
A bacteriophage, also known informally as a phage, is a virus that infects and replicates within bacteria and archaea. The term was derived from "bacteria" and the Greek φαγεῖν, meaning "to devour". Bacteriophages are composed of proteins that encapsulate a DNA or RNA genome, and may have structures that are either simple or elaborate. Their genomes may encode as few as four genes and as many as hundreds of genes. Phages replicate within the bacterium following the injection of their genome into its cytoplasm.
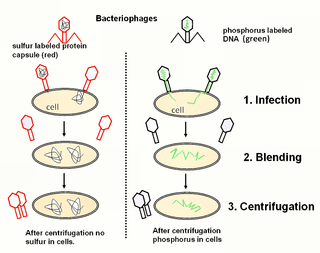
The Hershey–Chase experiments were a series of experiments conducted in 1952 by Alfred Hershey and Martha Chase that helped to confirm that DNA is genetic material.

Enterobacteria phage λ is a bacterial virus, or bacteriophage, that infects the bacterial species Escherichia coli. It was discovered by Esther Lederberg in 1950. The wild type of this virus has a temperate life cycle that allows it to either reside within the genome of its host through lysogeny or enter into a lytic phase, during which it kills and lyses the cell to produce offspring. Lambda strains, mutated at specific sites, are unable to lysogenize cells; instead, they grow and enter the lytic cycle after superinfecting an already lysogenized cell.
Virulence is a pathogen's or microorganism's ability to cause damage to a host.
Viral evolution is a subfield of evolutionary biology and virology that is specifically concerned with the evolution of viruses. Viruses have short generation times, and many—in particular RNA viruses—have relatively high mutation rates. Although most viral mutations confer no benefit and often even prove deleterious to viruses, the rapid rate of viral mutation combined with natural selection allows viruses to quickly adapt to changes in their host environment. In addition, because viruses typically produce many copies in an infected host, mutated genes can be passed on to many offspring quickly. Although the chance of mutations and evolution can change depending on the type of virus, viruses overall have high chances for mutations.

Transduction is the process by which foreign DNA is introduced into a cell by a virus or viral vector. An example is the viral transfer of DNA from one bacterium to another and hence an example of horizontal gene transfer. Transduction does not require physical contact between the cell donating the DNA and the cell receiving the DNA, and it is DNase resistant. Transduction is a common tool used by molecular biologists to stably introduce a foreign gene into a host cell's genome.

The lytic cycle is one of the two cycles of viral reproduction, the other being the lysogenic cycle. The lytic cycle results in the destruction of the infected cell and its membrane. Bacteriophages that only use the lytic cycle are called virulent phages.

Escherichia virus T4 is a species of bacteriophages that infect Escherichia coli bacteria. It is a double-stranded DNA virus in the subfamily Tevenvirinae of the family Straboviridae. T4 is capable of undergoing only a lytic life cycle and not the lysogenic life cycle. The species was formerly named T-even bacteriophage, a name which also encompasses, among other strains, Enterobacteria phage T2, Enterobacteria phage T4 and Enterobacteria phage T6.

Filamentous bacteriophages are a family of viruses (Inoviridae) that infect bacteria, or bacteriophages. They are named for their filamentous shape, a worm-like chain, about 6 nm in diameter and about 1000-2000 nm long. This distinctive shape reflects their method of replication: the coat of the virion comprises five types of viral protein, which are located in the inner membrane of the host bacterium during phage assembly, and these proteins are added to the nascent virion's DNA as it is extruded through the membrane. The simplicity of filamentous phages makes them an appealing model organism for research in molecular biology, and they have also shown promise as tools in nanotechnology and immunology.
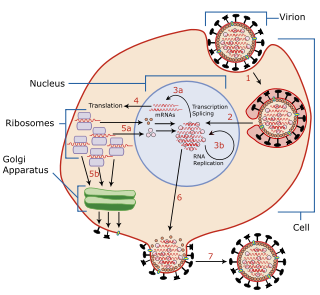
Viral replication is the formation of biological viruses during the infection process in the target host cells. Viruses must first get into the cell before viral replication can occur. Through the generation of abundant copies of its genome and packaging these copies, the virus continues infecting new hosts. Replication between viruses is greatly varied and depends on the type of genes involved in them. Most DNA viruses assemble in the nucleus while most RNA viruses develop solely in cytoplasm.

