In genetics, an operon is a functioning unit of DNA containing a cluster of genes under the control of a single promoter. The genes are transcribed together into an mRNA strand and either translated together in the cytoplasm, or undergo splicing to create monocistronic mRNAs that are translated separately, i.e. several strands of mRNA that each encode a single gene product. The result of this is that the genes contained in the operon are either expressed together or not at all. Several genes must be co-transcribed to define an operon.

In molecular biology, a riboswitch is a regulatory segment of a messenger RNA molecule that binds a small molecule, resulting in a change in production of the proteins encoded by the mRNA. Thus, an mRNA that contains a riboswitch is directly involved in regulating its own activity, in response to the concentrations of its effector molecule. The discovery that modern organisms use RNA to bind small molecules, and discriminate against closely related analogs, expanded the known natural capabilities of RNA beyond its ability to code for proteins, catalyze reactions, or to bind other RNA or protein macromolecules.
Charles Yanofsky was an American geneticist on the faculty of Stanford University who contributed to the establishment of the one gene-one enzyme hypothesis and discovered attenuation, a riboswitch mechanism in which messenger RNA changes shape in response to a small molecule and thus alters its binding ability for the regulatory region of a gene or operon.

In molecular genetics, a repressor is a DNA- or RNA-binding protein that inhibits the expression of one or more genes by binding to the operator or associated silencers. A DNA-binding repressor blocks the attachment of RNA polymerase to the promoter, thus preventing transcription of the genes into messenger RNA. An RNA-binding repressor binds to the mRNA and prevents translation of the mRNA into protein. This blocking or reducing of expression is called repression.
In genetics, attenuation is a regulatory mechanism for some bacterial operons that results in premature termination of transcription. The canonical example of attenuation used in many introductory genetics textbooks, is ribosome-mediated attenuation of the trp operon. Ribosome-mediated attenuation of the trp operon relies on the fact that, in bacteria, transcription and translation proceed simultaneously. Attenuation involves a provisional stop signal (attenuator), located in the DNA segment that corresponds to the leader sequence of mRNA. During attenuation, the ribosome becomes stalled (delayed) in the attenuator region in the mRNA leader. Depending on the metabolic conditions, the attenuator either stops transcription at that point or allows read-through to the structural gene part of the mRNA and synthesis of the appropriate protein.

Amino acid synthesis is the set of biochemical processes by which the amino acids are produced. The substrates for these processes are various compounds in the organism's diet or growth media. Not all organisms are able to synthesize all amino acids. For example, humans can synthesize 11 of the 20 standard amino acids. These 11 are called the non-essential amino acids).

The gcvB RNA gene encodes a small non-coding RNA involved in the regulation of a number of amino acid transport systems as well as amino acid biosynthetic genes. The GcvB gene is found in enteric bacteria such as Escherichia coli. GcvB regulates genes by acting as an antisense binding partner of the mRNAs for each regulated gene. This binding is dependent on binding to a protein called Hfq. Transcription of the GcvB RNA is activated by the adjacent GcvA gene and repressed by the GcvR gene. A deletion of GcvB RNA from Y. pestis changed colony shape as well as reducing growth. It has been shown by gene deletion that GcvB is a regulator of acid resistance in E. coli. GcvB enhances the ability of the bacterium to survive low pH by upregulating the levels of the alternate sigma factor RpoS. A polymeric form of GcvB has recently been identified. Interaction of GcvB with small RNA SroC triggers the degradation of GcvB by RNase E, lifting the GcvB-mediated mRNA repression of its target genes.

The SAM-II riboswitch is an RNA element found predominantly in Alphaproteobacteria that binds S-adenosyl methionine (SAM). Its structure and sequence appear to be unrelated to the SAM riboswitch found in Gram-positive bacteria. This SAM riboswitch is located upstream of the metA and metC genes in Agrobacterium tumefaciens, and other methionine and SAM biosynthesis genes in other alpha-proteobacteria. Like the other SAM riboswitch, it probably functions to turn off expression of these genes in response to elevated SAM levels. A significant variant of SAM-II riboswitches was found in Pelagibacter ubique and related marine bacteria and called SAM-V. Also, like many structured RNAs, SAM-II riboswitches can tolerate long loops between their stems.

SerC leader is a putative regulatory RNA structure found upstream of the serC-serA operon in some alpha-proteobacteria. The final stem of the structure overlaps the ribosome binding site of the serC reading frame.

suhB, also known as mmgR, is a non-coding RNA found multiple times in the Agrobacterium tumefaciens genome and related alpha-proteobacteria. Other non-coding RNAs uncovered in the same analysis include speF, ybhL, metA, and serC.

