Related Research Articles

Proteasomes are protein complexes which degrade unneeded or damaged proteins by proteolysis, a chemical reaction that breaks peptide bonds. Enzymes that help such reactions are called proteases.
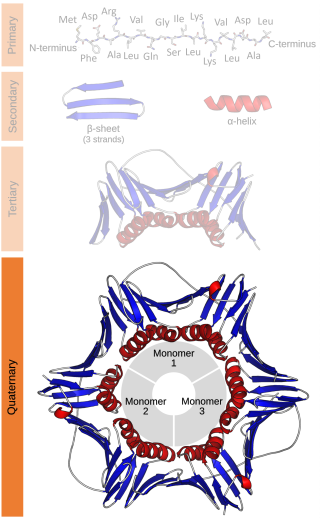
Protein quaternary structure is the fourth classification level of protein structure. Protein quaternary structure refers to the structure of proteins which are themselves composed of two or more smaller protein chains. Protein quaternary structure describes the number and arrangement of multiple folded protein subunits in a multi-subunit complex. It includes organizations from simple dimers to large homooligomers and complexes with defined or variable numbers of subunits. In contrast to the first three levels of protein structure, not all proteins will have a quaternary structure since some proteins function as single units. Protein quaternary structure can also refer to biomolecular complexes of proteins with nucleic acids and other cofactors.

ATPases (EC 3.6.1.3, Adenosine 5'-TriPhosphatase, adenylpyrophosphatase, ATP monophosphatase, triphosphatase, SV40 T-antigen, ATP hydrolase, complex V (mitochondrial electron transport), (Ca2+ + Mg2+)-ATPase, HCO3−-ATPase, adenosine triphosphatase) are a class of enzymes that catalyze the decomposition of ATP into ADP and a free phosphate ion or the inverse reaction. This dephosphorylation reaction releases energy, which the enzyme (in most cases) harnesses to drive other chemical reactions that would not otherwise occur. This process is widely used in all known forms of life.

ATP synthase is an enzyme that catalyzes the formation of the energy storage molecule adenosine triphosphate (ATP) using adenosine diphosphate (ADP) and inorganic phosphate (Pi). ATP synthase is a molecular machine. The overall reaction catalyzed by ATP synthase is:

In structural biology, a protein subunit is a polypeptide chain or single protein molecule that assembles with others to form a protein complex. Large assemblies of proteins such as viruses often use a small number of types of protein subunits as building blocks.
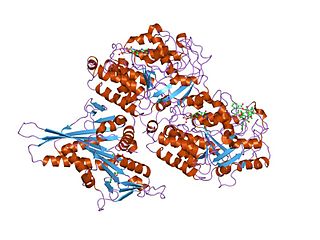
Tubulin in molecular biology can refer either to the tubulin protein superfamily of globular proteins, or one of the member proteins of that superfamily. α- and β-tubulins polymerize into microtubules, a major component of the eukaryotic cytoskeleton. Microtubules function in many essential cellular processes, including mitosis. Tubulin-binding drugs kill cancerous cells by inhibiting microtubule dynamics, which are required for DNA segregation and therefore cell division.
Phycoerythrin (PE) is a red protein-pigment complex from the light-harvesting phycobiliprotein family, present in cyanobacteria, red algae and cryptophytes, accessory to the main chlorophyll pigments responsible for photosynthesis.The red pigment is due to the prosthetic group, phycoerythrobilin, which gives phycoerythrin its red color.

In chemistry and biochemistry, an oligomer is a molecule that consists of a few repeating units which could be derived, actually or conceptually, from smaller molecules, monomers. The name is composed of Greek elements oligo-, "a few" and -mer, "parts". An adjective form is oligomeric.

Protein structure is the three-dimensional arrangement of atoms in an amino acid-chain molecule. Proteins are polymers – specifically polypeptides – formed from sequences of amino acids, which are the monomers of the polymer. A single amino acid monomer may also be called a residue, which indicates a repeating unit of a polymer. Proteins form by amino acids undergoing condensation reactions, in which the amino acids lose one water molecule per reaction in order to attach to one another with a peptide bond. By convention, a chain under 30 amino acids is often identified as a peptide, rather than a protein. To be able to perform their biological function, proteins fold into one or more specific spatial conformations driven by a number of non-covalent interactions, such as hydrogen bonding, ionic interactions, Van der Waals forces, and hydrophobic packing. To understand the functions of proteins at a molecular level, it is often necessary to determine their three-dimensional structure. This is the topic of the scientific field of structural biology, which employs techniques such as X-ray crystallography, NMR spectroscopy, cryo-electron microscopy (cryo-EM) and dual polarisation interferometry, to determine the structure of proteins.
Aromatic-ring-hydroxylating dioxygenases (ARHD) incorporate two atoms of dioxygen (O2) into their substrates in the dihydroxylation reaction. The product is (substituted) cis-1,2-dihydroxycyclohexadiene, which is subsequently converted to (substituted) benzene glycol by a cis-diol dehydrogenase.
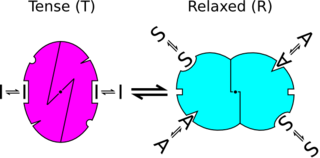
In biochemistry, the Monod–Wyman–Changeux model describes allosteric transitions of proteins made up of identical subunits. It was proposed by Jean-Pierre Changeux in his PhD thesis, and described by Jacques Monod, Jeffries Wyman, and Jean-Pierre Changeux. It contrasts with the sequential model and substrate presentation.

