Related Research Articles

Bacterial conjugation is the transfer of genetic material between bacterial cells by direct cell-to-cell contact or by a bridge-like connection between two cells. This takes place through a pilus. It is a parasexual mode of reproduction in bacteria.
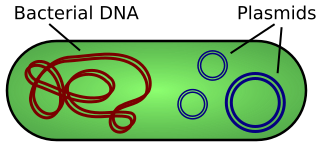
A plasmid is a small, extrachromosomal DNA molecule within a cell that is physically separated from chromosomal DNA and can replicate independently. They are most commonly found as small circular, double-stranded DNA molecules in bacteria; however, plasmids are sometimes present in archaea and eukaryotic organisms. Plasmids often carry useful genes, such as antibiotic resistance and virulence. While chromosomes are large and contain all the essential genetic information for living under normal conditions, plasmids are usually very small and contain additional genes for special circumstances.

FtsZ is a protein encoded by the ftsZ gene that assembles into a ring at the future site of bacterial cell division. FtsZ is a prokaryotic homologue of the eukaryotic protein tubulin. The initials FtsZ mean "Filamenting temperature-sensitive mutant Z." The hypothesis was that cell division mutants of E. coli would grow as filaments due to the inability of the daughter cells to separate from one another. FtsZ is found in almost all bacteria, many archaea, all chloroplasts and some mitochondria, where it is essential for cell division. FtsZ assembles the cytoskeletal scaffold of the Z ring that, along with additional proteins, constricts to divide the cell in two.

Mycobacterium smegmatis is an acid-fast bacterial species in the phylum Actinomycetota and the genus Mycobacterium. It is 3.0 to 5.0 μm long with a bacillus shape and can be stained by Ziehl–Neelsen method and the auramine-rhodamine fluorescent method. It was first reported in November 1884, who found a bacillus with the staining appearance of tubercle bacilli in syphilitic chancres. Subsequent to this, Alvarez and Tavel found organisms similar to that described by Lustgarten also in normal genital secretions (smegma). This organism was later named M. smegmatis.

A tumour inducing (Ti) plasmid is a plasmid found in pathogenic species of Agrobacterium, including A. tumefaciens, A. rhizogenes, A. rubi and A. vitis.

Prokaryotic DNA Replication is the process by which a prokaryote duplicates its DNA into another copy that is passed on to daughter cells. Although it is often studied in the model organism E. coli, other bacteria show many similarities. Replication is bi-directional and originates at a single origin of replication (OriC). It consists of three steps: Initiation, elongation, and termination.
An origin of transfer (oriT) is a short sequence ranging from 40-500 base pairs in length that is necessary for the transfer of DNA from a gram-negative bacterial donor to recipient during bacterial conjugation. The transfer of DNA is a critical component for antimicrobial resistance within bacterial cells and the oriT structure and mechanism within plasmid DNA is complementary to its function in bacterial conjugation. The first oriT to be identified and cloned was on the RK2 (IncP) conjugative plasmid, which was done by Guiney and Helinski in 1979.
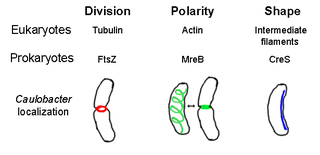
The prokaryotic cytoskeleton is the collective name for all structural filaments in prokaryotes. It was once thought that prokaryotic cells did not possess cytoskeletons, but advances in visualization technology and structure determination led to the discovery of filaments in these cells in the early 1990s. Not only have analogues for all major cytoskeletal proteins in eukaryotes been found in prokaryotes, cytoskeletal proteins with no known eukaryotic homologues have also been discovered. Cytoskeletal elements play essential roles in cell division, protection, shape determination, and polarity determination in various prokaryotes.
Segrosomes are protein complexes that ensure accurate segregation (partitioning) of plasmids or chromosomes during bacterial cell division.

A toxin-antitoxin system consists of a "toxin" and a corresponding "antitoxin", usually encoded by closely linked genes. The toxin is usually a protein while the antitoxin can be a protein or an RNA. Toxin-antitoxin systems are widely distributed in prokaryotes, and organisms often have them in multiple copies. When these systems are contained on plasmids – transferable genetic elements – they ensure that only the daughter cells that inherit the plasmid survive after cell division. If the plasmid is absent in a daughter cell, the unstable antitoxin is degraded and the stable toxic protein kills the new cell; this is known as 'post-segregational killing' (PSK).

