
In genetics, a promoter is a sequence of DNA to which proteins bind to initiate transcription of a single RNA transcript from the DNA downstream of the promoter. The RNA transcript may encode a protein (mRNA), or can have a function in and of itself, such as tRNA or rRNA. Promoters are located near the transcription start sites of genes, upstream on the DNA . Promoters can be about 100–1000 base pairs long, the sequence of which is highly dependent on the gene and product of transcription, type or class of RNA polymerase recruited to the site, and species of organism.

The ykkC/yxkD leader is a conserved RNA structure found upstream of the ykkC and yxkD genes in Bacillus subtilis and related genes in other bacteria. The function of this family is unclear for many years although it has been suggested that it may function to switch on efflux pumps and detoxification systems in response to harmful environmental molecules. The Thermoanaerobacter tengcongensis sequence AE013027 overlaps with that of purine riboswitch suggesting that the two riboswitches may work in conjunction to regulate the upstream gene which codes for TTE0584 (Q8RC62), a member of the permease family.
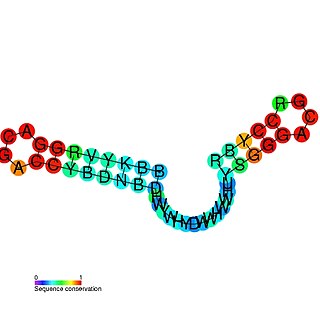
The mini-ykkC RNA motif was discovered as a putative RNA structure that is conserved in bacteria. The motif consists of two conserved stem-loops whose terminal loops contain the RNA sequence ACGR, where R represents either A or G. Mini-ykkC RNAs are widespread in Pseudomonadota, but some are predicted in other phyla of bacteria. It was expected that the RNAs are cis-regulatory elements, because they are typically located upstream of protein-coding genes.

The ATPC RNA motif is a conserved RNA structure found in certain cyanobacteria. It is apparently ubiquitous in Prochlorococcus marinus, and is present in many species in the genus Synechococcus. The RNA is always found within an operon encoding subunits of ATP synthase, and it is always located downstream of the gene encoding the A subunit of ATP synthase, and upstream of the C subunit gene. This location is consistent with a cis-regulatory element, but also with a non-coding RNA that is transcribed with the ATP synthase genes.

PhotoRC RNA motifs refer to conserved RNA structures that are associated with genes acting in the photosynthetic reaction centre of photosynthetic bacteria. Two such RNA classes were identified and called the PhotoRC-I and PhotoRC-II motifs. PhotoRC-I RNAs were detected in the genomes of some cyanobacteria. Although no PhotoRC-II RNA has been detected in cyanobacteria, one is found in the genome of a purified phage that infects cyanobacteria. Both PhotoRC-I and PhotoRC-II RNAs are present in sequences derived from DNA that was extracted from uncultivated marine bacteria.
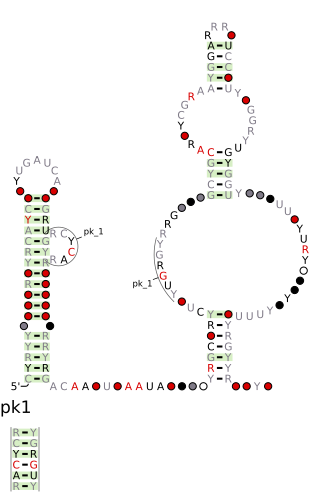
The DUF2800 RNA motif is a conserved RNA structure that was discovered by bioinformatics. DUF2800 motif RNAs are found in Bacillota. DUF2800 RNAs are also predicted in the phyla Actinomycetota and Synergistota, although these RNAs are likely the result of recent horizontal gene transfer or conceivably sequence contamination.

The DUF805 RNA motif is a conserved RNA structure that was discovered by bioinformatics. The motif is subdivided into the DUF805 motif and the DUF805b motif, which have similar, but distinct secondary structures. Together, these motifs are found in Bacteroidota, Chlorobiota, and Pseudomonadota.

The folE RNA motif, now known as the THF-II riboswitch, is a conserved RNA structure that was discovered by bioinformatics. folE motifs are found in Alphaproteobacteria.

