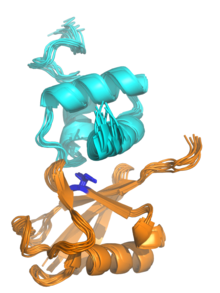Proteasomes are protein complexes which degrade ubiquitin-tagged proteins by proteolysis, a chemical reaction that breaks peptide bonds. Enzymes that help such reactions are called proteases.

Ubiquitin is a small regulatory protein found in most tissues of eukaryotic organisms, i.e., it is found ubiquitously. It was discovered in 1975 by Gideon Goldstein and further characterized throughout the late 1970s and 1980s. Four genes in the human genome code for ubiquitin: UBB, UBC, UBA52 and RPS27A.
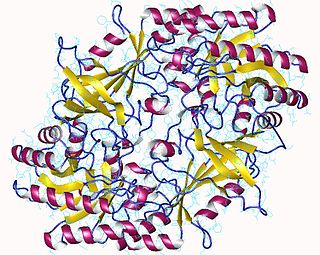
The enzyme ornithine decarboxylase catalyzes the decarboxylation of ornithine to form putrescine. This reaction is the committed step in polyamine synthesis. In humans, this protein has 461 amino acids and forms a homodimer.

A ubiquitin ligase is a protein that recruits an E2 ubiquitin-conjugating enzyme that has been loaded with ubiquitin, recognizes a protein substrate, and assists or directly catalyzes the transfer of ubiquitin from the E2 to the protein substrate. In simple and more general terms, the ligase enables movement of ubiquitin from a ubiquitin carrier to another protein by some mechanism. The ubiquitin, once it reaches its destination, ends up being attached by an isopeptide bond to a lysine residue, which is part of the target protein. E3 ligases interact with both the target protein and the E2 enzyme, and so impart substrate specificity to the E2. Commonly, E3s polyubiquitinate their substrate with Lys48-linked chains of ubiquitin, targeting the substrate for destruction by the proteasome. However, many other types of linkages are possible and alter a protein's activity, interactions, or localization. Ubiquitination by E3 ligases regulates diverse areas such as cell trafficking, DNA repair, and signaling and is of profound importance in cell biology. E3 ligases are also key players in cell cycle control, mediating the degradation of cyclins, as well as cyclin dependent kinase inhibitor proteins. The human genome encodes over 600 putative E3 ligases, allowing for tremendous diversity in substrates.

Endoplasmic-reticulum-associated protein degradation (ERAD) designates a cellular pathway which targets misfolded proteins of the endoplasmic reticulum for ubiquitination and subsequent degradation by a protein-degrading complex, called the proteasome.

ADP-ribosylation is the addition of one or more ADP-ribose moieties to a protein. It is a reversible post-translational modification that is involved in many cellular processes, including cell signaling, DNA repair, gene regulation and apoptosis. Improper ADP-ribosylation has been implicated in some forms of cancer. It is also the basis for the toxicity of bacterial compounds such as cholera toxin, diphtheria toxin, and others.

Proteasome subunit alpha type-3 also known as macropain subunit C8 and proteasome component C8 is a protein that in humans is encoded by the PSMA3 gene. This protein is one of the 17 essential subunits that contributes to the complete assembly of 20S proteasome complex.

UV excision repair protein RAD23 homolog A is a protein that in humans is encoded by the RAD23A gene.

Proteasome subunit beta type-3, also known as 20S proteasome subunit beta-3, is a protein that in humans is encoded by the PSMB3 gene. This protein is one of the 17 essential subunits that contribute to the complete assembly of the 20S proteasome complex. In particular, proteasome subunit beta type-2, along with other beta subunits, assemble into two heptameric rings and subsequently a proteolytic chamber for substrate degradation. The eukaryotic proteasome recognizes degradable proteins, including damaged proteins for protein quality control purpose or key regulatory protein components for dynamic biological processes.

UV excision repair protein RAD23 homolog B is a protein that in humans is encoded by the RAD23B gene.

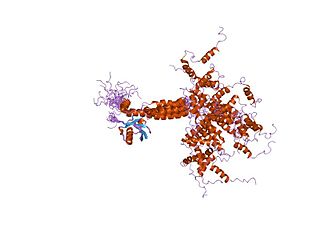
Valosin-containing protein (VCP) or transitional endoplasmic reticulum ATPase also known as p97 in mammals and CDC48 in S. cerevisiae, is an enzyme that in humans is encoded by the VCP gene. The TER ATPase is an ATPase enzyme present in all eukaryotes and archaebacteria. Its main function is to segregate protein molecules from large cellular structures such as protein assemblies, organelle membranes and chromatin, and thus facilitate the degradation of released polypeptides by the multi-subunit protease proteasome.

