Succinate dehydrogenase (SDH) or succinate-coenzyme Q reductase (SQR) or respiratory Complex II is an enzyme complex, found in many bacterial cells and in the inner mitochondrial membrane of eukaryotes. It is the only enzyme that participates in both the citric acid cycle and the electron transport chain. Histochemical analysis showing high succinate dehydrogenase in muscle demonstrates high mitochondrial content and high oxidative potential.
The gene rpoS encodes the sigma factor sigma-38, a 37.8 kD protein in Escherichia coli. Sigma factors are proteins that regulate transcription in bacteria. Sigma factors can be activated in response to different environmental conditions. rpoS is transcribed in late exponential phase, and RpoS is the primary regulator of stationary phase genes. RpoS is a central regulator of the general stress response and operates in both a retroactive and a proactive manner: it not only allows the cell to survive environmental challenges, but it also prepares the cell for subsequent stresses (cross-protection). The transcriptional regulator CsgD is central to biofilm formation, controlling the expression of the curli structural and export proteins, and the diguanylate cyclase, adrA, which indirectly activates cellulose production. The rpoS gene most likely originated in the gammaproteobacteria.

DicF RNA is a non-coding RNA that is an antisense inhibitor of cell division gene ftsZ. DicF is bound by the Hfq protein which enhances its interaction with its targets. Pathogenic E. coli strains possess multiple copies of sRNA DicF in their genomes, while no-pathogenic strains do not.DicF and Hfq are both necessary to reduce FtsZ protein levels, leading to cell filamentation under anaerobic conditions.

DsrA RNA is a non-coding RNA that regulates both transcription, by overcoming transcriptional silencing by the nucleoid-associated H-NS protein, and translation, by promoting efficient translation of the stress sigma factor, RpoS. These two activities of DsrA can be separated by mutation: the first of three stem-loops of the 85 nucleotide RNA is necessary for RpoS translation but not for anti-H-NS action, while the second stem-loop is essential for antisilencing and less critical for RpoS translation. The third stem-loop, which behaves as a transcription terminator, can be substituted by the trp transcription terminator without loss of either DsrA function. The sequence of the first stem-loop of DsrA is complementary with the upstream leader portion of RpoS messenger RNA, suggesting that pairing of DsrA with the RpoS message might be important for translational regulation. The structures of DsrA and DsrA/rpoS complex were studied by NMR. The study concluded that the sRNA contains a dynamic conformational equilibrium for its second stem–loop which might be an important mechanism for DsrA to regulate the translations of its multiple target mRNAs.

The gcvB RNA gene encodes a small non-coding RNA involved in the regulation of a number of amino acid transport systems as well as amino acid biosynthetic genes. The GcvB gene is found in enteric bacteria such as Escherichia coli. GcvB regulates genes by acting as an antisense binding partner of the mRNAs for each regulated gene. This binding is dependent on binding to a protein called Hfq. Transcription of the GcvB RNA is activated by the adjacent GcvA gene and repressed by the GcvR gene. A deletion of GcvB RNA from Y. pestis changed colony shape as well as reducing growth. It has been shown by gene deletion that GcvB is a regulator of acid resistance in E. coli. GcvB enhances the ability of the bacterium to survive low pH by upregulating the levels of the alternate sigma factor RpoS. A polymeric form of GcvB has recently been identified. Interaction of GcvB with small RNA SroC triggers the degradation of GcvB by RNase E, lifting the GcvB-mediated mRNA repression of its target genes.
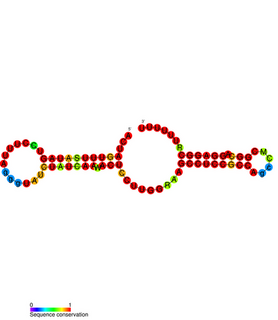
The RprA RNA gene encodes a 106 nucleotide regulatory non-coding RNA. Translational regulation of the stationary phase sigma factor RpoS is mediated by the formation of a double-stranded RNA stem-loop structure in the upstream region of the rpoS messenger RNA, occluding the translation initiation site. Clones carrying rprA increased the translation of RpoS. As with DsrA, RprA is predicted to form three stem-loops. Thus, at least two small RNAs, DsrA and RprA, participate in the positive regulation of RpoS translation. RprA also appears to bind to the RpoS leader. RprA is non-essential. Wasserman et al. demonstrated that this RNA is bound by the Hfq protein. Binding to Hfq alters the conformation of RprA. In the presence of Hfq the stability of RprA is influenced by the osmolarity of the cell, this is dependent on the endoribonuclease RNase E.

