Related Research Articles

A Morpholino, also known as a Morpholino oligomer and as a phosphorodiamidate Morpholino oligomer (PMO), is a type of oligomer molecule used in molecular biology to modify gene expression. Its molecular structure contains DNA bases attached to a backbone of methylenemorpholine rings linked through phosphorodiamidate groups. Morpholinos block access of other molecules to small specific sequences of the base-pairing surfaces of ribonucleic acid (RNA). Morpholinos are used as research tools for reverse genetics by knocking down gene function.
The epithelial–mesenchymal transition (EMT) is a process by which epithelial cells lose their cell polarity and cell–cell adhesion, and gain migratory and invasive properties to become mesenchymal stem cells; these are multipotent stromal cells that can differentiate into a variety of cell types. EMT is essential for numerous developmental processes including mesoderm formation and neural tube formation. EMT has also been shown to occur in wound healing, in organ fibrosis and in the initiation of metastasis in cancer progression.
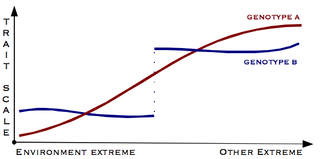
Canalisation is a measure of the ability of a population to produce the same phenotype regardless of variability of its environment or genotype. It is a form of evolutionary robustness. The term was coined in 1942 by C. H. Waddington to capture the fact that "developmental reactions, as they occur in organisms submitted to natural selection...are adjusted so as to bring about one definite end-result regardless of minor variations in conditions during the course of the reaction". He used this word rather than robustness to consider that biological systems are not robust in quite the same way as, for example, engineered systems.
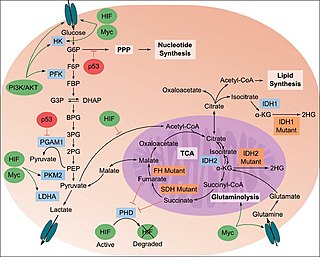
The study of the tumor metabolism, also known as tumor metabolome describes the different characteristic metabolic changes in tumor cells. The characteristic attributes of the tumor metabolome are high glycolytic enzyme activities, the expression of the pyruvate kinase isoenzyme type M2, increased channeling of glucose carbons into synthetic processes, such as nucleic acid, amino acid and phospholipid synthesis, a high rate of pyrimidine and purine de novo synthesis, a low ratio of Adenosine triphosphate and Guanosine triphosphate to Cytidine triphosphate and Uridine triphosphate, low Adenosine monophosphate levels, high glutaminolytic capacities, release of immunosuppressive substances and dependency on methionine.

Chemogenomics, or chemical genomics, is the systematic screening of targeted chemical libraries of small molecules against individual drug target families with the ultimate goal of identification of novel drugs and drug targets. Typically some members of a target library have been well characterized where both the function has been determined and compounds that modulate the function of those targets have been identified. Other members of the target family may have unknown function with no known ligands and hence are classified as orphan receptors. By identifying screening hits that modulate the activity of the less well characterized members of the target family, the function of these novel targets can be elucidated. Furthermore, the hits for these targets can be used as a starting point for drug discovery. The completion of the human genome project has provided an abundance of potential targets for therapeutic intervention. Chemogenomics strives to study the intersection of all possible drugs on all of these potential targets.
Genetic assimilation is a process described by Conrad H. Waddington by which a phenotype originally produced in response to an environmental condition, such as exposure to a teratogen, later becomes genetically encoded via artificial selection or natural selection. Despite superficial appearances, this does not require the (Lamarckian) inheritance of acquired characters, although epigenetic inheritance could potentially influence the result. Waddington stated that genetic assimilation overcomes the barrier to selection imposed by what he called canalization of developmental pathways; he supposed that the organism's genetics evolved to ensure that development proceeded in a certain way regardless of normal environmental variations.

