
Transfer-messenger RNA is a bacterial RNA molecule with dual tRNA-like and messenger RNA-like properties. The tmRNA forms a ribonucleoprotein complex (tmRNP) together with Small Protein B (SmpB), Elongation Factor Tu (EF-Tu), and ribosomal protein S1. In trans-translation, tmRNA and its associated proteins bind to bacterial ribosomes which have stalled in the middle of protein biosynthesis, for example when reaching the end of a messenger RNA which has lost its stop codon. The tmRNA is remarkably versatile: it recycles the stalled ribosome, adds a proteolysis-inducing tag to the unfinished polypeptide, and facilitates the degradation of the aberrant messenger RNA. In the majority of bacteria these functions are carried out by standard one-piece tmRNAs. In other bacterial species, a permuted ssrA gene produces a two-piece tmRNA in which two separate RNA chains are joined by base-pairing.
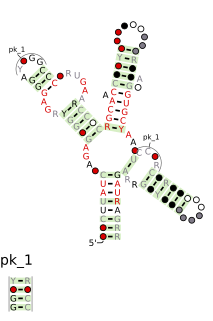
The SAM riboswitch is found upstream of a number of genes which code for proteins involved in methionine or cysteine biosynthesis in Gram-positive bacteria. Two SAM riboswitches in Bacillus subtilis that were experimentally studied act at the level of transcription termination control. The predicted secondary structure consists of a complex stem-loop region followed by a single stem-loop terminator region. An alternative and mutually exclusive form involves bases in the 3' segment of helix 1 with those in the 5' region of helix 5 to form a structure termed the anti-terminator form. When SAM is unbound, the anti-terminator sequence sequesters the terminator sequence so the terminator is unable to form, allowing the polymerase read-through the downstream gene. When S-Adenosyl methionine (SAM) is bound to the aptamer, the anti-terminator is sequestered by an anti-anti-terminator; the terminator forms and terminates the transcription. However, many SAM riboswitches are likely to regulate gene expression at the level of translation.

Usually found in gram-positive bacteria, the T box leader sequence is an RNA element that controls gene expression through the regulation of translation by binding directly to a specific tRNA and sensing its aminoacylation state. This interaction controls expression of downstream aminoacyl-tRNA synthetase genes, amino acid biosynthesis, and uptake-related genes in a negative feedback loop. The uncharged tRNA acts as the effector for transcription antitermination of genes in the T-box leader family. The anticodon of a specific tRNA base pairs to a specifier sequence within the T-box motif, and the NCCA acceptor tail of the tRNA base pairs to a conserved bulge in the T-box antiterminator hairpin.

The ykkC/yxkD leader is a conserved RNA structure found upstream of the ykkC and yxkD genes in Bacillus subtilis and related genes in other bacteria. The function of this family is unclear for many years although it has been suggested that it may function to switch on efflux pumps and detoxification systems in response to harmful environmental molecules. The Thermoanaerobacter tengcongensis sequence AE013027 overlaps with that of purine riboswitch suggesting that the two riboswitches may work in conjunction to regulate the upstream gene which codes for TTE0584 (Q8RC62), a member of the permease family.
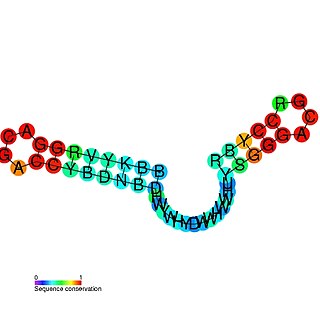
The mini-ykkC RNA motif was discovered as a putative RNA structure that is conserved in bacteria. The motif consists of two conserved stem-loops whose terminal loops contain the RNA sequence ACGR, where R represents either A or G. Mini-ykkC RNAs are widespread in Proteobacteria, but some are predicted in other phyla of bacteria. It was expected that the RNAs are cis-regulatory elements, because they are typically located upstream of protein-coding genes.