Succinyl coenzyme A synthetase is an enzyme that catalyzes the reversible reaction of succinyl-CoA to succinate. The enzyme facilitates the coupling of this reaction to the formation of a nucleoside triphosphate molecule from an inorganic phosphate molecule and a nucleoside diphosphate molecule. It plays a key role as one of the catalysts involved in the citric acid cycle, a central pathway in cellular metabolism, and it is located within the mitochondrial matrix of a cell.
Gastric hydrogen potassium ATPase, also known as H+/K+ ATPase, is an enzyme which functions to acidify the stomach. It is a member of the P-type ATPases, also known as E1-E2 ATPases due to its two states.

Bisphosphoglycerate mutase is an enzyme expressed in erythrocytes and placental cells. It is responsible for the catalytic synthesis of 2,3-Bisphosphoglycerate (2,3-BPG) from 1,3-bisphosphoglycerate. BPGM also has a mutase and a phosphatase function, but these are much less active, in contrast to its glycolytic cousin, phosphoglycerate mutase (PGM), which favors these two functions, but can also catalyze the synthesis of 2,3-BPG to a lesser extent.

Hexosaminidase is an enzyme involved in the hydrolysis of terminal N-acetyl-D-hexosamine residues in N-acetyl-β-D-hexosaminides.

Phosphorylase kinase (PhK) is a serine/threonine-specific protein kinase which activates glycogen phosphorylase to release glucose-1-phosphate from glycogen. PhK phosphorylates glycogen phosphorylase at two serine residues, triggering a conformational shift which favors the more active glycogen phosphorylase “a” form over the less active glycogen phosphorylase b.
Nitrile hydratases are mononuclear iron or non-corrinoid cobalt enzymes that catalyse the hydration of diverse nitriles to their corresponding amides:

The enzyme anthranilate synthase catalyzes the chemical reaction

Fatty-acyl-CoA Synthase, or more commonly known as yeast fatty acid synthase, is an enzyme complex responsible for fatty acid biosynthesis, and is of Type I Fatty Acid Synthesis (FAS). Yeast fatty acid synthase plays a pivotal role in fatty acid synthesis. It is a 2.6 MDa barrel shaped complex and is composed of two, unique multi-functional subunits: alpha and beta. Together, the alpha and beta units are arranged in an α6β6 structure. The catalytic activities of this enzyme complex involves a coordination system of enzymatic reactions between the alpha and beta subunits. The enzyme complex therefore consists of six functional centers for fatty acid synthesis.
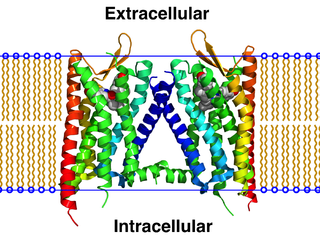
A GPCR oligomer is a protein complex that consists of a small number of G protein-coupled receptors (GPCRs). It is held together by covalent bonds or by intermolecular forces. The subunits within this complex are called protomers, while unconnected receptors are called monomers. Receptor homomers consist of identical protomers, while heteromers consist of different protomers.
References
- ↑ Chetverin, A.B. (1986). "Evidence for a diprotomeric structure of Na, K-ATPase: Accurate determination of protein concentration and quantitative end-group analysis". FEBS Lett. 196 (1): 121–125. doi: 10.1016/0014-5793(86)80225-3 . PMID 3002859.
- ↑ P. M. Lalli, B. A. Iglesias, H. E. Toma, G. F. de Sa, R. J. Daroda, J. C. Silva Filho, J. E. Szulejko, K. Araki and M. N. Eberlin, J. Mass Spectrom., 2012, 47, 712–719.
- ↑ C. Lapthorn, T. J. Dines, B. Z. Chowdhry, G. L. Perkins and F. S. Pullen, Rapid Commun. Mass Spectrom., 2013, 27, 2399–2410.
- ↑ Buxbaum, E. (2007). Fundamentals of protein structure and function. New York: Springer. pp. 105–120. ISBN 978-0-387-26352-6.
- ↑ J. Am. Chem. Soc., 2009, 131 (3), pp 1174–1181
- ↑ J. Phys. Chem. A, 2011, 115 (26), pp 7625–7632