Related Research Articles
A phylogenetic tree, phylogeny or evolutionary tree is a graphical representation which shows the evolutionary history between a set of species or taxa during a specific time. In other words, it is a branching diagram or a tree showing the evolutionary relationships among various biological species or other entities based upon similarities and differences in their physical or genetic characteristics. In evolutionary biology, all life on Earth is theoretically part of a single phylogenetic tree, indicating common ancestry. Phylogenetics is the study of phylogenetic trees. The main challenge is to find a phylogenetic tree representing optimal evolutionary ancestry between a set of species or taxa. Computational phylogenetics focuses on the algorithms involved in finding optimal phylogenetic tree in the phylogenetic landscape.

BioPerl is a collection of Perl modules that facilitate the development of Perl scripts for bioinformatics applications. It has played an integral role in the Human Genome Project.
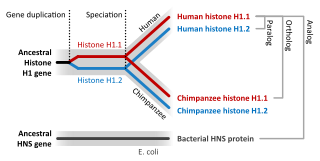
Sequence homology is the biological homology between DNA, RNA, or protein sequences, defined in terms of shared ancestry in the evolutionary history of life. Two segments of DNA can have shared ancestry because of three phenomena: either a speciation event (orthologs), or a duplication event (paralogs), or else a horizontal gene transfer event (xenologs).
A phylogenetic network is any graph used to visualize evolutionary relationships between nucleotide sequences, genes, chromosomes, genomes, or species. They are employed when reticulation events such as hybridization, horizontal gene transfer, recombination, or gene duplication and loss are believed to be involved. They differ from phylogenetic trees by the explicit modeling of richly linked networks, by means of the addition of hybrid nodes instead of only tree nodes. Phylogenetic trees are a subset of phylogenetic networks. Phylogenetic networks can be inferred and visualised with software such as SplitsTree, the R-package, phangorn, and, more recently, Dendroscope. A standard format for representing phylogenetic networks is a variant of Newick format which is extended to support networks as well as trees.
Computational phylogenetics, phylogeny inference, or phylogenetic inference focuses on computational and optimization algorithms, heuristics, and approaches involved in phylogenetic analyses. The goal is to find a phylogenetic tree representing optimal evolutionary ancestry between a set of genes, species, or taxa. Maximum likelihood, parsimony, Bayesian, and minimum evolution are typical optimality criteria used to assess how well a phylogenetic tree topology describes the sequence data. Nearest Neighbour Interchange (NNI), Subtree Prune and Regraft (SPR), and Tree Bisection and Reconnection (TBR), known as tree rearrangements, are deterministic algorithms to search for optimal or the best phylogenetic tree. The space and the landscape of searching for the optimal phylogenetic tree is known as phylogeny search space.

Noah Aubrey Rosenberg is a geneticist working in evolutionary biology, mathematical phylogenetics, and population genetics, and is the Stanford Professor of Population Genetics and Society. His research focuses on mathematical modeling and statistical methods for genetics and evolution and he is the editor-in-chief of Theoretical Population Biology.

Masatoshi Nei was a Japanese-born American evolutionary biologist.
Ancestral reconstruction is the extrapolation back in time from measured characteristics of individuals, populations, or species to their common ancestors. It is an important application of phylogenetics, the reconstruction and study of the evolutionary relationships among individuals, populations or species to their ancestors. In the context of evolutionary biology, ancestral reconstruction can be used to recover different kinds of ancestral character states of organisms that lived millions of years ago. These states include the genetic sequence, the amino acid sequence of a protein, the composition of a genome, a measurable characteristic of an organism (phenotype), and the geographic range of an ancestral population or species. This is desirable because it allows us to examine parts of phylogenetic trees corresponding to the distant past, clarifying the evolutionary history of the species in the tree. Since modern genetic sequences are essentially a variation of ancient ones, access to ancient sequences may identify other variations and organisms which could have arisen from those sequences. In addition to genetic sequences, one might attempt to track the changing of one character trait to another, such as fins turning to legs.
Bayesian inference of phylogeny combines the information in the prior and in the data likelihood to create the so-called posterior probability of trees, which is the probability that the tree is correct given the data, the prior and the likelihood model. Bayesian inference was introduced into molecular phylogenetics in the 1990s by three independent groups: Bruce Rannala and Ziheng Yang in Berkeley, Bob Mau in Madison, and Shuying Li in University of Iowa, the last two being PhD students at the time. The approach has become very popular since the release of the MrBayes software in 2001, and is now one of the most popular methods in molecular phylogenetics.

