Synechocystis sp. PCC6803 is a strain of unicellular, freshwater cyanobacteria. Synechocystis sp. PCC6803 is capable of both phototrophic growth by oxygenic photosynthesis during light periods and heterotrophic growth by glycolysis and oxidative phosphorylation during dark periods. Gene expression is regulated by a circadian clock and the organism can effectively anticipate transitions between the light and dark phases.

The Cyano-S1 RNA motif is a conserved RNA structure present in some species of Cyanobacteria. Cyano-S1 RNAs are consistently found upstream of genes encoding ribosomal protein S1, a subunit of the ribosome. Therefore, they are presumed to be ribosomal protein leaders, i.e., cis-regulatory elements to which the ribosomal protein S1 binds, thereby controlling its expression levels.
Iron stress repressed RNA (IsrR) is a cis-encoded antisense RNA which regulates the expression of the photosynthetic protein isiA. IsiA expression is activated by the Ferric uptake regulator protein (Fur) under iron stress conditions. IsiA enhances photosynthesis by forming a ring around photosystem I which acts as an additional antenna complex.

The Cyano-2 RNA motif is a conserved RNA structure identified by bioinformatics. Cyano-2 RNAs are found in Cyanobacterial species classified within the genus Synechococcus. Many terminal loops in the two conserved stem-loops contain the nucleotide sequence GCGA, and these sequences might in some cases form stable GNRA tetraloops. Since the two stem-loops are somewhat distant from one another it is possible that they represent two independent non-coding RNAs that are often or always co-transcribed. The region one thousand base pairs upstream of predicted Cyano-2 RNAs is usually devoid of annotated features such as RNA or protein-coding genes. This absence of annotated genes within one thousand base pairs is relatively unusual within bacteria.

The Downstream-peptide motif refers to a conserved RNA structure identified by bioinformatics in the cyanobacterial genera Synechococcus and Prochlorococcus and one phage that infects such bacteria. It was also detected in marine samples of DNA from uncultivated bacteria, which are presumably other species of cyanobacteria.

The glutamine riboswitch is a conserved RNA structure that was predicted by bioinformatics. It is present in a variety of lineages of cyanobacteria, as well as some phages that infect cyanobacteria. It is also found in DNA extracted from uncultivated bacteria living in the ocean that are presumably species of cyanobacteria.

