
Thrombin is a serine protease, an enzyme that, in humans, is encoded by the F2 gene. Prothrombin is proteolytically cleaved to form thrombin in the clotting process. Thrombin in turn acts as a serine protease that converts soluble fibrinogen into insoluble strands of fibrin, as well as catalyzing many other coagulation-related reactions.
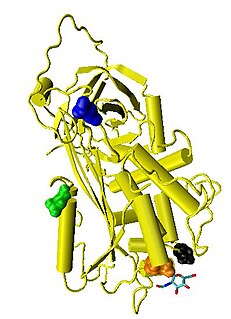
Antithrombin (AT) is a small glycoprotein that inactivates several enzymes of the coagulation system. It is a 432-amino-acid protein produced by the liver. It contains three disulfide bonds and a total of four possible glycosylation sites. α-Antithrombin is the dominant form of antithrombin found in blood plasma and has an oligosaccharide occupying each of its four glycosylation sites. A single glycosylation site remains consistently un-occupied in the minor form of antithrombin, β-antithrombin. Its activity is increased manyfold by the anticoagulant drug heparin, which enhances the binding of antithrombin to factor IIa (Thrombin) and factor Xa.
In biology and biochemistry, protease inhibitors, or antiproteases, are molecules that inhibit the function of proteases. Many naturally occurring protease inhibitors are proteins.

Hirudin is a naturally occurring peptide in the salivary glands of blood-sucking leeches that has a blood anticoagulant property. This is fundamental for the leeches’ habit of feeding on blood, since it keeps a host's blood flowing after the worm's initial puncture of the skin.
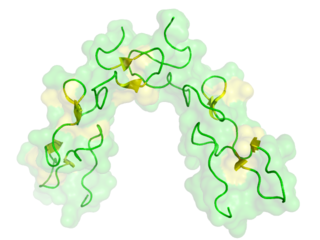
Disintegrins are a family of small proteins from viper venoms that function as potent inhibitors of both platelet aggregation and integrin-dependent cell adhesion.

In structural biology, a beta-propeller (β-propeller) is a type of all-β protein architecture characterized by 4 to 8 highly symmetrical blade-shaped beta sheets arranged toroidally around a central axis. Together the beta-sheets form a funnel-like active site.
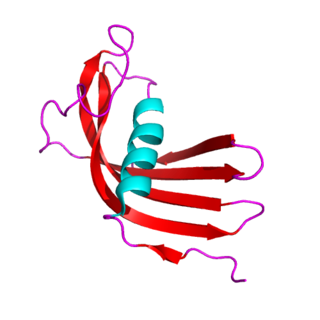
The cystatins are a family of cysteine protease inhibitors which share a sequence homology and a common tertiary structure of an alpha helix lying on top of an anti-parallel beta sheet. The family is subdivided as described below.

A leucine-rich repeat (LRR) is a protein structural motif that forms an α/β horseshoe fold. It is composed of repeating 20–30 amino acid stretches that are unusually rich in the hydrophobic amino acid leucine. These tandem repeats commonly fold together to form a solenoid protein domain, termed leucine-rich repeat domain. Typically, each repeat unit has beta strand-turn-alpha helix structure, and the assembled domain, composed of many such repeats, has a horseshoe shape with an interior parallel beta sheet and an exterior array of helices. One face of the beta sheet and one side of the helix array are exposed to solvent and are therefore dominated by hydrophilic residues. The region between the helices and sheets is the protein's hydrophobic core and is tightly sterically packed with leucine residues.
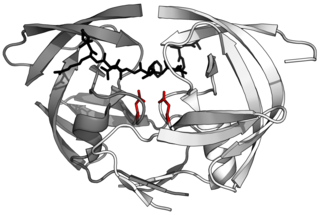
Aspartic proteases are a catalytic type of protease enzymes that use an activated water molecule bound to one or more aspartate residues for catalysis of their peptide substrates. In general, they have two highly conserved aspartates in the active site and are optimally active at acidic pH. Nearly all known aspartyl proteases are inhibited by pepstatin.

