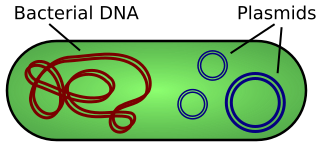
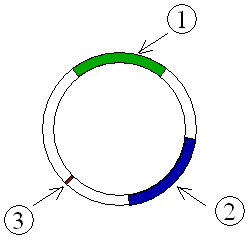
A plasmid is a small, extrachromosomal DNA molecule within a cell that is physically separated from chromosomal DNA and can replicate independently. They are most commonly found as small circular, double-stranded DNA molecules in bacteria; however, plasmids are sometimes present in archaea and eukaryotic organisms. In nature, plasmids often carry genes that benefit the survival of the organism and confer selective advantage such as antibiotic resistance. While chromosomes are large and contain all the essential genetic information for living under normal conditions, plasmids are usually very small and contain only additional genes that may be useful in certain situations or conditions. Artificial plasmids are widely used as vectors in molecular cloning, serving to drive the replication of recombinant DNA sequences within host organisms. In the laboratory, plasmids may be introduced into a cell via transformation.
A bacterial artificial chromosome (BAC) is a DNA construct, based on a functional fertility plasmid, used for transforming and cloning in bacteria, usually E. coli. F-plasmids play a crucial role because they contain partition genes that promote the even distribution of plasmids after bacterial cell division. The bacterial artificial chromosome's usual insert size is 150–350 kbp. A similar cloning vector called a PAC has also been produced from the DNA of P1 bacteriophage.
An inverted repeat is a single stranded sequence of nucleotides followed downstream by its reverse complement. The intervening sequence of nucleotides between the initial sequence and the reverse complement can be any length including zero. When the intervening length is zero, the composite sequence is a palindromic sequence. For example, 5'---TTACGnnnnnnCGTAA---3' is an inverted repeat sequence.

Transduction is the process by which foreign DNA is introduced into a cell by a virus or viral vector. An example is the viral transfer of DNA from one bacterium to another and hence an example of horizontal gene transfer. Transduction does not require physical contact between the cell donating the DNA and the cell receiving the DNA, and it is DNase resistant. Transduction is a common tool used by molecular biologists to stably introduce a foreign gene into a host cell's genome.

Antisense RNA (asRNA), also referred to as antisense transcript, natural antisense transcript (NAT) or antisense oligonucleotide, is a single stranded RNA that is complementary to a protein coding messenger RNA (mRNA) with which it hybridizes, and thereby blocks its translation into protein. asRNAs have been found in both prokaryotes and eukaryotes, antisense transcripts can be classified into short and long non-coding RNAs (ncRNAs). The primary function of asRNA is regulating gene expression. asRNAs may also be produced synthetically and have found wide spread use as research tools for gene knockdown. They may also have therapeutic applications.

A tumour inducing (Ti) plasmid is a plasmid found in pathogenic species of Agrobacterium, including A. tumefaciens, A. rhizogenes, A. rubi and A. vitis.

pBR322 is a plasmid and was one of the first widely used E. coli cloning vectors. Created in 1977 in the laboratory of Herbert Boyer at the University of California, San Francisco, it was named after Francisco Bolivar Zapata, the postdoctoral researcher who constructed it. The p stands for "plasmid," and BR for "Bolivar" and "Rodriguez."
Cis-regulatory elements (CREs) are regions of non-coding DNA which regulate the transcription of neighboring genes. CREs are vital components of genetic regulatory networks, which in turn control morphogenesis, the development of anatomy, and other aspects of embryonic development, studied in evolutionary developmental biology.