M13 is one of the Ff phages, a member of the family filamentous bacteriophage (inovirus). Ff phages are composed of circular single-stranded DNA (ssDNA), which in the case of the m13 phage is 6407 nucleotides long and is encapsulated in approximately 2700 copies of the major coat protein p8, and capped with about 5 copies each of four different minor coat proteins. The minor coat protein p3 attaches to the receptor at the tip of the F pilus of the host Escherichia coli. The life cycle is relatively short, with the early phage progeny exiting the cell ten minutes after infection. Ff phages are chronic phage, releasing their progeny without killing the host cells. The infection causes turbid plaques in E. coli lawns, of intermediate opacity in comparison to regular lysis plaques. However, a decrease in the rate of cell growth is seen in the infected cells. The replicative form of M13 is circular double-stranded DNA similar to plasmids that are used for many recombinant DNA processes, and the virus has also been used for phage display, directed evolution, nanostructures and nanotechnology applications.

Lysogeny, or the lysogenic cycle, is one of two cycles of viral reproduction. Lysogeny is characterized by integration of the bacteriophage nucleic acid into the host bacterium's genome or formation of a circular replicon in the bacterial cytoplasm. In this condition the bacterium continues to live and reproduce normally, while the bacteriophage lies in a dormant state in the host cell. The genetic material of the bacteriophage, called a prophage, can be transmitted to daughter cells at each subsequent cell division, and later events can release it, causing proliferation of new phages via the lytic cycle.

Bacteriophage T7 is a bacteriophage, a virus that infects bacteria. It infects most strains of Escherichia coli and relies on these hosts to propagate. Bacteriophage T7 has a lytic life cycle, meaning that it destroys the cell it infects. It also possesses several properties that make it an ideal phage for experimentation: its purification and concentration have produced consistent values in chemical analyses; it can be rendered noninfectious by exposure to UV light; and it can be used in phage display to clone RNA binding proteins.
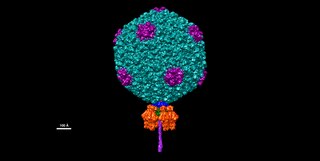
Salmonella virus P22 is a bacteriophage in the Podoviridae family that infects Salmonella typhimurium. Like many phages, it has been used in molecular biology to induce mutations in cultured bacteria and to introduce foreign genetic material. P22 has been used in generalized transduction and is an important tool for investigating Salmonella genetics.
In molecular cloning, a vector is any particle used as a vehicle to artificially carry a foreign nucleic sequence – usually DNA – into another cell, where it can be replicated and/or expressed. A vector containing foreign DNA is termed recombinant DNA. The four major types of vectors are plasmids, viral vectors, cosmids, and artificial chromosomes. Of these, the most commonly used vectors are plasmids. Common to all engineered vectors are an origin of replication, a multicloning site, and a selectable marker.

A virus is a submicroscopic infectious agent that replicates only inside the living cells of an organism. Viruses infect all life forms, from animals and plants to microorganisms, including bacteria and archaea. Viruses are found in almost every ecosystem on Earth and are the most numerous type of biological entity. Since Dmitri Ivanovsky's 1892 article describing a non-bacterial pathogen infecting tobacco plants and the discovery of the tobacco mosaic virus by Martinus Beijerinck in 1898, more than 11,000 of the millions of virus species have been described in detail. The study of viruses is known as virology, a subspeciality of microbiology.

A corynebacteriophage is a DNA-containing bacteriophage specific for bacteria of genus Corynebacterium as its host. Corynebacterium diphtheriae virus strain Corynebacterium diphtheriae phage introduces toxigenicity into strains of Corynebacterium diphtheriae as it encodes diphtheria toxin, it has subtypes beta c and beta vir. According to proposed taxonomic classification, corynephages β and ω are unclassified members of the genus Lambdavirus, family Siphoviridae.

Bacteriophage P2, scientific name Peduovirus P2, is a temperate phage that infects E. coli. It is a tailed virus with a contractile sheath and is thus classified in the genus Peduovirus, family Peduoviridae within class Caudoviricetes. This genus of viruses includes many P2-like phages as well as the satellite phage P4.
Escherichia virus CC31, formerly known as Enterobacter virus CC31, is a dsDNA bacteriophage of the subfamily Tevenvirinae responsible for infecting the bacteria family of Enterobacteriaceae. It is one of two discovered viruses of the genus Karamvirus, diverging away from the previously discovered T4virus, as a clonal complex (CC). CC31 was first isolated from Escherichia coli B strain S/6/4 and is primarily associated with Escherichia, even though is named after Enterobacter.
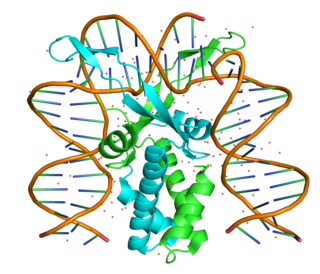
The integration host factor (IHF) is a bacterial DNA binding protein complex that facilitates genetic recombination, replication, and transcription by binding to specific DNA sequences and bending the DNA. It also facilitates the integration of foreign DNA into the host genome. It is a heterodimeric complex composed of two homologous subunits IHFalpha and IHFbeta.
