RyhB RNA is a 90 nucleotide RNA that down-regulates a set of iron-storage and iron-using proteins when iron is limiting; it is itself negatively regulated by the ferric uptake repressor protein, Fur.

Spot 42 (spf) RNA is a regulatory non-coding bacterial small RNA encoded by the spf gene. Spf is found in gammaproteobacteria and the majority of experimental work on Spot42 has been performed in Escherichia coli and recently in Aliivibrio salmonicida. In the cell Spot42 plays essential roles as a regulator in carbohydrate metabolism and uptake, and its expression is activated by glucose, and inhibited by the cAMP-CRP complex.

In molecular biology the ArcZ RNA is a small non-coding RNA (ncRNA). It is the functional product of a gene which is not translated into protein. ArcZ is an Hfq binding RNA that functions as an antisense regulator of a number of protein coding genes.

The Hfq protein encoded by the hfq gene was discovered in 1968 as an Escherichia coli host factor that was essential for replication of the bacteriophage Qβ. It is now clear that Hfq is an abundant bacterial RNA binding protein which has many important physiological roles that are usually mediated by interacting with Hfq binding sRNA.

Listeria monocytogenes is a gram positive bacterium and causes many food-borne infections such as Listeriosis. This bacteria is ubiquitous in the environment where it can act as either a saprophyte when free living within the environment or as a pathogen when entering a host organism. Many non-coding RNAs have been identified within the bacteria genome where several of these have been classified as novel non-coding RNAs and may contribute to pathogenesis.

MicX sRNA is a small non-coding RNA found in Vibrio cholerae. It was given the name MicX as it has a similar function to MicA, MicC and MicF in E. coli. MicX sRNA negatively regulates an outer membrane protein and also a component of an ABC transporter. These interactions were predicted and then confirmed using a DNA microarray.
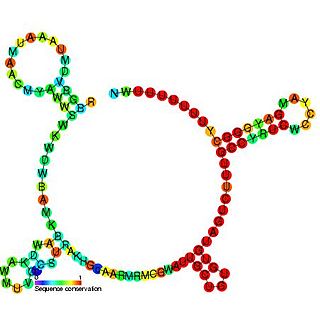
FnrS RNA is a family of Hfq-binding small RNA whose expression is upregulated in response to anaerobic conditions. It is named FnrS because its expression is strongly dependent on fumarate and nitrate reductase regulator (FNR), a direct oxygen availability sensor.
Rsa RNAs are non-coding RNAs found in the bacterium Staphylococcus aureus. The shared name comes from their discovery, and does not imply homology. Bioinformatics scans identified the 16 Rsa RNA families named RsaA-K and RsaOA-OG. Others, RsaOH-OX, were found thanks to an RNomic approach. Although the RNAs showed varying expression patterns, many of the newly discovered RNAs were shown to be Hfq-independent and most carried a C-rich motif (UCCC).

The gab operon is responsible for the conversion of γ-aminobutyrate (GABA) to succinate. The gab operon comprises three structural genes – gabD, gabT and gabP – that encode for a succinate semialdehyde dehydrogenase, GABA transaminase and a GABA permease respectively. There is a regulatory gene csiR, downstream of the operon, that codes for a putative transcriptional repressor and is activated when nitrogen is limiting.
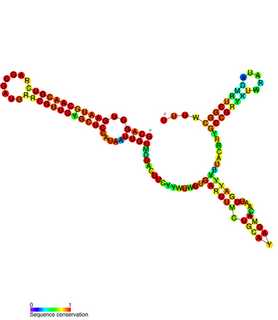
RNA thermometers (RNATs) regulate gene expression in response to temperature, allowing pathogens such as Neisseria meningitidis to switch on silent genes after entering the host organism. However the temperature for expression of Neisseria virulence-associated traits is 42 °C while other bacterial pathogen RNATs require 37 °C. This is probably because N. meningitidis is an obligate commensal of the human nasopharynx and becomes pathogenic during inflammation due to viral infection. Three independent RNA thermosensors were identified in the 5'UTRs of genes needed for: capsule biosynthesis (cssA), the expression of factor H binding protein (fHbp) and sialylation of lipopolysaccharide, which is essential for bacterial resistance against immune killing (lst). The very different nucleotide sequence and predicted inhibitory structures of the three RNATs indicate that they have evolved independently.