The 5′ untranslated region is the region of an mRNA that is directly upstream from the initiation codon. This region is important for the regulation of translation of a transcript by differing mechanisms in viruses, prokaryotes and eukaryotes. While called untranslated, the 5′ UTR or a portion of it is sometimes translated into a protein product. This product can then regulate the translation of the main coding sequence of the mRNA. In many organisms, however, the 5′ UTR is completely untranslated, instead forming complex secondary structure to regulate translation.
An internal ribosome entry site, abbreviated IRES, is an RNA element that allows for translation initiation in cap-independent manner, as part of the greater process of protein synthesis. In prokaryotic translation, initiation typically occurs at the 5' end of rRNA molecules, since 5' cap recognition is required for the assembly of the initiation complex. The location for IRES elements is often in the 5'UTR, but can also occur elsewhere in tRNAs.
Eukaryotic translation is the biological process by which transfer RNA is translated into proteins in eukaryotes. It consists of four phases: gene regulation, elongation, termination, and recycling.
The Kozak consensus sequence is a nucleic acid motif that functions as the protein translation initiation site in most eukaryotic mRNA transcripts. Regarded as the optimum sequence for initiating translation in eukaryotes, the sequence is an integral aspect of protein regulation and overall cellular health as well as having implications in human disease. It ensures that a protein is correctly translated from the genetic message, mediating ribosome assembly and translation initiation. A wrong start site can result in non-functional proteins. As it has become more studied, expansions of the nucleotide sequence, bases of importance, and notable exceptions have arisen. The sequence was named after the scientist who discovered it, Marilyn Kozak. Kozak discovered the sequence through a detailed analysis of DNA genomic sequences.
Eukaryotic initiation factors (eIFs) are proteins or protein complexes involved in the initiation phase of eukaryotic translation. These proteins help stabilize the formation of ribosomal preinitiation complexes around the start codon and are an important input for post-transcription gene regulation. Several initiation factors form a complex with the small 40S ribosomal subunit and Met-tRNAiMet called the 43S preinitiation complex. Additional factors of the eIF4F complex recruit the 43S PIC to the five-prime cap structure of the mRNA, from which the 43S particle scans 5'-->3' along the mRNA to reach an AUG start codon. Recognition of the start codon by the Met-tRNAiMet promotes gated phosphate and eIF1 release to form the 48S preinitiation complex, followed by large 60S ribosomal subunit recruitment to form the 80S ribosome. There exist many more eukaryotic initiation factors than prokaryotic initiation factors, reflecting the greater biological complexity of eukaryotic translation. There are at least twelve eukaryotic initiation factors, composed of many more polypeptides, and these are described below.

This family represents the internal ribosome entry site (IRES) of the Picornaviruses. IRES elements allow cap and end-independent translation of mRNA in the host cell. The IRES achieves this by mediating the internal initiation of translation by recruiting a ribosomal 43S pre-initiation complex directly to the initiation codon and eliminates the requirement for the eukaryotic initiation factor eIF4F.
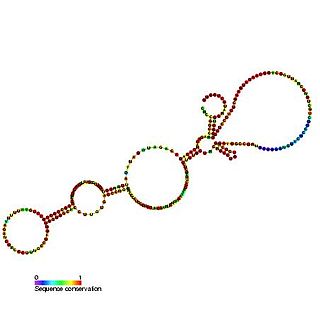

The BiP internal ribosome entry site (IRES) is an RNA element present in the 5' UTR of the mRNA of BiP protein and allows cap-independent translation. BiP protein expression has been found to be significantly enhanced by the heat shock response due to internal ribosome entry site (IRES)-dependent translation. It is thought that this translational mechanism is essential for the survival of cells under stress.

This family represents the internal ribosome entry site (IRES) of the hepatitis A virus. HAV IRES is a 450 nucleotide long sequence located in the 735 nt long 5’ UTR of Hepatitis A viral RNA genome. IRES elements allow cap and end-independent translation of mRNA in the host cell. The IRES achieves this by mediating the internal initiation of translation by recruiting a ribosomal 40S pre-initiation complex directly to the initiation codon and eliminates the requirement for eukaryotic initiation factor, eIF4F.
The Hepatitis C virus internal ribosome entry site, or HCV IRES, is an RNA structure within the 5'UTR of the HCV genome that mediates cap-independent translation initiation.
The insulin-like growth factor II (IGF-II) internal ribosome entry site IRES is found in the 5' UTR of IGF-II leader 2 mRNA. This RNA element allows cap-independent translation of the mRNA and it is thought that this family may facilitate a continuous IGF-II production in rapidly dividing cells during development. Ribosomal scanning on human insulin-like growth factor II (IGF-II) is hard to comprehend due to one open reading frame and the ability for the hormone to fold into a stable structure.

