Ploidy

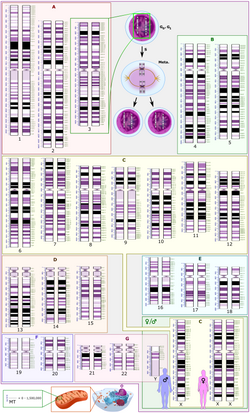
Ploidy refers to the number of complete sets of chromosomes in a cell.
Humans typically have a gene dosage of two. Because they are diploid, they have two sets of 23 different chromosomes. The number of copies of chromosomes generally correlates to the number of copies of a gene present in the genome. For example, the gene that codes for the beta-subunit of hemoglobin (HBB) is located on chromosome 11. Humans have 2 copies of chromosome 11, so they have 2 copies of the HBB gene. [3]
Aneuploidy
If an individual has an abnormal number of chromosomes, then it is called aneuploidy. Aneuploidy is very common in humans, with around 20-40% of all conceptions making a embryo displaying aneuploidy. [4] Most aneuploidy events are fatal and lead to miscarriage. However, there are a few exceptions, including Down Syndrome and intersex conditions. Down Syndrome is caused by trisomy 21, which means having three copies of chromosome 21. Thus gene dosage is increased by 50% for the genes on that chromosome. Though not fully understood, it is thought that the increased expression of genes on chromosome 21 is the cause of some of the characteristic traits of Down syndrome. The intersex condition, Turner syndrome, occurs when a female only has one X chromosome, so she has one sex chromosome. Klinefelter syndrome is another intersex condition where a male has two X chromosomes and one Y chromosome, and so three sex chromosomes. All of these syndromes have characteristic changes in either appearance and/or behavior.
Not all species are diploid like humans [see Polyploidy]. For example, some species of strawberries are octoploid. Those species have eight copies of each chromosome, so they would have eight copies of each gene if that gene has only one copy per chromosome. Some species of wheat are hexaploid, and some species of watermelon are triploid.
Ploidy in prokaryotes
Prokaryotes reproduce through asexual reproduction, usually by binary fission. The bacterial chromosome is present only in one copy per cell. However, there still can be variation in gene dosage due to DNA replication, which starts at the origin of replication and ends at the termination site. The genes that are closer to the origin site are replicated first and are consequently present in two copies in the cell for a longer time than the genes that are closer to the termination site. These slight gene dosage differences are responsible for variation in gene expression depending on their position on the chromosome. [5]