In molecular biology the SeqA protein is found in bacteria and archaea. The function of this protein is highly important in DNA replication. The protein negatively regulates the initiation of DNA replication at the origin of replication, in Escherichia coli, OriC. Additionally the protein plays a further role in sequestration. The importance of this protein is vital, without its help in DNA replication, cell division and other crucial processes could not occur. This protein domain is thought to be part of a much larger protein complex which includes other proteins such as SeqB.
The archaellum is a unique structure on the cell surface of many archaea that allows for swimming motility. The archaellum consists of a rigid helical filament that is attached to the cell membrane by a molecular motor. This molecular motor – composed of cytosolic, membrane, and pseudo-periplasmic proteins – is responsible for the assembly of the filament and, once assembled, for its rotation. The rotation of the filament propels archaeal cells in liquid medium, in a manner similar to the propeller of a boat. The bacterial analog of the archaellum is the flagellum, which is also responsible for their swimming motility and can also be compared to a rotating corkscrew. Although the movement of archaella and flagella is sometimes described as "whip-like", this is incorrect, as only cilia from Eukaryotes move in this manner. Indeed, even "flagellum" is a misnomer, as bacterial flagella also work as propeller-like structures.
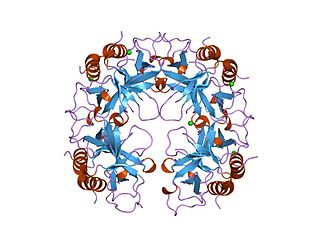
The CcdA/CcdB Type II Toxin-antitoxin system is one example of the bacterial toxin-antitoxin (TA) systems that encode two proteins, one a potent inhibitor of cell proliferation (toxin) and the other its specific antidote (antitoxin). These systems preferentially guarantee growth of plasmid-carrying daughter cells in a bacterial population by killing newborn bacteria that have not inherited a plasmid copy at cell division.
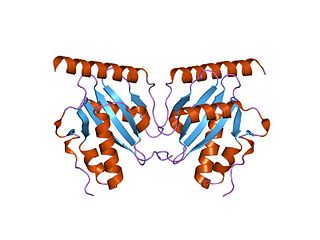
The parDE type II toxin-antitoxin system is one example of the bacterial toxin-antitoxin (TA) systems that encode two proteins, one a potent inhibitor of cell proliferation (toxin) and the other its specific antidote (antitoxin). These systems preferentially guarantee growth of plasmid-carrying daughter cells in a bacterial population by killing newborn bacteria that have not inherited a plasmid copy at cell division.

