
In molecular biology, a riboswitch is a regulatory segment of a messenger RNA molecule that binds a small molecule, resulting in a change in production of the proteins encoded by the mRNA. Thus, an mRNA that contains a riboswitch is directly involved in regulating its own activity, in response to the concentrations of its effector molecule. The discovery that modern organisms use RNA to bind small molecules, and discriminate against closely related analogs, expanded the known natural capabilities of RNA beyond its ability to code for proteins, catalyze reactions, or to bind other RNA or protein macromolecules.
Bacterial translation is the process by which messenger RNA is translated into proteins in bacteria.

The ykkC/yxkD leader is a conserved RNA structure found upstream of the ykkC and yxkD genes in Bacillus subtilis and related genes in other bacteria. The function of this family is unclear for many years although it has been suggested that it may function to switch on efflux pumps and detoxification systems in response to harmful environmental molecules. The Thermoanaerobacter tengcongensis sequence AE013027 overlaps with that of purine riboswitch suggesting that the two riboswitches may work in conjunction to regulate the upstream gene which codes for TTE0584 (Q8RC62), a member of the permease family.

The signal recognition particle RNA, is part of the signal recognition particle (SRP) ribonucleoprotein complex. SRP recognizes the signal peptide and binds to the ribosome, halting protein synthesis. SRP-receptor is a protein that is embedded in a membrane, and which contains a transmembrane pore. When the SRP-ribosome complex binds to SRP-receptor, SRP releases the ribosome and drifts away. The ribosome resumes protein synthesis, but now the protein is moving through the SRP-receptor transmembrane pore.

PreQ1-II riboswitches form a class of riboswitches that specifically bind pre-queuosine1 (PreQ1), a precursor of the modified nucleoside queuosine. They are found in certain species of Streptococcus and Lactococcus, and were originally identified as a conserved RNA secondary structure called the "COG4708 motif". All known members of this riboswitch class appear to control members of COG4708 genes. These genes are predicted to encode membrane-bound proteins and have been proposed to be a transporter of preQ1, or a related metabolite, based on their association with preQ1-binding riboswitches. PreQ1-II riboswitches have no apparent similarities in sequence or structure to preQ1-I riboswitches, a previously discovered class of preQ1-binding riboswitches. PreQ1 thus joins S-adenosylmethionine as the second metabolite to be found that is the ligand of more than one riboswitch class.

The Moco RNA motif is a conserved RNA structure that is presumed to be a riboswitch that binds molybdenum cofactor or the related tungsten cofactor. Genetic experiments support the hypothesis that the Moco RNA motif corresponds to a genetic control element that responds to changing concentrations of molybdenum or tungsten cofactor. As these cofactors are not available in purified form, in vitro binding assays cannot be performed. However, the genetic data, complex structure of the RNA and the failure to detect a protein involved in the regulation suggest that the Moco RNA motif corresponds to a class of riboswitches.

The Cyano-S1 RNA motif is a conserved RNA structure present in some species of Cyanobacteria. Cyano-S1 RNAs are consistently found upstream of genes encoding ribosomal protein S1, a subunit of the ribosome. Therefore, they are presumed to be ribosomal protein leaders, i.e., cis-regulatory elements to which the ribosomal protein S1 binds, thereby controlling its expression levels.
The lacto-2 RNA motif is an RNA structure that is conserved amongst bacteria within the order Lactobacillales. The motif consists of a stem-loop whose stem is interrupted by many internal loops and bulges. Nucleotide identities in many places are conserved, and one internal loop in particular is highly conserved.

The L17 downstream element RNA motif is a conserved RNA structure identified in bacteria by bioinformatics. All known L17 downstream elements were detected immediately downstream of genes encoding the L17 subunit of the ribosome, and therefore might be in the 3' untranslated regions of these genes. The element is found in a variety of lactic acid bacteria and in the genus Listeria.