Matrix metalloproteinase-1 (MMP-1) also known as interstitial collagenase and fibroblast collagenase is an enzyme that in humans is encoded by the MMP1 gene. The gene is part of a cluster of MMP genes which localize to chromosome 11q22.3. MMP-1 was the first vertebrate collagenase both purified to homogeneity as a protein, and cloned as a cDNA. MMP-1 has an estimated molecular weight of 54 kDa.
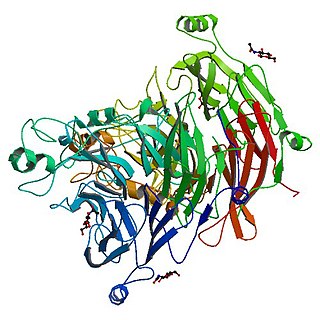
The Sema domain is a structural domain of semaphorins, which are a large family of secreted and transmembrane proteins, some of which function as repellent signals during axon guidance. Sema domains also occur in the hepatocyte growth factor receptor, Plexin-A3 and in viral proteins.
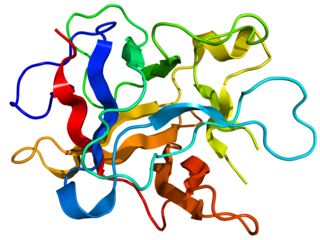
Kunitz soybean trypsin inhibitor is a type of protein contained in legume seeds which functions as a protease inhibitor. Kunitz-type Soybean Trypsin Inhibitors are usually specific for either trypsin or chymotrypsin. They are thought to protect seeds against consumption by animal predators.
Sushi domain is an evolutionarily conserved protein domain. It is also known as Complement control protein (CCP) modules or short consensus repeats (SCR). The name derives from the visual similarity of the domain to nigiri sushi when the primary structure is drawn showing the loops created by the disulfide bonds.
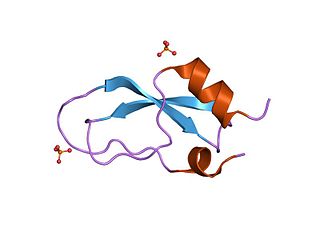
Kunitz domains are the active domains of proteins that inhibit the function of protein degrading enzymes or, more specifically, domains of Kunitz-type are protease inhibitors. They are relatively small with a length of about 50 to 60 amino acids and a molecular weight of 6 kDa. Examples of Kunitz-type protease inhibitors are aprotinin, Alzheimer's amyloid precursor protein (APP), and tissue factor pathway inhibitor (TFPI). Kunitz STI protease inhibitor, the trypsin inhibitor initially studied by Moses Kunitz, was extracted from soybeans.
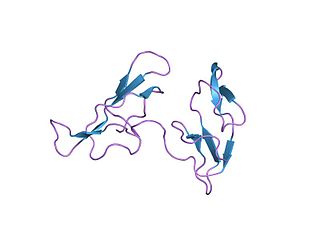
In molecular biology, the Bowman–Birk protease inhibitor family of proteins consists of eukaryotic proteinase inhibitors, belonging to MEROPS inhibitor family I12, clan IF. They mainly inhibit serine peptidases of the S1 family, but also inhibit S3 peptidases.
The Kazal domain is an evolutionary conserved protein domain usually indicative of serine protease inhibitors. However, kazal-like domains are also seen in the extracellular part of agrins, which are not known to be protease inhibitors.

In molecular biology, the carboxypeptidase A inhibitor family is a family of proteins which is represented by the well-characterised metallocarboxypeptidase A inhibitor (MCPI) from potatoes, which belongs to the MEROPS inhibitor family I37, clan IE. It inhibits metallopeptidases belonging to MEROPS peptidase family M14, carboxypeptidase A. In Russet Burbank potatoes, it is a mixture of approximately equal amounts of two polypeptide chains containing 38 or 39 amino acid residues. The chains differ in their amino terminal sequence only and are resistant to fragmentation by proteases. The structure of the complex between bovine carboxypeptidase A and the 39-amino-acid carboxypeptidase A inhibitor from potatoes has been determined at 2.5-Angstrom resolution.

In molecular biology, a carbohydrate-binding module (CBM) is a protein domain found in carbohydrate-active enzymes. The majority of these domains have carbohydrate-binding activity. Some of these domains are found on cellulosomal scaffoldin proteins. CBMs were previously known as cellulose-binding domains. CBMs are classified into numerous families, based on amino acid sequence similarity. There are currently 64 families of CBM in the CAZy database.

In molecular biology, D-stereospecific aminopeptidase (D-aminopeptidase) EC 3.4.11.19 is an enzyme which catalyses the release of an N-terminal D-amino acid from a peptide, Xaa-|-Yaa-, in which Xaa is preferably D-Ala, D-Ser or D-Thr. D-amino acid amides and methyl esters also are hydrolyzed, as is glycine amide.

Pacifastin is a family of serine proteinase inhibitors found in arthropods. Pacifastin inhibits the serine peptidases trypsin and chymotrypsin.


















