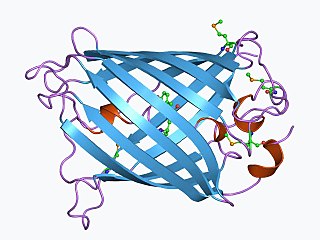
The green fluorescent protein (GFP) is a protein that exhibits bright green fluorescence when exposed to light in the blue to ultraviolet range. The label GFP traditionally refers to the protein first isolated from the jellyfish Aequorea victoria and is sometimes called avGFP. However, GFPs have been found in other organisms including corals, sea anemones, zoanithids, copepods and lancelets.

In molecular biology and biotechnology, a fluorescent tag, also known as a fluorescent label or fluorescent probe, is a molecule that is attached chemically to aid in the detection of a biomolecule such as a protein, antibody, or amino acid. Generally, fluorescent tagging, or labeling, uses a reactive derivative of a fluorescent molecule known as a fluorophore. The fluorophore selectively binds to a specific region or functional group on the target molecule and can be attached chemically or biologically. Various labeling techniques such as enzymatic labeling, protein labeling, and genetic labeling are widely utilized. Ethidium bromide, fluorescein and green fluorescent protein are common tags. The most commonly labelled molecules are antibodies, proteins, amino acids and peptides which are then used as specific probes for detection of a particular target.
A fluorophore is a fluorescent chemical compound that can re-emit light upon light excitation. Fluorophores typically contain several combined aromatic groups, or planar or cyclic molecules with several π bonds.
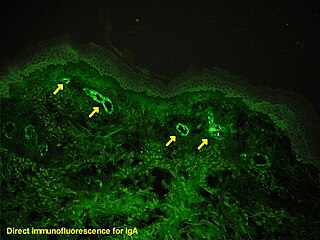
Immunofluorescence is a technique used for light microscopy with a fluorescence microscope and is used primarily on biological samples. This technique uses the specificity of antibodies to their antigen to target fluorescent dyes to specific biomolecule targets within a cell, and therefore allows visualization of the distribution of the target molecule through the sample. The specific region an antibody recognizes on an antigen is called an epitope. There have been efforts in epitope mapping since many antibodies can bind the same epitope and levels of binding between antibodies that recognize the same epitope can vary. Additionally, the binding of the fluorophore to the antibody itself cannot interfere with the immunological specificity of the antibody or the binding capacity of its antigen. Immunofluorescence is a widely used example of immunostaining and is a specific example of immunohistochemistry. This technique primarily makes use of fluorophores to visualise the location of the antibodies.
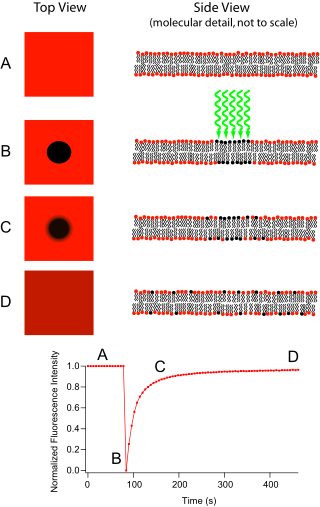
Fluorescence recovery after photobleaching (FRAP) is a method for determining the kinetics of diffusion through tissue or cells. It is capable of quantifying the two-dimensional lateral diffusion of a molecularly thin film containing fluorescently labeled probes, or to examine single cells. This technique is very useful in biological studies of cell membrane diffusion and protein binding. In addition, surface deposition of a fluorescing phospholipid bilayer allows the characterization of hydrophilic surfaces in terms of surface structure and free energy.

Förster resonance energy transfer (FRET), fluorescence resonance energy transfer, resonance energy transfer (RET) or electronic energy transfer (EET) is a mechanism describing energy transfer between two light-sensitive molecules (chromophores). A donor chromophore, initially in its electronic excited state, may transfer energy to an acceptor chromophore through nonradiative dipole–dipole coupling. The efficiency of this energy transfer is inversely proportional to the sixth power of the distance between donor and acceptor, making FRET extremely sensitive to small changes in distance.

A fluorescence microscope is an optical microscope that uses fluorescence instead of, or in addition to, scattering, reflection, and attenuation or absorption, to study the properties of organic or inorganic substances. "Fluorescence microscope" refers to any microscope that uses fluorescence to generate an image, whether it is a simple set up like an epifluorescence microscope or a more complicated design such as a confocal microscope, which uses optical sectioning to get better resolution of the fluorescence image.