The manA RNA motif refers to a conserved RNA structure that was identified by bioinformatics. Instances of the manA RNA motif were detected in bacteria in the genus Photobacterium and phages that infect certain kinds of cyanobacteria. However, most predicted manA RNA sequences are derived from DNA collected from uncultivated marine bacteria. Almost all manA RNAs are positioned such that they might be in the 5' untranslated regions of protein-coding genes, and therefore it was hypothesized that manA RNAs function as cis-regulatory elements. Given the relative complexity of their secondary structure, and their hypothesized cis-regulatory role, they might be riboswitches.
The wcaG RNA motif is an RNA structure conserved in some bacteria that was detected by bioinformatics. wcaG RNAs are found in certain phages that infect cyanobacteria. Most known wcaG RNAs were found in sequences of DNA extracted from uncultivated marine bacteria. wcaG RNAs might function as cis-regulatory elements, in view of their consistent location in the possible 5' untranslated regions of genes. It was suggested the wcaG RNAs might further function as riboswitches.
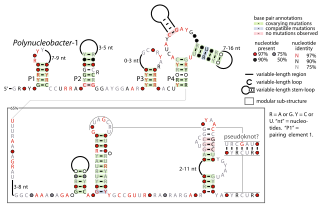
The Polynucleobacter-1 RNA motif is a conserved RNA structure that was identified by bioinformatics. The RNA structure is predominantly located in genome sequences derived from DNA extracted from uncultivated marine samples. However it was also predicted in the genome of Polynucleobacter species QLW-P1DMWA-1, a kind of betaproteobacteria. The RNAs are often located near to a conserved gene that might be homologous to a gene found in a phage that infects cyanobacteria. However, it is unknown if the RNA is used by phages.
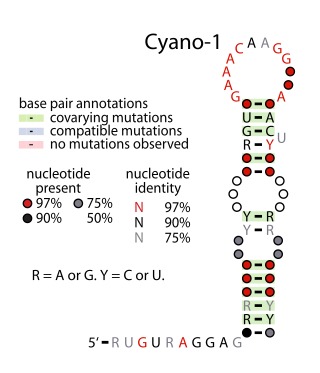
Yfr2 is a family of non-coding RNAs. Members of the Yrf2 family have been identified in almost all studied species of cyanobacteria. The family was identified through a bioinformatics screen of published cyanobacterial genomes, having previously been grouped in a family of Yfr2–5.
PtaRNA1 is a family of non-coding RNAs. Homologs of PtaRNA1 can be found in the bacterial families, Betaproteobacteria and Gammaproteobacteria. In all cases the PtaRNA1 is located anti-sense to a short protein-coding gene. In Xanthomonas campestris pv. vesicatoria, this gene is annotated as XCV2162 and is included in the plasmid toxin family of proteins.
Bacterial small RNAs (bsRNA) are small RNAs produced by bacteria; they are 50- to 500-nucleotide non-coding RNA molecules, highly structured and containing several stem-loops. Numerous sRNAs have been identified using both computational analysis and laboratory-based techniques such as Northern blotting, microarrays and RNA-Seq in a number of bacterial species including Escherichia coli, the model pathogen Salmonella, the nitrogen-fixing alphaproteobacterium Sinorhizobium meliloti, marine cyanobacteria, Francisella tularensis, Streptococcus pyogenes, the pathogen Staphylococcus aureus, and the plant pathogen Xanthomonas oryzae pathovar oryzae. Bacterial sRNAs affect how genes are expressed within bacterial cells via interaction with mRNA or protein, and thus can affect a variety of bacterial functions like metabolism, virulence, environmental stress response, and structure.
αr9 is a family of bacterial small non-coding RNAs with representatives in a broad group of α-proteobacteria from the order Hyphomicrobiales. The first member of this family (Smr9C) was found in a Sinorhizobium meliloti 1021 locus located in the chromosome (C). Further homology and structure conservation analysis have identified full-length Smr9C homologs in several nitrogen-fixing symbiotic rhizobia, in the plant pathogens belonging to Agrobacterium species as well as in a broad spectrum of Brucella species. αr9C RNA species are 144-158 nt long and share a well defined common secondary structure consisting of seven conserved regions. Most of the αr9 transcripts can be catalogued as trans-acting sRNAs expressed from well-defined promoter regions of independent transcription units within intergenic regions (IGRs) of the α-proteobacterial genomes.
αr14 is a family of bacterial small non-coding RNAs with representatives in a broad group of α-proteobacteria. The first member of this family (Smr14C2) was found in a Sinorhizobium meliloti 1021 locus located in the chromosome (C). It was later renamed NfeR1 and shown to be highly expressed in salt stress and during the symbiotic interaction on legume roots. Further homology and structure conservation analysis identified 2 other chromosomal copies and 3 plasmidic ones. Moreover, full-length Smr14C homologs have been identified in several nitrogen-fixing symbiotic rhizobia, in the plant pathogens belonging to Agrobacterium species as well as in a broad spectrum of Brucella species. αr14C RNA species are 115-125 nt long and share a well defined common secondary structure. Most of the αr14 transcripts can be catalogued as trans-acting sRNAs expressed from well-defined promoter regions of independent transcription units within intergenic regions (IGRs) of the α-proteobacterial genomes.
αr15 is a family of bacterial small non-coding RNAs with representatives in a broad group of α-proteobacteria from the order Rhizobiales. The first members of this family were found tandemly arranged in the same intergenic region (IGR) of the Sinorhizobium meliloti 1021 chromosome (C). Further homology and structure conservation analysis have identified full-length Smr15C1 and Smr15C2 homologs in several nitrogen-fixing symbiotic rhizobia, in the plant pathogens belonging to Agrobacterium species as well as in a broad spectrum of Brucella species. The Smr15C1 and Smr15C2 homologs are also encoded in tandem within the same IGR region of Rhizobium and Agrobacterium species, whereas in Brucella species the αr15C loci are spread in the IGRs of Chromosome I. Moreover, this analysis also identified a third αr15 loci in extrachromosomal replicons of the mentioned nitrogen-fixing α-proteobacteria and in the Chromosome II of Brucella species. αr15 RNA species are 99-121 nt long and share a well defined common secondary structure consisting of three stem loops. The transcripts of the αr15 family can be catalogued as trans-acting sRNAs encoded by independent transcription units with recognizable promoter and transcription termination signatures within intergenic regions (IGRs) of the α-proteobacterial genomes.
In molecular biology, Cyanobacterial non-coding RNAs are non-coding RNAs which have been identified in species of cyanobacteria. Large scale screens have identified 21 Yfr in the marine cyanobacterium Prochlorococcus and related species such as Synechococcus. These include the Yfr1 and Yfr2 RNAs. In Prochlorococcus and Synechocystis, non-coding RNAs have been shown to regulate gene expression. NsiR4, widely conserved throughout the cyanobacterial phylum, has been shown to be involved in nitrogen assimilation control in Synechocystis sp. PCC 6803 and in the filamentous, nitrogen-fixing Anabaena sp. PCC 7120.
Non-coding RNAs have been discovered using both experimental and bioinformatic approaches. Bioinformatic approaches can be divided into three main categories. The first involves homology search, although these techniques are by definition unable to find new classes of ncRNAs. The second category includes algorithms designed to discover specific types of ncRNAs that have similar properties. Finally, some discovery methods are based on very general properties of RNA, and are thus able to discover entirely new kinds of ncRNAs.
Microbial DNA barcoding is the use of DNA metabarcoding to characterize a mixture of microorganisms. DNA metabarcoding is a method of DNA barcoding that uses universal genetic markers to identify DNA of a mixture of organisms.
DIMPL is a bioinformatic pipeline that enables the extraction and selection of bacterial GC-rich intergenic regions (IGRs) that are enriched for structured non-coding RNAs (ncRNAs). The method of enriching bacterial IGRs for ncRNA motif discovery was first reported for a study in "Genome-wide discovery of structured noncoding RNAs in bacteria".









