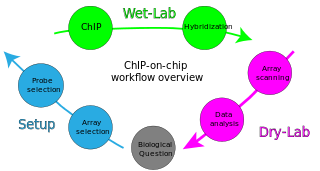
ChIP-on-chip is a technology that combines chromatin immunoprecipitation ('ChIP') with DNA microarray ("chip"). Like regular ChIP, ChIP-on-chip is used to investigate interactions between proteins and DNA in vivo. Specifically, it allows the identification of the cistrome, the sum of binding sites, for DNA-binding proteins on a genome-wide basis. Whole-genome analysis can be performed to determine the locations of binding sites for almost any protein of interest. As the name of the technique suggests, such proteins are generally those operating in the context of chromatin. The most prominent representatives of this class are transcription factors, replication-related proteins, like origin recognition complex protein (ORC), histones, their variants, and histone modifications.
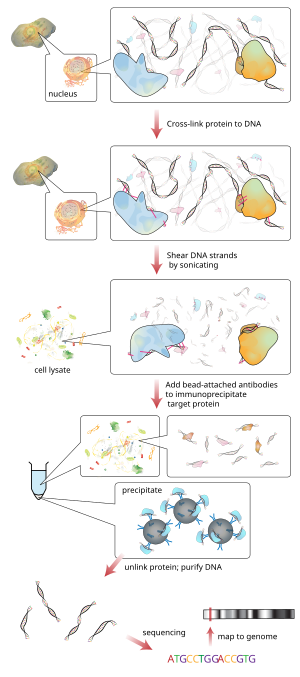
ChIP-sequencing, also known as ChIP-seq, is a method used to analyze protein interactions with DNA. ChIP-seq combines chromatin immunoprecipitation (ChIP) with massively parallel DNA sequencing to identify the binding sites of DNA-associated proteins. It can be used to map global binding sites precisely for any protein of interest. Previously, ChIP-on-chip was the most common technique utilized to study these protein–DNA relations.
Epigenomics is the study of the complete set of epigenetic modifications on the genetic material of a cell, known as the epigenome. The field is analogous to genomics and proteomics, which are the study of the genome and proteome of a cell. Epigenetic modifications are reversible modifications on a cell's DNA or histones that affect gene expression without altering the DNA sequence. Epigenomic maintenance is a continuous process and plays an important role in stability of eukaryotic genomes by taking part in crucial biological mechanisms like DNA repair. Plant flavones are said to be inhibiting epigenomic marks that cause cancers. Two of the most characterized epigenetic modifications are DNA methylation and histone modification. Epigenetic modifications play an important role in gene expression and regulation, and are involved in numerous cellular processes such as in differentiation/development and tumorigenesis. The study of epigenetics on a global level has been made possible only recently through the adaptation of genomic high-throughput assays.

ChIP-exo is a chromatin immunoprecipitation based method for mapping the locations at which a protein of interest binds to the genome. It is a modification of the ChIP-seq protocol, improving the resolution of binding sites from hundreds of base pairs to almost one base pair. It employs the use of exonucleases to degrade strands of the protein-bound DNA in the 5'-3' direction to within a small number of nucleotides of the protein binding site. The nucleotides of the exonuclease-treated ends are determined using some combination of DNA sequencing, microarrays, and PCR. These sequences are then mapped to the genome to identify the locations on the genome at which the protein binds.
H3K27ac is an epigenetic modification to the DNA packaging protein histone H3. It is a mark that indicates acetylation of the lysine residue at N-terminal position 27 of the histone H3 protein.
H3K9me3 is an epigenetic modification to the DNA packaging protein Histone H3. It is a mark that indicates the tri-methylation at the 9th lysine residue of the histone H3 protein and is often associated with heterochromatin.
H3K4me1 is an epigenetic modification to the DNA packaging protein Histone H3. It is a mark that indicates the mono-methylation at the 4th lysine residue of the histone H3 protein and often associated with gene enhancers.