The nuclear receptor 4A2 (NR4A2) also known as nuclear receptor related 1 protein (NURR1) is a protein that in humans is encoded by the NR4A2 gene. NR4A2 is a member of the nuclear receptor family of intracellular transcription factors.

Telomerase reverse transcriptase is a catalytic subunit of the enzyme telomerase, which, together with the telomerase RNA component (TERC), comprises the most important unit of the telomerase complex.

Nuclear factor erythroid 2-related factor 2 (NRF2), also known as nuclear factor erythroid-derived 2-like 2, is a transcription factor that in humans is encoded by the NFE2L2 gene. NRF2 is a basic leucine zipper (bZIP) protein that may regulate the expression of antioxidant proteins that protect against oxidative damage triggered by injury and inflammation, according to preliminary research. In vitro, NRF2 binds to antioxidant response elements (AREs) in the promoter regions of genes encoding cytoprotective proteins. NRF2 induces the expression of heme oxygenase 1 in vitro leading to an increase in phase II enzymes. NRF2 also inhibits the NLRP3 inflammasome.

MAP kinase-activated protein kinase 2 is an enzyme that in humans is encoded by the MAPKAPK2 gene.

Mitogen-activated protein kinase 7 also known as MAP kinase 7 is an enzyme that in humans is encoded by the MAPK7 gene.

DNA damage-inducible transcript 3, also known as C/EBP homologous protein (CHOP), is a pro-apoptotic transcription factor that is encoded by the DDIT3 gene. It is a member of the CCAAT/enhancer-binding protein (C/EBP) family of DNA-binding transcription factors. The protein functions as a dominant-negative inhibitor by forming heterodimers with other C/EBP members, preventing their DNA binding activity. The protein is implicated in adipogenesis and erythropoiesis and has an important role in the cell's stress response.

Histone deacetylase 9 is an enzyme that in humans is encoded by the HDAC9 gene.

Transcription factor Sp7, also called osterix (Osx), is a protein that in humans is encoded by the SP7 gene. It is a member of the Sp family of zinc-finger transcription factors It is highly conserved among bone-forming vertebrate species It plays a major role, along with Runx2 and Dlx5 in driving the differentiation of mesenchymal precursor cells into osteoblasts and eventually osteocytes. Sp7 also plays a regulatory role by inhibiting chondrocyte differentiation maintaining the balance between differentiation of mesenchymal precursor cells into ossified bone or cartilage. Mutations of this gene have been associated with multiple dysfunctional bone phenotypes in vertebrates. During development, a mouse embryo model with Sp7 expression knocked out had no formation of bone tissue. Through the use of GWAS studies, the Sp7 locus in humans has been strongly associated with bone mass density. In addition there is significant genetic evidence for its role in diseases such as Osteogenesis imperfecta (OI).
A gene signature or gene expression signature is a single or combined group of genes in a cell with a uniquely characteristic pattern of gene expression that occurs as a result of an altered or unaltered biological process or pathogenic medical condition. This is not to be confused with the concept of gene expression profiling. Activating pathways in a regular physiological process or a physiological response to a stimulus results in a cascade of signal transduction and interactions that elicit altered levels of gene expression, which is classified as the gene signature of that physiological process or response. The clinical applications of gene signatures breakdown into prognostic, diagnostic and predictive signatures. The phenotypes that may theoretically be defined by a gene expression signature range from those that predict the survival or prognosis of an individual with a disease, those that are used to differentiate between different subtypes of a disease, to those that predict activation of a particular pathway. Ideally, gene signatures can be used to select a group of patients for whom a particular treatment will be effective.
Phenotypic screening is a type of screening used in biological research and drug discovery to identify substances such as small molecules, peptides, or RNAi that alter the phenotype of a cell or an organism in a desired manner. Phenotypic screening must be followed up with identification and validation, often through the use of chemoproteomics, to identify the mechanisms through which a phenotypic hit works.
Chemical genetics is the investigation of the function of proteins and signal transduction pathways in cells by the screening of chemical libraries of small molecules. Chemical genetics is analogous to classical genetic screen where random mutations are introduced in organisms, the phenotype of these mutants is observed, and finally the specific gene mutation (genotype) that produced that phenotype is identified. In chemical genetics, the phenotype is disturbed not by introduction of mutations, but by exposure to small molecule tool compounds. Phenotypic screening of chemical libraries is used to identify drug targets or to validate those targets in experimental models of disease. Recent applications of this topic have been implicated in signal transduction, which may play a role in discovering new cancer treatments. Chemical genetics can serve as a unifying study between chemistry and biology. The approach was first proposed by Tim Mitchison in 1994 in an opinion piece in the journal Chemistry & Biology entitled "Towards a pharmacological genetics".

