
The alpha helix (α-helix) is a common motif in the secondary structure of proteins and is a right hand-helix conformation in which every backbone N−H group hydrogen bonds to the backbone C=O group of the amino acid located four residues earlier along the protein sequence.

Ubiquitin is a small regulatory protein found in most tissues of eukaryotic organisms, i.e., it is found ubiquitously. It was discovered in 1975 by Gideon Goldstein and further characterized throughout the late 1970s and 1980s. Four genes in the human genome code for ubiquitin: UBB, UBC, UBA52 and RPS27A.

The globins are a superfamily of heme-containing globular proteins, involved in binding and/or transporting oxygen. These proteins all incorporate the globin fold, a series of eight alpha helical segments. Two prominent members include myoglobin and hemoglobin. Both of these proteins reversibly bind oxygen via a heme prosthetic group. They are widely distributed in many organisms.

Peripheral membrane proteins are membrane proteins that adhere only temporarily to the biological membrane with which they are associated. These proteins attach to integral membrane proteins, or penetrate the peripheral regions of the lipid bilayer. The regulatory protein subunits of many ion channels and transmembrane receptors, for example, may be defined as peripheral membrane proteins. In contrast to integral membrane proteins, peripheral membrane proteins tend to collect in the water-soluble component, or fraction, of all the proteins extracted during a protein purification procedure. Proteins with GPI anchors are an exception to this rule and can have purification properties similar to those of integral membrane proteins.
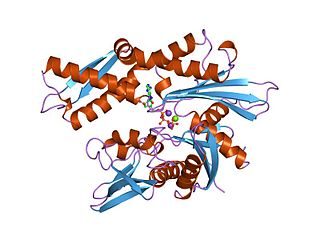
The 70 kilodalton heat shock proteins are a family of conserved ubiquitously expressed heat shock proteins. Proteins with similar structure exist in virtually all living organisms. Intracellularly localized Hsp70s are an important part of the cell's machinery for protein folding, performing chaperoning functions, and helping to protect cells from the adverse effects of physiological stresses. Additionally, membrane-bound Hsp70s have been identified as a potential target for cancer therapies and their extracellularly localized counterparts have been identified as having both membrane-bound and membrane-free structures.
DnaG is a bacterial DNA primase and is encoded by the dnaG gene. The enzyme DnaG, and any other DNA primase, synthesizes short strands of RNA known as oligonucleotides during DNA replication. These oligonucleotides are known as primers because they act as a starting point for DNA synthesis. DnaG catalyzes the synthesis of oligonucleotides that are 10 to 60 nucleotides long, however most of the oligonucleotides synthesized are 11 nucleotides. These RNA oligonucleotides serve as primers, or starting points, for DNA synthesis by bacterial DNA polymerase III. DnaG is important in bacterial DNA replication because DNA polymerase cannot initiate the synthesis of a DNA strand, but can only add nucleotides to a preexisting strand. DnaG synthesizes a single RNA primer at the origin of replication. This primer serves to prime leading strand DNA synthesis. For the other parental strand, the lagging strand, DnaG synthesizes an RNA primer every few kilobases (kb). These primers serve as substrates for the synthesis of Okazaki fragments.

A ubiquitin ligase is a protein that recruits an E2 ubiquitin-conjugating enzyme that has been loaded with ubiquitin, recognizes a protein substrate, and assists or directly catalyzes the transfer of ubiquitin from the E2 to the protein substrate. In simple and more general terms, the ligase enables movement of ubiquitin from a ubiquitin carrier to another thing by some mechanism. The ubiquitin, once it reaches its destination, ends up being attached by an isopeptide bond to a lysine residue, which is part of the target protein. E3 ligases interact with both the target protein and the E2 enzyme, and so impart substrate specificity to the E2. Commonly, E3s polyubiquitinate their substrate with Lys48-linked chains of ubiquitin, targeting the substrate for destruction by the proteasome. However, many other types of linkages are possible and alter a protein's activity, interactions, or localization. Ubiquitination by E3 ligases regulates diverse areas such as cell trafficking, DNA repair, and signaling and is of profound importance in cell biology. E3 ligases are also key players in cell cycle control, mediating the degradation of cyclins, as well as cyclin dependent kinase inhibitor proteins. The human genome encodes over 600 putative E3 ligases, allowing for tremendous diversity in substrates.
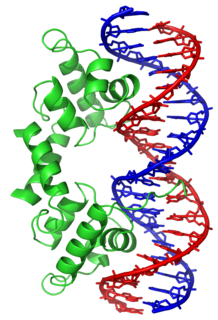
Helix-turn-helix is a DNA-binding protein (DBP). The helix-turn-helix (HTH) is a major structural motif capable of binding DNA. Each monomer incorporates two α helices, joined by a short strand of amino acids, that bind to the major groove of DNA. The HTH motif occurs in many proteins that regulate gene expression. It should not be confused with the helix–loop–helix motif.

Protein–protein interactions (PPIs) are physical contacts of high specificity established between two or more protein molecules as a result of biochemical events steered by interactions that include electrostatic forces, hydrogen bonding and the hydrophobic effect. Many are physical contacts with molecular associations between chains that occur in a cell or in a living organism in a specific biomolecular context.

A leucine zipper is a common three-dimensional structural motif in proteins. They were first described by Landschulz and collaborators in 1988 when they found that an enhancer binding protein had a very characteristic 30-amino acid segment and the display of these amino acid sequences on an idealized alpha helix revealed a periodic repetition of leucine residues at every seventh position over a distance covering eight helical turns. The polypeptide segments containing these periodic arrays of leucine residues were proposed to exist in an alpha-helical conformation and the leucine side chains from one alpha helix interdigitate with those from the alpha helix of a second polypeptide, facilitating dimerization.