H3K79me2 is an epigenetic modification to the DNA packaging protein Histone H3. It is a mark that indicates the di-methylation at the 79th lysine residue of the histone H3 protein. H3K79me2 is detected in the transcribed regions of active genes.
H4K5ac is an epigenetic modification to the DNA packaging protein histone H4. It is a mark that indicates the acetylation at the 5th lysine residue of the histone H4 protein. H4K5 is the closest lysine residue to the N-terminal tail of histone H4. It is enriched at the transcription start site (TSS) and along gene bodies. Acetylation of histone H4K5 and H4K12ac is enriched at centromeres.
H4K8ac, representing an epigenetic modification to the DNA packaging protein histone H4, is a mark indicating the acetylation at the 8th lysine residue of the histone H4 protein. It has been implicated in the prevalence of malaria.
H4K12ac is an epigenetic modification to the DNA packaging protein histone H4. It is a mark that indicates the acetylation at the 12th lysine residue of the histone H4 protein. H4K12ac is involved in learning and memory. It is possible that restoring this modification could reduce age-related decline in memory.
H3K14ac is an epigenetic modification to the DNA packaging protein Histone H3. It is a mark that indicates the acetylation at the 14th lysine residue of the histone H3 protein.
H3K9ac is an epigenetic modification to the DNA packaging protein Histone H3. It is a mark that indicates the acetylation at the 9th lysine residue of the histone H3 protein.
H3K36me2 is an epigenetic modification to the DNA packaging protein Histone H3. It is a mark that indicates the di-methylation at the 36th lysine residue of the histone H3 protein.

MNase-seq, short for micrococcal nuclease digestion with deep sequencing, is a molecular biological technique that was first pioneered in 2006 to measure nucleosome occupancy in the C. elegans genome, and was subsequently applied to the human genome in 2008. Though, the term ‘MNase-seq’ had not been coined until a year later, in 2009. Briefly, this technique relies on the use of the non-specific endo-exonuclease micrococcal nuclease, an enzyme derived from the bacteria Staphylococcus aureus, to bind and cleave protein-unbound regions of DNA on chromatin. DNA bound to histones or other chromatin-bound proteins may remain undigested. The uncut DNA is then purified from the proteins and sequenced through one or more of the various Next-Generation sequencing methods.
H3R17me2 is an epigenetic modification to the DNA packaging protein histone H3. It is a mark that indicates the di-methylation at the 17th arginine residue of the histone H3 protein. In epigenetics, arginine methylation of histones H3 and H4 is associated with a more accessible chromatin structure and thus higher levels of transcription. The existence of arginine demethylases that could reverse arginine methylation is controversial.
H3R26me2 is an epigenetic modification to the DNA packaging protein histone H3. It is a mark that indicates the di-methylation at the 26th arginine residue of the histone H3 protein. In epigenetics, arginine methylation of histones H3 and H4 is associated with a more accessible chromatin structure and thus higher levels of transcription. The existence of arginine demethylases that could reverse arginine methylation is controversial.
H3R8me2 is an epigenetic modification to the DNA packaging protein histone H3. It is a mark that indicates the di-methylation at the 8th arginine residue of the histone H3 protein. In epigenetics, arginine methylation of histones H3 and H4 is associated with a more accessible chromatin structure and thus higher levels of transcription. The existence of arginine demethylases that could reverse arginine methylation is controversial.
H3R2me2 is an epigenetic modification to the DNA packaging protein histone H3. It is a mark that indicates the di-methylation at the 2nd arginine residue of the histone H3 protein. In epigenetics, arginine methylation of histones H3 and H4 is associated with a more accessible chromatin structure and thus higher levels of transcription. The existence of arginine demethylases that could reverse arginine methylation is controversial.
H3T45P is an epigenetic modification to the DNA packaging protein histone H3. It is a mark that indicates the phosphorylation the 45th threonine residue of the histone H3 protein.