A regulatory sequence is a segment of a nucleic acid molecule which is capable of increasing or decreasing the expression of specific genes within an organism. Regulation of gene expression is an essential feature of all living organisms and viruses.

The Argonaute protein family, first discovered for its evolutionarily conserved stem cell function, plays a central role in RNA silencing processes as essential components of the RNA-induced silencing complex (RISC). RISC is responsible for the gene silencing phenomenon known as RNA interference (RNAi). Argonaute proteins bind different classes of small non-coding RNAs, including microRNAs (miRNAs), small interfering RNAs (siRNAs) and Piwi-interacting RNAs (piRNAs). Small RNAs guide Argonaute proteins to their specific targets through sequence complementarity, which then leads to mRNA cleavage, translation inhibition, and/or the initiation of mRNA decay.

In cell biology, a paraspeckle is an irregularly shaped compartment of the cell, approximately 0.2-1 μm in size, found in the nucleus' interchromatin space. First documented in HeLa cells, where there are generally 10-30 per nucleus, Paraspeckles are now known to also exist in all human primary cells, transformed cell lines and tissue sections. Their name is derived from their distribution in the nucleus; the "para" is short for parallel and the "speckle" refers to the splicing speckles to which they are always in close proximity. Their function is still not fully understood, but they are thought to regulate gene expression by sequestrating proteins or mRNAs with inverted repeats in their 3′ UTRs.
In the field of molecular biology the 6S RNA is a non-coding RNA that was one of the first to be identified and sequenced. What it does in the bacterial cell was unknown until recently. In the early 2000s scientists found out the function of 6S RNA to be as a regulator of sigma 70-dependent gene transcription. All bacterial RNA polymerases have a subunit called a sigma factor. The sigma factors are important because they control how DNA promoter binding and RNA transcription start sites. Sigma 70 was the first one to be discovered in Escherichia coli.

In molecular biology, SNORD115 is a non-coding RNA (ncRNA) molecule known as a small nucleolar RNA which usually functions in guiding the modification of other non-coding RNAs. This type of modifying RNA is usually located in the nucleolus of the eukaryotic cell which is a major site of snRNA biogenesis. HBII-52 refers to the human gene, whereas RBII-52 is used for the rat gene and MBII-52 is used for naming the mouse gene.

The ykkC/yxkD leader is a conserved RNA structure found upstream of the ykkC and yxkD genes in Bacillus subtilis and related genes in other bacteria. The function of this family is unclear for many years although it has been suggested that it may function to switch on efflux pumps and detoxification systems in response to harmful environmental molecules. The Thermoanaerobacter tengcongensis sequence AE013027 overlaps with that of purine riboswitch suggesting that the two riboswitches may work in conjunction to regulate the upstream gene which codes for TTE0584 (Q8RC62), a member of the permease family.

Eukaryotic initiation factor 4A-III is a protein that in humans is encoded by the EIF4A3 gene.

Putative RNA-binding protein 15 is a protein that in humans is encoded by the RBM15 gene. It is an RNA-binding protein that acts as a key regulator of N6-Methyladenosine (m6A) methylation of RNAs

DNA-directed RNA polymerase, mitochondrial is an enzyme that in humans is encoded by the POLRMT gene.

The sucA RNA motif is a conserved RNA structure found in bacteria of the order Burkholderiales. RNAs within this motif are always found in the presumed 5' UTR of sucA genes. sucA encodes a subunit of an enzyme that participates in the citric acid cycle by synthesizing succinyl-CoA from 2-oxoglutarate. A part of the conserved structure overlaps predicted Shine-Dalgarno sequences of the downstream sucA genes. Because of the RNA motif's consistent gene association and a possible mechanism for sequestering the ribosome binding site, it was proposed that the sucA RNA motif corresponds to a cis-regulatory element. Its relatively complex secondary structure could indicate that it is a riboswitch. However, the function of this RNA motif remains unknown.

The ATPC RNA motif is a conserved RNA structure found in certain cyanobacteria. It is apparently ubiquitous in Prochlorococcus marinus, and is present in many species in the genus Synechococcus. The RNA is always found within an operon encoding subunits of ATP synthase, and it is always located downstream of the gene encoding the A subunit of ATP synthase, and upstream of the C subunit gene. This location is consistent with a cis-regulatory element, but also with a non-coding RNA that is transcribed with the ATP synthase genes.

