather than a partnered protein.
Cold shock response is a series of cardio-respiratory responses caused by sudden immersion in cold water.
Translational regulation refers to the control of the levels of protein synthesized from its mRNA. This regulation is vastly important to the cellular response to stressors, growth cues, and differentiation. In comparison to transcriptional regulation, it results in much more immediate cellular adjustment through direct regulation of protein concentration. The corresponding mechanisms are primarily targeted on the control of ribosome recruitment on the initiation codon, but can also involve modulation of peptide elongation, termination of protein synthesis, or ribosome biogenesis. While these general concepts are widely conserved, some of the finer details in this sort of regulation have been proven to differ between prokaryotic and eukaryotic organisms.

The JUMPstart RNA motif describes a conserved RNA-based secondary structure associated with JUMPstart elements. The 39-base-pair JUMPstart sequence describes a conserved element upstream of genes that participate in polysaccharide synthesis. The JUMPstart element has been shown to function as an RNA, and is present in the 5' untranslated regions of the genes it regulates.

FourU thermometers are a class of non-coding RNA thermometers found in Salmonella. They are named 'FourU' due to the four highly conserved uridine nucleotides found directly opposite the Shine-Dalgarno sequence on hairpin II (pictured). RNA thermometers such as FourU control regulation of temperature via heat shock proteins in many prokaryotes. FourU thermometers are relatively small RNA molecules, only 57 nucleotides in length, and have a simple two-hairpin structure.

MicX sRNA is a small non-coding RNA found in Vibrio cholerae. It was given the name MicX as it has a similar function to MicA, MicC and MicF in E. coli. MicX sRNA negatively regulates an outer membrane protein and also a component of an ABC transporter. These interactions were predicted and then confirmed using a DNA microarray.
Bacterial small RNAs (sRNA) are small RNAs produced by bacteria; they are 50- to 500-nucleotide non-coding RNA molecules, highly structured and containing several stem-loops. Numerous sRNAs have been identified using both computational analysis and laboratory-based techniques such as Northern blotting, microarrays and RNA-Seq in a number of bacterial species including Escherichia coli, the model pathogen Salmonella, the nitrogen-fixing alphaproteobacterium Sinorhizobium meliloti, marine cyanobacteria, Francisella tularensis, Streptococcus pyogenes, the pathogen Staphylococcus aureus, and the plant pathogen Xanthomonas oryzae pathovar oryzae. Bacterial sRNAs affect how genes are expressed within bacterial cells via interaction with mRNA or protein, and thus can affect a variety of bacterial functions like metabolism, virulence, environmental stress response, and structure.

An RNA thermometer is a temperature-sensitive non-coding RNA molecule which regulates gene expression. RNA thermometers often regulate genes required during either a heat shock or cold shock response, but have been implicated in other regulatory roles such as in pathogenicity and starvation.
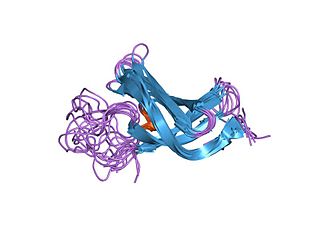
In molecular biology, the cold-shock domain (CSD) is a protein domain of about 70 amino acids which has been found in prokaryotic and eukaryotic DNA-binding proteins. Part of this domain is highly similar to the RNP-1 RNA-binding motif.
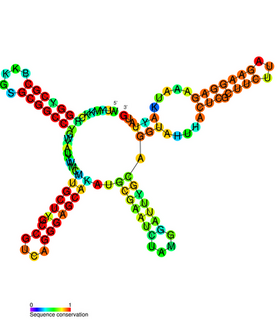
The IbpB thermometer is an RNA thermometer element found in the ibpAB operon. The operon contains two heat-shock genes, encoding inclusion body binding proteins A and B (IbpA/B), and is the most drastically upregulated operon under heat-shock in Escherichia coli.
The gene rpoE encodes the sigma factor sigma-24, a protein in Escherichia coli and other species of bacteria. Depending on the bacterial species, this gene may be referred to as sigE.
Bacterial small RNAs (sRNA) are an important class of regulatory molecules in bacteria such as Brucella. They are often bound to the chaperone protein Hfq, which allows them to interact with mRNA(s). In Brucella suis 1330 RNA sequencing identified a novel list of 33 sRNAs and 62 Hfq-associated mRNAs. In Brucella melitensis eight novel sRNA genes were identified using bioinformatic and experimental approach. One of them BSR0602 was found to modulate the intracellular survival of B. melitensis. In another large-scale deep sequencing study 1321 sRNAs were identified in B. melitensis. BSR0441 sRNA was further investigated in this study and shown to play role in the intracellular survival. sRNA BM-sr0117 from Brucella melitensis was identified and shown to be bound to and cleaved by Bm-RNase III. AbcR and AbcR2 were studied B. abortus. Seven novel sRNAs were validated and their interaction with a putative target sequence was verified in B. abortus.