In biology, a gene is a sequence of nucleotides in DNA or RNA that encodes the synthesis of a gene product, either RNA or protein.
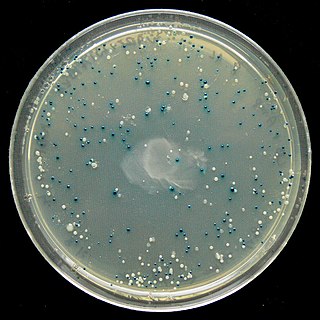
The blue–white screen is a screening technique that allows for the rapid and convenient detection of recombinant bacteria in vector-based molecular cloning experiments. DNA of interest is ligated into a vector. The vector is then inserted into a competent host cell viable for transformation, which are then grown in the presence of X-gal. Cells transformed with vectors containing recombinant DNA will produce white colonies; cells transformed with non-recombinant plasmids grow into blue colonies. This method of screening is usually performed using a suitable bacterial strain, but other organisms such as yeast may also be used.
fis is an E. coli gene encoding the Fis protein. The regulation of this gene is more complex than most other genes in the E. coli genome, as Fis is an important protein which regulates expression of other genes. It is supposed that fis is regulated by H-NS, IHF and CRP. It also regulates its own expression (autoregulation). Fis is one of the most abundant DNA binding proteins in Escherichia coli under nutrient-rich growth conditions.
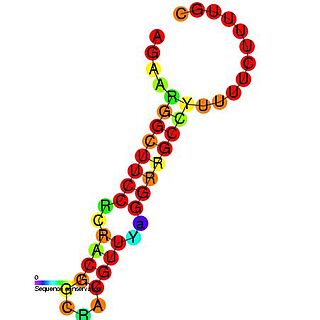
In molecular biology ctRNA is a plasmid encoded noncoding RNA that binds to the mRNA of repB and causes translational inhibition. ctRNA is encoded by plasmids and functions in rolling circle replication to maintain a low copy number. In Corynebacterium glutamicum, it achieves this by antisense pairing with the mRNA of RepB, a replication initiation protein. In Enterococcus faecium the plasmid pJB01 contains three open reading frames, copA, repB, and repC. The pJB01 ctRNA is coded on the opposite strand from the copA/repB intergenic region and partially overlaps an atypical ribosome binding site for repB.
Iterons are directly repeated DNA sequences which play an important role in regulation of plasmid copy number in bacterial cells. It is one among the three negative regulatory elements found in plasmids which control its copy number. The others include antisense RNAs and ctRNAs. Iterons complex with cognate replication (Rep) initiator proteins to achieve the required regulatory effect.
In molecular cloning, a vector is a DNA molecule used as a vehicle to artificially carry foreign genetic material into another cell, where it can be replicated and/or expressed. A vector containing foreign DNA is termed recombinant DNA. The four major types of vectors are plasmids, viral vectors, cosmids, and artificial chromosomes. Of these, the most commonly used vectors are plasmids. Common to all engineered vectors are an origin of replication, a multicloning site, and a selectable marker.
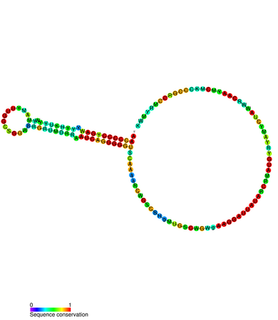
In plasmids, the regulatory region of repBA gene forms a pseudoknot. The repA gene, which encodes a protein likely to function as an initiator for replication, and the repB gene are translationally coupled. The leader sequence of the repA mRNA contains two complementary sequences of 8 bases. Base-pairing between these two sequences forms a pseudoknot which is essential for translation. The first of these complementary sequences is found within a stem-loop, which forms a target for RNAI. Binding of RNAI to this stem-loop inhibits pseudoknot formation and translation of RepA.
Plasmid-mediated resistance is the transfer of antibiotic resistance genes which are carried on plasmids. The plasmids can be transferred between bacteria within the same species or between different species via conjugation. Plasmids often carry multiple antibiotic resistance genes, contributing to the spread of multidrug-resistance (MDR). Antibiotic resistance mediated by MDR plasmids severely limits the treatment options for the infections caused by Gram-negative bacteria, especially family Enterobacteriaceae. The global spread of MDR plasmids has been enhanced by selective pressure from antibiotic usage in human and veterinary medicine.
ColE1 is a plasmid found in bacteria. Its name derives from the fact that it carries a gene for colicin E1. It also codes for immunity from this product with the imm gene. In addition, the plasmid has a series of mobility (mob) genes. Its size and the presence of a single EcoRI recognition site caused it to be considered as a vector candidate.
Self-complementary adeno-associated virus (scAAV) is a viral vector engineered from the naturally occurring adeno-associated virus (AAV) to be used as a tool for gene therapy. Use of recombinant AAV (rAAV) has been successful in clinical trials addressing a variety of diseases. This lab-made progeny of rAAV is termed "self-complementary" because the coding region has been designed to form an intra-molecular double-stranded DNA template. A rate-limiting step for the standard AAV genome involves the second-strand synthesis since the typical AAV genome is a single-stranded DNA template. However, this is not the case for scAAV genomes. Upon infection, rather than waiting for cell mediated synthesis of the second strand, the two complementary halves of scAAV will associate to form one double stranded DNA (dsDNA) unit that is ready for immediate replication and transcription. The caveat of this construct is that instead of the full coding capacity found in rAAV (4.7-6kb) scAAV can only hold about half of that amount (≈2.4kb).
The bacterial, archaeal and plant plastid code is the DNA code used by bacteria, archaea, prokaryotic viruses and chloroplast proteins. It is essentially the same as the standard code, however there are some variations in alternative start codons.
Plasmids must regulate their copy number to ensure that they do not excessively burden the host or become lost during cell division. Plasmids may be either high copy number plasmids or low copy number plasmids; the regulation mechanisms between these two types are often significantly different. Biotechnology applications may involve engineering plasmids to allow a very high copy number. For example, pBR322 is a low copy number plasmid from which several very high copy number cloning vectors have been derived.