Aptamers are short sequences of artificial DNA, RNA, XNA, or peptide that bind a specific target molecule, or family of target molecules. They exhibit a range of affinities, with variable levels of off-target binding and are sometimes classified as chemical antibodies. Aptamers and antibodies can be used in many of the same applications, but the nucleic acid-based structure of aptamers, which are mostly oligonucleotides, is very different from the amino acid-based structure of antibodies, which are proteins. This difference can make aptamers a better choice than antibodies for some purposes.
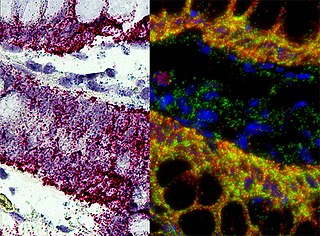
In situ hybridization (ISH) is a type of hybridization that uses a labeled complementary DNA, RNA or modified nucleic acids strand to localize a specific DNA or RNA sequence in a portion or section of tissue or if the tissue is small enough, in the entire tissue, in cells, and in circulating tumor cells (CTCs). This is distinct from immunohistochemistry, which usually localizes proteins in tissue sections.

In optics, photobleaching is the photochemical alteration of a dye or a fluorophore molecule such that it is permanently unable to fluoresce. This is caused by cleaving of covalent bonds or non-specific reactions between the fluorophore and surrounding molecules. Such irreversible modifications in covalent bonds are caused by transition from a singlet state to the triplet state of the fluorophores. The number of excitation cycles to achieve full bleaching varies. In microscopy, photobleaching may complicate the observation of fluorescent molecules, since they will eventually be destroyed by the light exposure necessary to stimulate them into fluorescing. This is especially problematic in time-lapse microscopy.

Bimolecular fluorescence complementation is a technology typically used to validate protein interactions. It is based on the association of fluorescent protein fragments that are attached to components of the same macromolecular complex. Proteins that are postulated to interact are fused to unfolded complementary fragments of a fluorescent reporter protein and expressed in live cells. Interaction of these proteins will bring the fluorescent fragments within proximity, allowing the reporter protein to reform in its native three-dimensional structure and emit its fluorescent signal. This fluorescent signal can be detected and located within the cell using an inverted fluorescence microscope that allows imaging of fluorescence in cells. In addition, the intensity of the fluorescence emitted is proportional to the strength of the interaction, with stronger levels of fluorescence indicating close or direct interactions and lower fluorescence levels suggesting interaction within a complex. Therefore, through the visualisation and analysis of the intensity and distribution of fluorescence in these cells, one can identify both the location and interaction partners of proteins of interest.

Systematic evolution of ligands by exponential enrichment (SELEX), also referred to as in vitro selection or in vitro evolution, is a combinatorial chemistry technique in molecular biology for producing oligonucleotides of either single-stranded DNA or RNA that specifically bind to a target ligand or ligands. These single-stranded DNA or RNA are commonly referred to as aptamers. Although SELEX has emerged as the most commonly used name for the procedure, some researchers have referred to it as SAAB and CASTing SELEX was first introduced in 1990. In 2015, a special issue was published in the Journal of Molecular Evolution in the honor of quarter century of the discovery of SELEX.
Fluorescence Loss in Photobleaching (FLIP) is a fluorescence microscopy technique used to examine movement of molecules inside cells and membranes. A cell membrane is typically labeled with a fluorescent dye to allow for observation. A specific area of this labeled section is then bleached several times using the beam of a confocal laser scanning microscope. After each imaging scan, bleaching occurs again. This occurs several times, to ensure that all accessible fluorophores are bleached since unbleached fluorophores are exchanged for bleached fluorophores, causing movement through the cell membrane. The amount of fluorescence from that region is then measured over a period of time to determine the results of the photobleaching on the cell as a whole.