Roderic Dugald Morton Page is a New Zealand-born evolutionary biologist at the University of Glasgow, Scotland, and the author of several books. As of 2015 he is professor at the University of Glasgow and was editor of the journal Systematic Biology until the end of 2007. His main interests are in phylogenetics, evolutionary biology and bioinformatics.
TREE-PUZZLE is a computer program used to construct phylogenetic trees from sequence data by maximum likelihood analysis. Branch lengths can be calculated with and without the molecular clock hypothesis.

SplitsTree is a popular freeware program for inferring phylogenetic trees, phylogenetic networks, or, more generally, splits graphs, from various types of data such as a sequence alignment, a distance matrix or a set of trees. SplitsTree implements published methods such as split decomposition, neighbor-net, consensus networks, super networks methods or methods for computing hybridization or simple recombination networks. It uses the NEXUS file format. The splits graph is defined using a special data block.
Biological data visualization is a branch of bioinformatics concerned with the application of computer graphics, scientific visualization, and information visualization to different areas of the life sciences. This includes visualization of sequences, genomes, alignments, phylogenies, macromolecular structures, systems biology, microscopy, and magnetic resonance imaging data. Software tools used for visualizing biological data range from simple, standalone programs to complex, integrated systems.

NeighborNet is an algorithm for constructing phylogenetic networks which is loosely based on the neighbor joining algorithm. Like neighbor joining, the method takes a distance matrix as input, and works by agglomerating clusters. However, the NeighborNet algorithm can lead to collections of clusters which overlap and do not form a hierarchy, and are represented using a type of phylogenetic network called a splits graph. If the distance matrix satisfies the Kalmanson combinatorial conditions then Neighbor-net will return the corresponding circular ordering. The method is implemented in the SplitsTree and R/Phangorn packages.
T-REX is a freely available web server, developed at the department of Computer Science of the Université du Québec à Montréal, dedicated to the inference, validation and visualization of phylogenetic trees and phylogenetic networks. The T-REX web server allows the users to perform several popular methods of phylogenetic analysis as well as some new phylogenetic applications for inferring, drawing and validating phylogenetic trees and networks.
Horizontal or lateral gene transfer is the transmission of portions of genomic DNA between organisms through a process decoupled from vertical inheritance. In the presence of HGT events, different fragments of the genome are the result of different evolutionary histories. This can therefore complicate investigations of the evolutionary relatedness of lineages and species. Also, as HGT can bring into genomes radically different genotypes from distant lineages, or even new genes bearing new functions, it is a major source of phenotypic innovation and a mechanism of niche adaptation. For example, of particular relevance to human health is the lateral transfer of antibiotic resistance and pathogenicity determinants, leading to the emergence of pathogenic lineages.
Arndt von Haeseler is a German bioinformatician and evolutionary biologist. He is the scientific director of the Max F. Perutz Laboratories at the Vienna Biocenter and a professor of bioinformatics at the University of Vienna and the Medical University of Vienna.

In phylogenetics, reconciliation is an approach to connect the history of two or more coevolving biological entities. The general idea of reconciliation is that a phylogenetic tree representing the evolution of an entity can be drawn within another phylogenetic tree representing an encompassing entity to reveal their interdependence and the evolutionary events that have marked their shared history. The development of reconciliation approaches started in the 1980s, mainly to depict the coevolution of a gene and a genome, and of a host and a symbiont, which can be mutualist, commensalist or parasitic. It has also been used for example to detect horizontal gene transfer, or understand the dynamics of genome evolution.
References
- ↑ Birth year from Library of Congress catalog entry, retrieved 2022-03-05
- 1 2 "Katharina Huber", People, University of East Anglia, retrieved 2022-03-05
- ↑ Katharina T. Huber at the Mathematics Genealogy Project
- ↑ Reviews of Basic Phylogenetic Combinatorics:
- Fletez-Brant, Kipper (September 2014), "Review" (PDF), SIGACT News , 45 (3): 26–28, doi:10.1145/2670418.2670427, MR 3266629, S2CID 17544618
- van Iersel, Leo (March 2013), Systematic Biology, 62 (2): 346–348, JSTOR 23485144
{{citation}}: CS1 maint: untitled periodical (link) - Morrison, David A. (March 2013), The Quarterly Review of Biology, 88 (1): 33, doi:10.1086/669305, JSTOR 10.1086/669305
{{citation}}: CS1 maint: untitled periodical (link) - Willson, Stephen J., Mathematical Reviews, MR 2893879
{{citation}}: CS1 maint: untitled periodical (link)
- ↑ Popescu, Andrei-Alin; Huber, Katharina T.; Paradis, Emmanuel (April 2012), "ape 3.0: New tools for distance-based phylogenetics and evolutionary analysis in R", Bioinformatics, 28 (11): 1536–1537, doi: 10.1093/bioinformatics/bts184 , PMID 22495750