The fluoride riboswitch is a conserved RNA structure identified by bioinformatics in a wide variety of bacteria and archaea. These RNAs were later shown to function as riboswitches that sense fluoride ions. These "fluoride riboswitches" increase expression of downstream genes when fluoride levels are elevated, and the genes are proposed to help mitigate the toxic effects of very high levels of fluoride.

The glutamine riboswitch is a conserved RNA structure that was predicted by bioinformatics. It is present in a variety of lineages of cyanobacteria, as well as some phages that infect cyanobacteria. It is also found in DNA extracted from uncultivated bacteria living in the ocean that are presumably species of cyanobacteria.

The gyrA RNA motif is a conserved RNA structure identified by bioinformatics. The RNAs are present in multiple species of bacteria within the order Pseudomonadales. This order contains the genus Pseudomonas, which includes the opportunistic human pathogen Pseudomonas aeruginosa and Pseudomonas syringae, a plant pathogen.
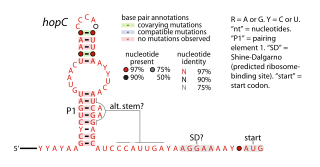
The hopC RNA motif is a predicted cis-regulatory element identified by a bioinformatic screen for conserved RNA secondary structures. hopC RNAs are exclusively found within bacteria classified within the genus Helicobacter, some of which are human pathogens that infect the stomach and can cause ulcers.

The manA RNA motif refers to a conserved RNA structure that was identified by bioinformatics. Instances of the manA RNA motif were detected in bacteria in the genus Photobacterium and phages that infect certain kinds of cyanobacteria. However, most predicted manA RNA sequences are derived from DNA collected from uncultivated marine bacteria. Almost all manA RNAs are positioned such that they might be in the 5' untranslated regions of protein-coding genes, and therefore it was hypothesized that manA RNAs function as cis-regulatory elements. Given the relative complexity of their secondary structure, and their hypothesized cis-regulatory role, they might be riboswitches.
The wcaG RNA motif is an RNA structure conserved in some bacteria that was detected by bioinformatics. wcaG RNAs are found in certain phages that infect cyanobacteria. Most known wcaG RNAs were found in sequences of DNA extracted from uncultivated marine bacteria. wcaG RNAs might function as cis-regulatory elements, in view of their consistent location in the possible 5' untranslated regions of genes. It was suggested the wcaG RNAs might further function as riboswitches.

The pfl RNA motif refers to a conserved RNA structure present in some bacteria and originally discovered using bioinformatics. pfl RNAs are consistently present in genomic locations that likely correspond to the 5' untranslated regions of protein-coding genes. This arrangement in bacteria is commonly associated with cis-regulatory elements. Moreover, they are in presumed 5' UTRs of multiple non-homologous genes, suggesting that they function only in these locations. Additional evidence of cis-regulatory function came from the observation that predicted rho-independent transcription terminators overlap pfl RNAs. This overlap suggests that the alternate secondary structures of pfl RNA and the transcription terminator stem-loops compete with each other, and this is a common mechanism for cis gene control in bacteria.

N6-Methyladenosine (m6A) was originally identified and partially characterised in the 1970s, and is an abundant modification in mRNA and DNA. It is found within some viruses, and most eukaryotes including mammals, insects, plants and yeast. It is also found in tRNA, rRNA, and small nuclear RNA (snRNA) as well as several long non-coding RNA, such as Xist.
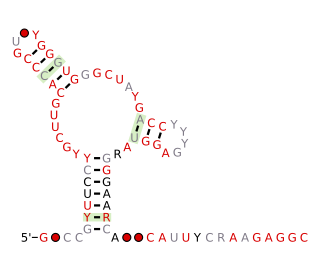
The Fibro-purF RNA motif is a conserved RNA structure that was discovered by bioinformatics. Fibro-purF motif RNAs are found in Fibrobacterota, a group of bacteria that are common in cow rumen. Additionally, the RNAs are found in metagenomic sequences of DNA isolated from cow rumen.

