The Mnt internal ribosome entry site (IRES) is an RNA element. Mnt is a transcriptional repressor related to the Myc/Mad family of transcription factors. It is thought that this IRES allows efficient Mnt synthesis when cap-dependent translation initiation is reduced.

The N-myc internal ribosome entry site (IRES) is an RNA element found in the n-myc gene. The myc family of genes when expressed are known to be involved in the control of cell growth, differentiation and apoptosis. n-myc mRNA has an alternative method of translation via an internal ribosome entry site where ribosomes are recruited to the IRES located in the 5' UTR thus bypassing the typical eukaryotic cap-dependent translation pathway.

This family represents the internal ribosome entry site (IRES) of the pestiviruses. The pestivirus IRES allows cap and end-independent translation of mRNA in the host cell. The IRES achieves this by mediating the internal initiation of translation by recruiting a ribosomal 43S pre-initiation complex directly to the initiation codon and eliminates the requirement for the eukaryotic initiation factor, eIF4F. The classical swine fever virus UTR described appears to be longer at the 5' end than other pestivirus UTRs. This family represents the conserved core.

The TrkB internal ribosome entry site (IRES) is an RNA element which is present in the 5' UTR sequence of the mRNA. TrkB is a neurotrophin receptor which is essential for the development and maintenance of the nervous system. The internal ribosome entry site IRES element allows cap-independent translation of TrkB which may be needed for efficient translation in neuronal dendrites.
A ribosome binding site, or ribosomal binding site (RBS), is a sequence of nucleotides upstream of the start codon of an mRNA transcript that is responsible for the recruitment of a ribosome during the initiation of translation. Mostly, RBS refers to bacterial sequences, although internal ribosome entry sites (IRES) have been described in mRNAs of eukaryotic cells or viruses that infect eukaryotes. Ribosome recruitment in eukaryotes is generally mediated by the 5' cap present on eukaryotic mRNAs.
The eukaryotic small ribosomal subunit (40S) is the smaller subunit of the eukaryotic 80S ribosomes, with the other major component being the large ribosomal subunit (60S). The "40S" and "60S" names originate from the convention that ribosomal particles are denoted according to their sedimentation coefficients in Svedberg units. It is structurally and functionally related to the 30S subunit of 70S prokaryotic ribosomes. However, the 40S subunit is much larger than the prokaryotic 30S subunit and contains many additional protein segments, as well as rRNA expansion segments.
Eukaryotic translation initiation factor 4 G (eIF4G) is a protein involved in eukaryotic translation initiation and is a component of the eIF4F cap-binding complex. Orthologs of eIF4G have been studied in multiple species, including humans, yeast, and wheat. However, eIF4G is exclusively found in domain Eukarya, and not in domains Bacteria or Archaea, which do not have capped mRNA. As such, eIF4G structure and function may vary between species, although the human eIF4G 1 has been the focus of extensive studies.
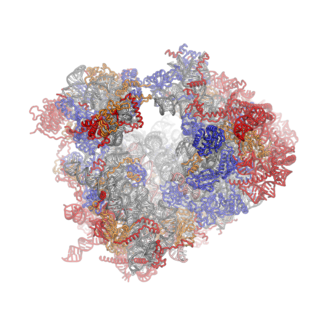
Ribosomes are a large and complex molecular machine that catalyzes the synthesis of proteins, referred to as translation. The ribosome selects aminoacylated transfer RNAs (tRNAs) based on the sequence of a protein-encoding messenger RNA (mRNA) and covalently links the amino acids into a polypeptide chain. Ribosomes from all organisms share a highly conserved catalytic center. However, the ribosomes of eukaryotes are much larger than prokaryotic ribosomes and subject to more complex regulation and biogenesis pathways. Eukaryotic ribosomes are also known as 80S ribosomes, referring to their sedimentation coefficients in Svedberg units, because they sediment faster than the prokaryotic (70S) ribosomes. Eukaryotic ribosomes have two unequal subunits, designated small subunit (40S) and large subunit (60S) according to their sedimentation coefficients. Both subunits contain dozens of ribosomal proteins arranged on a scaffold composed of ribosomal RNA (rRNA). The small subunit monitors the complementarity between tRNA anticodon and mRNA, while the large subunit catalyzes peptide bond formation.
The P-site is the second binding site for tRNA in the ribosome. The other two sites are the A-site (aminoacyl), which is the first binding site in the ribosome, and the E-site (exit), the third. During protein translation, the P-site holds the tRNA which is linked to the growing polypeptide chain. When a stop codon is reached, the peptidyl-tRNA bond of the tRNA located in the P-site is cleaved releasing the newly synthesized protein. During the translocation step of the elongation phase, the mRNA is advanced by one codon, coupled to movement of the tRNAs from the ribosomal A to P and P to E sites, catalyzed by elongation factor EF-G.