The Min System is a mechanism composed of three proteins MinC, MinD, and MinE used by E. coli as a means of properly localizing the septum prior to cell division. Each component participates in generating a dynamic oscillation of FtsZ protein inhibition between the two bacterial poles to precisely specify the mid-zone of the cell, allowing the cell to accurately divide in two. This system is known to function in conjunction with a second negative regulatory system, the nucleoid occlusion system (NO), to ensure proper spatial and temporal regulation of chromosomal segregation and division.
The MinE protein is one of three proteins of the Min system encoded by the minB operon required to generate pole to pole oscillations prior to bacterial cell division as a means of specifying the midzone of the cell, as seen in E.coli.
A plasmid partition system is a mechanism that ensures the stable inheritance of plasmids during bacterial cell division. Each plasmid has its independent replication system which controls the number of copies of the plasmid in a cell. The higher the copy number, the more likely the two daughter cells will contain the plasmid. Generally, each molecule of plasmid diffuses randomly, so the probability of having a plasmid-less daughter cell is 21−N, where N is the number of copies. For instance, if there are 2 copies of a plasmid in a cell, there is 50% chance of having one plasmid-less daughter cell. However, high-copy number plasmids have a cost for the hosting cell. This metabolic burden is lower for low-copy plasmids, but those have a higher probability of plasmid loss after a few generations. To control vertical transmission of plasmids, in addition to controlled-replication systems, bacterial plasmids use different maintenance strategies, such as multimer resolution systems, post-segregational killing systems, and partition systems.
The ParMRC system is a mechanism for sorting DNA plasmids to opposite ends of a bacterial cell during cell division. It has three components: ParM, an actin-like protein that forms a long filament to push two plasmids apart, ParR, which binds the plasmid to ParM and generates the ParM filament, and parC, which is a DNA sequence on the plasmid that anchors ParR to itself.
The locus of enterocyte effacement-encoded regulator (Ler) is a regulatory protein that controls bacterial pathogenicity of enteropathogenic Escherichia coli (EPEC) and enterohemorrhagic Escherichia coli (EHEC). More specifically, Ler regulates the locus of enterocyte effacement (LEE) pathogenicity island genes, which are responsible for creating intestinal attachment and effacing lesions and subsequent diarrhea: LEE1, LEE2, and LEE3. LEE1, 2, and 3 carry the information necessary for a type III secretion system. The transcript encoding the Ler protein is the open reading frame 1 on the LEE1 operon.
FtsK is a protein in E.Coli involved in bacterial cell division and chromosome segregation. It is one of the largest proteins, consisting of 1329 amino acids. FtsK stands for "Filament temperature sensitive mutant K" because cells expressing a mutant ftsK allele called ftsK44, which encodes an FtsK variant containing an G80A residue change in the second transmembrane segment, fail to divide at high temperatures and form long filaments instead. FtsK, specifically its C-terminal domain, functions as a DNA translocase, interacts with other cell division proteins, and regulates Xer-mediated recombination. FtsK belongs to the AAA superfamily and is present in most bacteria.
References
- ↑ Surtees, JA; Funnell, BE (2003). Plasmid and chromosome traffic control: how ParA and ParB drive partition. Vol. 56. pp. 145–80. doi:10.1016/s0070-2153(03)01010-x. ISBN 9780121531560. PMID 14584729.
{{cite book}}:|journal=ignored (help) - ↑ Rodionov, O; Lobocka, M; Yarmolinsky, M (Jan 22, 1999). "Silencing of genes flanking the P1 plasmid centromere". Science. 283 (5401): 546–9. Bibcode:1999Sci...283..546R. doi:10.1126/science.283.5401.546. PMID 9915704.
- ↑ Murray, H; Ferreira, H; Errington, J (Sep 2006). "The bacterial chromosome segregation protein Spo0J spreads along DNA from parS nucleation sites". Molecular Microbiology. 61 (5): 1352–61. doi:10.1111/j.1365-2958.2006.05316.x. hdl: 11449/701 . PMID 16925562. S2CID 17530813.
- ↑ Breier, AM; Grossman, AD (May 2007). "Whole-genome analysis of the chromosome partitioning and sporulation protein Spo0J (ParB) reveals spreading and origin-distal sites on the Bacillus subtilis chromosome". Molecular Microbiology. 64 (3): 703–18. doi: 10.1111/j.1365-2958.2007.05690.x . PMID 17462018.
- ↑ Sanchez, A; Cattoni, DI; Walter, JC; Rech, J; Parmeggiani, A; Nollmann, M; Bouet, JY (2015). "Stochastic Self-Assembly of ParB Proteins Builds the Bacterial DNA Segregation Apparatus". Cell Systems. 1 (2): 163–73. doi: 10.1016/j.cels.2015.07.013 . PMID 27135801.
- ↑ Bouet, JY; Ah-Seng, Y; Benmeradi, N; Lane, D (2007). "Polymerization of SopA partition ATPase: regulation by DNA binding and SopB". Molecular Microbiology. 63 (2): 468–81. doi: 10.1111/j.1365-2958.2006.05537.x . PMID 17166176.
- ↑ Castaing, JP; Bouet, JY; Lane, D (2008). "F plasmid partition depends on interaction of SopA with non-specific DNA". Molecular Microbiology. 70 (4): 1000–11. doi: 10.1111/j.1365-2958.2008.06465.x . PMID 18826408. S2CID 26612131.
- ↑ Bouet, JY; Funnell, BE (1999). "P1 ParA interacts with the P1 partition complex at parS and an ATP-ADP switch controls ParA activities". EMBO J. 18 (5): 1415–24. doi:10.1093/emboj/18.5.1415. PMC 1171231 . PMID 10064607.
- ↑ Walter, JC; Dorignac, J; Lorman, V; Rech, J; Bouet, JY; Nollmann, M; Palmeri, J; Parmeggiani, A; Geniet, F (2017). "Surfing on protein waves: proteophoresis as a mechanism for bacterial genome partitioning". Physical Review Letters. 119 (28101): 028101. arXiv: 1702.07372 . Bibcode:2017PhRvL.119b8101W. doi:10.1103/PhysRevLett.119.028101. PMID 28753349. S2CID 6762277.
- ↑ Ptacin, JL; Lee, SF; Garner, EC; Toro, E; Eckart, M; Comolli, LR; Moerner, WE; Shapiro, L (Aug 2010). "A spindle-like apparatus guides bacterial chromosome segregation". Nature Cell Biology. 12 (8): 791–8. doi:10.1038/ncb2083. PMC 3205914 . PMID 20657594.
- ↑ Ringgaard, S; van Zon, J; Howard, M; Gerdes, K (Nov 17, 2009). "Movement and equipositioning of plasmids by ParA filament disassembly". Proceedings of the National Academy of Sciences of the United States of America. 106 (46): 19369–74. Bibcode:2009PNAS..10619369R. doi: 10.1073/pnas.0908347106 . PMC 2775997 . PMID 19906997.
- ↑ Hwang, LC; Vecchiarelli, AG; Han, YW; Mizuuchi, M; Harada, Y; Funnell, BE; Mizuuchi, K (May 2, 2013). "ParA-mediated plasmid partition driven by protein pattern self-organization". The EMBO Journal. 32 (9): 1238–49. doi:10.1038/emboj.2013.34. PMC 3642677 . PMID 23443047.
- ↑ Vecchiarelli, AG; Hwang, LC; Mizuuchi, K (Apr 9, 2013). "Cell-free study of F plasmid partition provides evidence for cargo transport by a diffusion-ratchet mechanism". Proceedings of the National Academy of Sciences of the United States of America. 110 (15): E1390–7. Bibcode:2013PNAS..110E1390V. doi: 10.1073/pnas.1302745110 . PMC 3625265 . PMID 23479605.