Fluorescence is used in the life sciences generally as a non-destructive way of tracking or analysing biological molecules. Some proteins or small molecules in cells are naturally fluorescent, which is called intrinsic fluorescence or autofluorescence. Alternatively, specific or general proteins, nucleic acids, lipids or small molecules can be "labelled" with an extrinsic fluorophore, a fluorescent dye which can be a small molecule, protein or quantum dot. Several techniques exist to exploit additional properties of fluorophores, such as fluorescence resonance energy transfer, where the energy is passed non-radiatively to a particular neighbouring dye, allowing proximity or protein activation to be detected; another is the change in properties, such as intensity, of certain dyes depending on their environment allowing their use in structural studies.
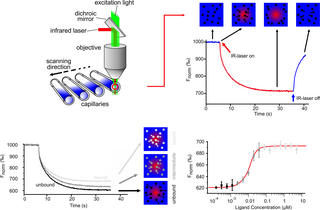
Microscale thermophoresis (MST) is a technology for the biophysical analysis of interactions between biomolecules. Microscale thermophoresis is based on the detection of a temperature-induced change in fluorescence of a target as a function of the concentration of a non-fluorescent ligand. The observed change in fluorescence is based on two distinct effects. On the one hand it is based on a temperature related intensity change (TRIC) of the fluorescent probe, which can be affected by binding events. On the other hand, it is based on thermophoresis, the directed movement of particles in a microscopic temperature gradient. Any change of the chemical microenvironment of the fluorescent probe, as well as changes in the hydration shell of biomolecules result in a relative change of the fluorescence detected when a temperature gradient is applied and can be used to determine binding affinities. MST allows measurement of interactions directly in solution without the need of immobilization to a surface.
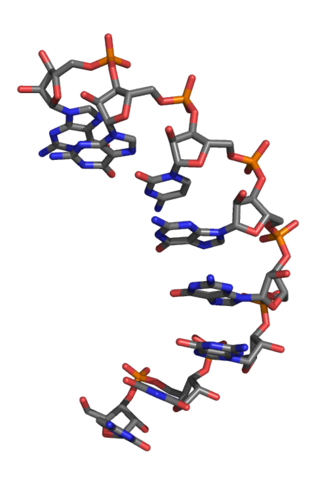
An L-ribonucleic acid aptamer is an RNA-like molecule built from L-ribose units. It is an artificial oligonucleotide named for being a mirror image of natural oligonucleotides. L-RNA aptamers are a form of aptamers. Due to their L-nucleotides, they are highly resistant to degradation by nucleases. L-RNA aptamers are considered potential drugs and are currently being tested in clinical trials.
Photo-activated localization microscopy and stochastic optical reconstruction microscopy (STORM) are widefield fluorescence microscopy imaging methods that allow obtaining images with a resolution beyond the diffraction limit. The methods were proposed in 2006 in the wake of a general emergence of optical super-resolution microscopy methods, and were featured as Methods of the Year for 2008 by the Nature Methods journal. The development of PALM as a targeted biophysical imaging method was largely prompted by the discovery of new species and the engineering of mutants of fluorescent proteins displaying a controllable photochromism, such as photo-activatible GFP. However, the concomitant development of STORM, sharing the same fundamental principle, originally made use of paired cyanine dyes. One molecule of the pair, when excited near its absorption maximum, serves to reactivate the other molecule to the fluorescent state.
Selective microfluidics-based ligand enrichment followed by sequencing (SMiLE-seq) is a technique developed for the rapid identification of DNA binding specificities and affinities of full length monomeric and dimeric transcription factors in a fast and semi-high-throughput fashion.
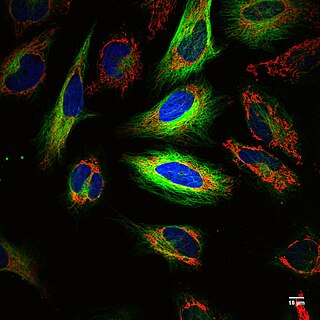
Fluorescence imaging is a type of non-invasive imaging technique that can help visualize biological processes taking place in a living organism. Images can be produced from a variety of methods including: microscopy, imaging probes, and spectroscopy.
RNA therapeutics are a new class of medications based on ribonucleic acid (RNA). Research has been working on clinical use since the 1990s, with significant success in cancer therapy in the early 2010s. In 2020 and 2021, mRNA vaccines have been developed globally for use in combating the coronavirus disease. The Pfizer–BioNTech COVID-19 vaccine was the first mRNA vaccine approved by a medicines regulator, followed by the Moderna COVID-19 vaccine, and others.

