The universal stress protein (USP) domain is a superfamily of conserved genes which can be found in bacteria, archaea, fungi, protozoa and plants. Proteins containing the domain are induced by many environmental stressors such as nutrient starvation, drought, extreme temperatures, high salinity, and the presence of uncouplers, antibiotics and metals.
Perturb-seq refers to a high-throughput method of performing single cell RNA sequencing (scRNA-seq) on pooled genetic perturbation screens. Perturb-seq combines multiplexed CRISPR mediated gene inactivations with single cell RNA sequencing to assess comprehensive gene expression phenotypes for each perturbation. Inferring a gene’s function by applying genetic perturbations to knock down or knock out a gene and studying the resulting phenotype is known as reverse genetics. Perturb-seq is a reverse genetics approach that allows for the investigation of phenotypes at the level of the transcriptome, to elucidate gene functions in many cells, in a massively parallel fashion.
Dedifferentiation is a transient process by which cells become less specialized and return to an earlier cell state within the same lineage. This suggests an increase in cell potency, meaning that, following dedifferentiation, a cell may possess the ability to re-differentiate into more cell types than it did before dedifferentiation. This is in contrast to differentiation, where differences in gene expression, morphology, or physiology arise in a cell, making its function increasingly specialized.
References
- ↑ Zhang, JD; Küng, E; Boess, F; Certa, U; Ebeling, M (24 April 2015). "Pathway reporter genes define molecular phenotypes of human cells". BMC Genomics. 16 (1): 342. doi: 10.1186/s12864-015-1532-2 . PMC 4415216 . PMID 25903797.
- ↑ Zhang, JD; Schindler, T; Küng, E; Ebeling, M; Certa, U (5 July 2014). "Highly sensitive amplicon-based transcript quantification by semiconductor sequencing". BMC Genomics. 15 (1): 565. doi: 10.1186/1471-2164-15-565 . PMC 4101174 . PMID 24997760.
- ↑ Zhang, JD; Berntenis, N; Roth, A; Ebeling, M (June 2014). "Data mining reveals a network of early-response genes as a consensus signature of drug-induced in vitro and in vivo toxicity". The Pharmacogenomics Journal. 14 (3): 208–16. doi:10.1038/tpj.2013.39. PMC 4034126 . PMID 24217556.
- ↑ Moffat, JG (18 May 2017). "Turning the Light On in the Phenotypic Drug Discovery Black Box". Cell Chemical Biology. 24 (5): 545–547. doi: 10.1016/j.chembiol.2017.05.005 . PMID 28525769.
- ↑ Drawnel, FM; Zhang, JD; Küng, E; Aoyama, N; Benmansour, F; Araujo Del Rosario, A; Jensen Zoffmann, S; Delobel, F; Prummer, M; Weibel, F; Carlson, C; Anson, B; Iacone, R; Certa, U; Singer, T; Ebeling, M; Prunotto, M (18 May 2017). "Molecular Phenotyping Combines Molecular Information, Biological Relevance, and Patient Data to Improve Productivity of Early Drug Discovery". Cell Chemical Biology. 24 (5): 624–634.e3. doi: 10.1016/j.chembiol.2017.03.016 . PMID 28434878.