The YbhL leader is a putative structured RNA element that is found upstream of the uncharacterized YbhL membrane protein in alpha-proteobacteria.

The eps-Associated RNA element is a conserved RNA motif associated with exopolysaccharide (eps) or capsule biosynthesis genes in a subset of bacteria classified within the order Bacillales. It was initially discovered in Bacillus subtilis, located between the second and third gene in the eps operon. Deletion of the EAR element impairs biofilm formation.
sRNA-Xcc1 is a family of trans-acting non-coding RNA. Homologs of sRNA-Xcc1 are found in a few bacterial strains belonging to alpha-proteobacteria, beta-proteobacteria, gamma-proteobacteria, and delta-proteobacteria. In Xanthomonascampestris pv. campestris, sRNA-Xcc1 is encoded by an integron gene cassette and is under the positive control of the virulence regulators HrpG and HrpX.
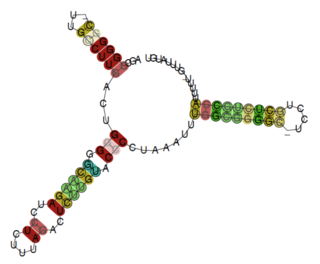
The Pseudomonas rpsL leader is a putative attenuator RNA element identified by bioinformatics searches within bacteria of the Pseudomonadaceae phylum. It is located upstream of the operon encoding ribosomal proteins S12 and S7, and presents a Rho-independent terminator at the 3' end. This RNA is presumed to operate as a non-coding ribosomal protein leader potentially interacting with the S12 or S7 proteins, which are encoded by the operon. The motif might be related to other rpsL leaders, such as the Rickettsia rpsL leader.

The Rickettsia rpsL leader is a putative attenuator element identified by bioinformatics within bacteria of the α-proteobacterial genus Rickettsia. It is located upstream of the operon encoding ribosomal proteins S12 and S7, and presents a Rho-independent terminator at the 3' end. This RNA is presumed to operate as a non-coding ribosomal protein leader potentially interacting with the S12 or S7 proteins, encoded by the operon. The motif might be related to other rpsL leaders, such as that from Pseudomonas.

The γ-proteobacterial rimP leader is a putative attenuator element identified by bioinformatics within bacteria of the γ-proteobacterial phylum. It is located upstream of the rimP-nusA-infB operon encoding RimP, a protein shown to be involved in the 30S ribosomal subunit maturation, NusA, a transcriptional factor controlling termination, and the translation initiation factor IF-2 respectively. The rimP-leader presents a Rho-independent terminator at the 3' end, corresponding to a highly conserved GGGc(...)gCCC motif. This motif is presumed to operate as a non-coding leader. Its mechanism remains unknown, but it is tempting to speculate a regulatory involvement of the NusA protein, which expression has been shown to lower the operon expression, and which is already involved in the attenuation of the Trp, His and S10 operons.
αr45 is a family of bacterial small non-coding RNAs with representatives in a broad group of α-proteobacteria from the order Hyphomicrobiales. The first member of this family (Smr45C) was found in a Sinorhizobium meliloti 1021 locus located in the chromosome (C). Further homology and structure conservation analysis identified homologs in several nitrogen-fixing symbiotic rhizobia, in the plant pathogens belonging to Agrobacterium species as well as in a broad spectrum of Brucella species, in Bartonella species, in several members of the Xanthobactereacea family, and in some representatives of the Beijerinckiaceae family. αr45C RNA species are 147-153 nt long and share a well defined common secondary structure. All of the αr45 transcripts can be catalogued as trans-acting sRNAs expressed from well-defined promoter regions of independent transcription units within intergenic regions (IGRs) of the α-proteobacterial genomes.
The gua operon is responsible for regulating the synthesis of guanosine mono phosphate (GMP), a purine nucleotide, from inosine monophosphate. It consists of two structural genes guaB (encodes for IMP dehydrogenase or and guaA apart from the promoter and operator region.
Transcription-translation coupling is a mechanism of gene expression regulation in which synthesis of an mRNA (transcription) is affected by its concurrent decoding (translation). In prokaryotes, mRNAs are translated while they are transcribed. This allows communication between RNA polymerase, the multisubunit enzyme that catalyzes transcription, and the ribosome, which catalyzes translation. Coupling involves both direct physical interactions between RNA polymerase and the ribosome, as well as ribosome-induced changes to the structure and accessibility of the intervening mRNA that affect transcription.