The leu/phe-leader RNA motif is a conserved RNA structure identified by bioinformatics. These RNAs function as peptide leaders. They contain a short open reading frame (ORF) that contains many codons for leucine or phenylalanine. Normally, expression of the downstream genes is suppressed. However, when cellular concentrations of the relevant amino acid is low, ribosome stalling leads to an alternate structure that enables downstream gene expression.

The msiK RNA motif describes a conserved RNA structure discovered using bioinformatics. The RNA is always found in the presumed 5' untranslated regions of genes annotated as msiK, and is therefore hypothesized to be an RNA-based cis-regulatory element that regulates these genes.

The rmf RNA motif is a conserved RNA structure that was originally detected using bioinformatics. rmf RNAs are consistently foundwithin species classified into the genus Pseudomonas, and is located potentially in the 5′ untranslated regions of rmf genes. These genes encodes the ribosome modulation factor protein, which affects the translation of genes by modifying ribosome structure in response to stress such as starvation. This ribosome modulation is a part of the stringent response in bacteria. The likely biological role of rmf RNAs is ambiguous. Since the RNA could be in the 5′ UTRs of protein-coding genes, it was hypothesized that it functions as a cis-regulatory element. This hypothesis is bolstered by the observation that ribosome modulation factor binds ribosomal RNA, and many cis-regulatory RNAs called ribosomal protein leaders participate in a feedback regulation mechanism by binding to proteins that normally bind to ribosomal RNA. However, since rmf RNAs are not very close to the rmf genes, they might function as non-coding RNAs.

The sucA-II RNA motif is a conserved RNA structure identified by bioinformatics. It is consistently found in the presumed 5' untranslated regions of sucA genes, which encode Oxoglutarate dehydrogenase enzymes that participate in the citric acid cycle. Given this arrangement, sucA-II RNAs might regulate the downstream sucA gene. This genetic arrangement is similar to the previously reported sucA RNA motif. However, sucA-II RNAs are found only in bacteria classified within the genus Pseudomonas, whereas the previously reported motif is found only in betaproteobacteria.

The sucC RNA motif is a conserved RNA structure discovered using bioinformatics. sucC RNAs are found in the genus Pseudomonas, and are consistently found in possible 5' untranslated regions of sucC genes. These genes encode Succinyl coenzyme A synthetase, and are hypothesised to be regulated by the sucC RNAs. sucC genes participate in the citric acid cycle, and another gene involved in the citric acid cycle, sucA, is also predicted to be regulated by a conserved RNA structure.

The engA RNA motif is a conserved RNA structure that was discovered by bioinformatics. engA motifs are found in bacteria within the genus Prevotella.
The leuA-Halobacteria RNA motif is a conserved RNA structure that was discovered by bioinformatics. leuA-Halobacteria motifs are found in Halobacteriaceae, a lineage of archaea.

The raiA RNA motif is a conserved RNA structure that was discovered by bioinformatics. raiA motif RNAs are found in Actinobacteria AND Firmicutes, and have many conserved features—including conserved nucleotide positions, conserved secondary structures and associated protein-coding genes—in both of these phyla. Some conserved features of the raiA RNA motif suggest that they function as cis-regulatory elements, but other aspects of the motif suggest otherwise.
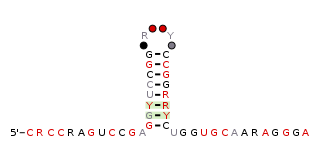
The raiA-hairpin RNA motif is a conserved RNA structure that was discovered by bioinformatics. raiA-hairpin motif RNAs are found in organism classified within the genus Streptomyces.

The Rothia-sucC RNA motif is a conserved RNA structure that was discovered by bioinformatics. Rothia-sucC motif RNAs are found in the Actinobacterial genus Rothia.

The uup RNA motif is a conserved RNA structure that was discovered by bioinformatics. uup motif RNAs are found in Firmicutes and Gammaproteobacteria.

