26S proteasome non-ATPase regulatory subunit 4, also as known as 26S Proteasome Regulatory Subunit Rpn10, is an enzyme that in humans is encoded by the PSMD4 gene. This protein is one of the 19 essential subunits that contributes to the complete assembly of 19S proteasome complex.

Proteasome subunit alpha type-6 is a protein that in humans is encoded by the PSMA6 gene. This protein is one of the 17 essential subunits that contributes to the complete assembly of 20S proteasome complex.

Proteasome subunit alpha type-7 also known as 20S proteasome subunit alpha-4 is a protein that in humans is encoded by the PSMA7 gene. This protein is one of the 17 essential subunits that contributes to the complete assembly of 20S proteasome complex.

26S proteasome non-ATPase regulatory subunit 2, also as known as 26S Proteasome Regulatory Subunit Rpn1, is an enzyme that in humans is encoded by the PSMD2 gene.

In molecular biology, protein aggregation is a phenomenon in which intrinsically-disordered or mis-folded proteins aggregate either intra- or extracellularly. Protein aggregates have been implicated in a wide variety of diseases known as amyloidoses, including ALS, Alzheimer's, Parkinson's and prion disease.

Prokaryotic ubiquitin-like protein (Pup) is a functional analog of ubiquitin found in the prokaryote Mycobacterium tuberculosis. Like ubiquitin, Pup serves to direct proteins to the proteasome for degradation in the Pup-proteasome system (PPS). However, the enzymology of ubiquitylation and pupylation is different, owing to their distinct evolutionary origins. In contrast to the three-step reaction of ubiquitylation, pupylation requires only two steps, and thus only two enzymes are involved in pupylation. The enzymes involved in pupylation are descended from glutamine synthetase.
In molecular biology, the Ubiquitin-Interacting Motif (UIM), or 'LALAL-motif', is a sequence motif of about 20 amino acid residues, which was first described in the 26S proteasome subunit PSD4/RPN-10 that is known to recognise ubiquitin. In addition, the UIM is found, often in tandem or triplet arrays, in a variety of proteins either involved in ubiquitination and ubiquitin metabolism, or known to interact with ubiquitin-like modifiers. Among the UIM proteins are two different subgroups of the UBP family of deubiquitinating enzymes, one F-box protein, one family of HECT-containing ubiquitin-ligases (E3s) from plants, and several proteins containing ubiquitin-associated UBA and/or UBX domains. In most of these proteins, the UIM occurs in multiple copies and in association with other domains such as UBA, UBX, ENTH domain, EH, VHS, SH3 domain, HECT, VWFA, EF-hand calcium-binding, WD-40, F-box, LIM, protein kinase, ankyrin, PX, phosphatidylinositol 3- and 4-kinase, C2 domain, OTU, DnaJ domain, RING-finger or FYVE-finger. UIMs have been shown to bind ubiquitin and to serve as a specific targeting signal important for monoubiquitination. Thus, UIMs may have several functions in ubiquitin metabolism each of which may require different numbers of UIMs.

Ubiquitin-binding domains (UBDs) are protein domains that recognise and bind non-covalently to ubiquitin through protein-protein interactions. As of 2019, a total of 29 types of UBDs had been identified in the human proteome. Most UBDs bind to ubiquitin only weakly, with binding affinities in the low to mid μM range. Proteins containing UBDs are known as ubiquitin-binding proteins or sometimes as "ubiquitin receptors".
UBXD8 is a protein in the Ubiquitin regulatory X (UBX) domain-containing protein family. The UBX domain contains many eukaryotic proteins that have similarities in amino acid sequence to the tiny protein modifier ubiquitin. UBXD8 engages in a molecular interaction with p97, a protein that is essential for the degradation of membrane proteins associated with the endoplasmic reticulum (ER) through the proteasome. Ubxd8 possesses a UBA domain, alongside the UBX domain, that could interact with polyubiquitin chains. Additionally, it possesses a UAS domain of undetermined function, and this protein is used as a protein sensor that detects long chain unsaturated fatty acids (FAs), having a vital function in regulating the balance of Fatty Acids within cells to maintain cellular homeostasis.