Ubiquitin-like modifier activating enzyme 1 (UBA1) is an enzyme which in humans is encoded by the UBA1 gene. UBA1 participates in ubiquitination and the NEDD8 pathway for protein folding and degradation, among many other biological processes. This protein has been linked to X-linked spinal muscular atrophy type 2, neurodegenerative diseases, and cancers.

A protein domain is a region of the protein's polypeptide chain that is self-stabilizing and that folds independently from the rest. Each domain forms a compact folded three-dimensional structure. Many proteins consist of several domains. One domain may appear in a variety of different proteins. Molecular evolution uses domains as building blocks and these may be recombined in different arrangements to create proteins with different functions. In general, domains vary in length from between about 50 amino acids up to 250 amino acids in length. The shortest domains, such as zinc fingers, are stabilized by metal ions or disulfide bridges. Domains often form functional units, such as the calcium-binding EF hand domain of calmodulin. Because they are independently stable, domains can be "swapped" by genetic engineering between one protein and another to make chimeric proteins.
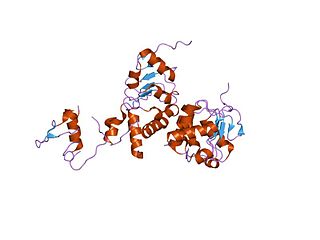
Nuclear RNA export factor 1, also known as NXF1 or TAP, is a protein which in humans is encoded by the NXF1 gene.

Cbl is a mammalian gene encoding the protein CBL which is an E3 ubiquitin-protein ligase involved in cell signalling and protein ubiquitination. Mutations to this gene have been implicated in a number of human cancers, particularly acute myeloid leukaemia.

Azurin is a small, periplasmic, bacterial blue copper protein found in Pseudomonas, Bordetella, or Alcaligenes bacteria. Azurin moderates single-electron transfer between enzymes associated with the cytochrome chain by undergoing oxidation-reduction between Cu(I) and Cu(II). Each monomer of an azurin tetramer has a molecular weight of approximately 14kDa, contains a single copper atom, is intensively blue, and has a fluorescence emission band centered at 308 nm.

The tetratricopeptide repeat (TPR) is a structural motif. It consists of a degenerate 34 amino acid tandem repeat identified in a wide variety of proteins. It is found in tandem arrays of 3–16 motifs, which form scaffolds to mediate protein–protein interactions and often the assembly of multiprotein complexes. These alpha-helix pair repeats usually fold together to produce a single, linear solenoid domain called a TPR domain. Proteins with such domains include the anaphase-promoting complex (APC) subunits cdc16, cdc23 and cdc27, the NADPH oxidase subunit p67-phox, hsp90-binding immunophilins, transcription factors, the protein kinase R (PKR), the major receptor for peroxisomal matrix protein import PEX5, protein arginine methyltransferase 9 (PRMT9), and mitochondrial import proteins.
DNA damage-binding protein or UV-DDB is a protein complex that is responsible for repair of UV-damaged DNA. This complex is composed of two protein subunits, a large subunit DDB1 (p127) and a small subunit DDB2 (p48). When cells are exposed to UV radiation, DDB1 moves from the cytosol to the nucleus and binds to DDB2, thus forming the UV-DDB complex. This complex formation is highly favorable and it is demonstrated by UV-DDB's binding preference and high affinity to the UV lesions in the DNA. This complex functions in nucleotide excision repair, recognising UV-induced (6-4) pyrimidine-pyrimidone photoproducts and cyclobutane pyrimidine dimers.

A pyrin domain is a protein domain and a subclass of protein motif known as the death fold, the 4th and most recently discovered member of the death domain superfamily (DDF). It was originally discovered in the pyrin protein, or marenostrin, encoded by MEFV. The mutation of the MEFV gene is the cause of the disease known as Familial Mediterranean Fever. The domain is encoded in 23 human proteins and at least 31 mouse genes.
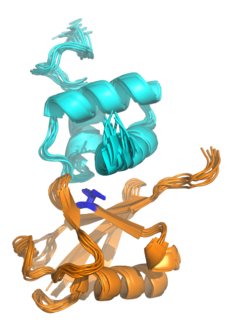
Ubiquitin-associated (UBA) domains are protein domains that non-covalently interact with ubiquitin through protein-protein interactions. Ubiquitin is a small protein that is covalently linked to other proteins as part of intracellular signaling pathways, often as a signal for protein degradation. UBA domains are among the most common ubiquitin-binding domains.
In molecular biology, the Ubiquitin-Interacting Motif (UIM), or 'LALAL-motif', is a sequence motif of about 20 amino acid residues, which was first described in the 26S proteasome subunit PSD4/RPN-10 that is known to recognise ubiquitin. In addition, the UIM is found, often in tandem or triplet arrays, in a variety of proteins either involved in ubiquitination and ubiquitin metabolism, or known to interact with ubiquitin-like modifiers. Among the UIM proteins are two different subgroups of the UBP family of deubiquitinating enzymes, one F-box protein, one family of HECT-containing ubiquitin-ligases (E3s) from plants, and several proteins containing ubiquitin-associated UBA and/or UBX domains. In most of these proteins, the UIM occurs in multiple copies and in association with other domains such as UBA, UBX, ENTH domain, EH, VHS, SH3 domain, HECT, VWFA, EF-hand calcium-binding, WD-40, F-box, LIM, protein kinase, ankyrin, PX, phosphatidylinositol 3- and 4-kinase, C2 domain, OTU, DnaJ domain, RING-finger or FYVE-finger. UIMs have been shown to bind ubiquitin and to serve as a specific targeting signal important for monoubiquitination. Thus, UIMs may have several functions in ubiquitin metabolism each of which may require different numbers of UIMs.


