The hya RNA motif is a conserved RNA structure that was discovered by bioinformatics. hya motif RNAs are found in Actinomycetota.
The leuA-Halobacteria RNA motif is a conserved RNA structure that was discovered by bioinformatics. leuA-Halobacteria motifs are found in Halobacteriaceae, a lineage of archaea.
malK RNA motifs are conserved RNA structures that were discovered by bioinformatics. They are defined by being consistently located upstream of malK genes, which encode an ATPase that is used by transporters whose ligand is likely a kind of sugar. Most of these genes are annotated either as transporting maltose or glycerol-3-phosphate, however the substrate of the transporters associated with malK motif RNAs has not been experimentally determined. All known types of malK RNA motif are generally located nearby to the Shine-Dalgarno sequence of the downstream gene.

The MDR-NUDIX RNA motif is a conserved RNA structure that was discovered by bioinformatics. The MDR-NUDIX motif is found in the poorly studied phylum TM7.

The nhaA-I RNA motif is a conserved RNA structure that was discovered by bioinformatics. nhaA-I motif RNAs are found in Acidobacteriota, alpha-, beta- and Gammaproteobacteria, Verrucomicrobiota and the tentative phylum NC10.

The nrdJ RNA motif is a conserved RNA structure that was discovered by bioinformatics. The nrdJ motif is found in the genus Streptomyces.
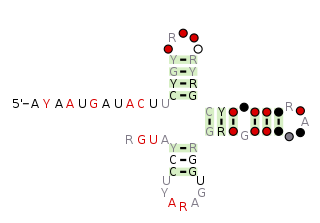
The pemK RNA motif is a conserved RNA structure that was discovered by bioinformatics. pemK motif RNAs are found in organisms within the phylum Bacillota, and is very widespread in this phylum.

The PGK RNA motif is a conserved RNA structure that was discovered by bioinformatics. PGK motif RNAs are found in metagenomic sequences isolated from the gastrointestinal tract of mammals. PGK RNAs have not yet been detected in a classified organism.
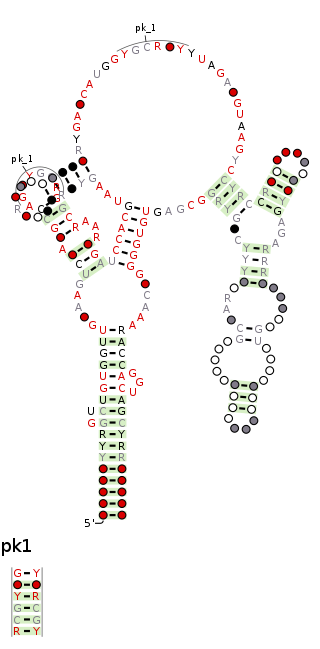
The raiA RNA motif is a conserved RNA structure that was discovered by bioinformatics. raiA motif RNAs are found in Actinomycetota and Bacillota, and have many conserved features—including conserved nucleotide positions, conserved secondary structures and associated protein-coding genes—in both of these phyla. Some conserved features of the raiA RNA motif suggest that they function as cis-regulatory elements, but other aspects of the motif suggest otherwise.

The sul1 RNA motif is a conserved RNA structure that was discovered by bioinformatics. Energetically stable tetraloops often occur in this motif. sul1 motif RNAs are found in Alphaproteobacteria.

The uup RNA motif is a conserved RNA structure that was discovered by bioinformatics. uup motif RNAs are found in Bacillota and Gammaproteobacteria.
The icd-II non-coding RNA (ncRNA) is an RNA motif proposed as a Strong Riboswitch Candidate (SRC). Icd-II ncRNA has been recognized by a comparative sequence analysis in GC-rich intergenic regions (IGR) of bacteria, using a pipeline call Discovery of Intergenic Motifs PipeLine (DIMPL). Icd-II ncRNA has been located upstream of the icd gene, which codes for an NADP+-dependent isocitrate dehydrogenase (IDH) enzyme. IDH is part of the citric acid cycle, and thus it participates in managing the carbon flux through this energy metabolism pathway. Icd-II ncRNA has been found in bacteria of the class beta proteobacteria, particularly in Polynucleobacter genus. Icd-II RNA secondary structure consists of a three-stem junction, where the ribosome binding site (RBS) of the adjacent open reading frame (ORF) is predicted to be involved in the first base-paired stem. It has been proposed that icd-II ncRNA can function as a riboswitch that regulates translation initiation of its associate ORF.

