The Cyano-S1 RNA motif is a conserved RNA structure present in some species of Cyanobacteria. Cyano-S1 RNAs are consistently found upstream of genes encoding ribosomal protein S1, a subunit of the ribosome. Therefore, they are presumed to be ribosomal protein leaders, i.e., cis-regulatory elements to which the ribosomal protein S1 binds, thereby controlling its expression levels.
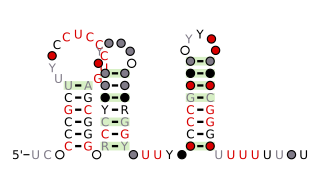
The anti-hemB RNA motif is a conserved RNA structure that was found in all known bacteria in the genus Burkholderia, and in a variety of other betaproteobacteria. The anti-hemB RNA motif consists primarily of two stem-loops, followed by a predicted rho-independent transcription termination stem-loop. As anti-hemB RNAs are generally not located in a 5' UTR, the RNAs are presumed to be non-coding RNAs. The terminator stem-loop implies that anti-hemB RNAs are transcribed as independent molecules.
The lacto-2 RNA motif is an RNA structure that is conserved amongst bacteria within the order Lactobacillales. The motif consists of a stem-loop whose stem is interrupted by many internal loops and bulges. Nucleotide identities in many places are conserved, and one internal loop in particular is highly conserved.

Yfr1 is a Cyanobacterial functional RNA that was identified by a comparative genome based screen for RNAs in cyanobacteria. Further analysis has shown that the RNA is well conserved and highly expressed in cyanobacteria. and is required for growth under several stress condition Bioinformatics research combined with follow-up experiments have shown that Yfr1 inhibits the translation of the proteins PMM1119 and PMM1121 by an antisense interaction by base pairing at the ribosomal binding site.

The Collinsella-1 RNA motif denotes a particular conserved RNA structure discovered by bioinformatics. Of the six sequences belonging to this motif that were originally identified, five are from uncultivated bacteria residing in the human gut, while only the sixth is in a cultivated species, Collinsella aerofaciens. The evidence supporting the stem-loops designated as "P1" and "P2" is ambiguous.

The Downstream-peptide motif refers to a conserved RNA structure identified by bioinformatics in the cyanobacterial genera Synechococcus and Prochlorococcus and one phage that infects such bacteria. It was also detected in marine samples of DNA from uncultivated bacteria, which are presumably other species of cyanobacteria.

The gabT RNA motif is the name of a conserved RNA structure identified by bioinformatics whose function is unknown. The gabT motif has been detected exclusively in bacteria within the genus Pseudomonas, and is found only upstream of gabT genes, and downstream to gabD genes.

The mraW RNA motif is a conserved, structured RNA found in certain bacteria. Specifically, it is predicted in many, though not all, species of actinobacteria, and especially within the genus Mycobacterium. Structurally, the motif consists of a hairpin with a highly conserved terminal loop sequence. mraW RNAs are consistently in the presumed 5' untranslated regions of mraW genes. These mraW genes likely form operons with immediately downstream ftsI genes, and multiple types of mur genes. These genes are associated with peptidoglycan synthesis, and it was hypothesized that the mraW RNA motif might regulate these genes.

The msiK RNA motif describes a conserved RNA structure discovered using bioinformatics. The RNA is always found in the presumed 5' untranslated regions of genes annotated as msiK, and is therefore hypothesized to be an RNA-based cis-regulatory element that regulates these genes.

The pan RNA motif defines a conserved RNA structure that was identified using bioinformatics. pan motif RNAs are present in three phyla: Chloroflexi, Firmicutes and Proteobacteria, although within the latter phylum they are only known in deltaproteobacteria. A pan RNA is present in the Firmicute Bacillus subtilis, which is one of the most extensively studied bacteria.

PhotoRC RNA motifs refer to conserved RNA structures that are associated with genes acting in the photosynthetic reaction centre of photosynthetic bacteria. Two such RNA classes were identified and called the PhotoRC-I and PhotoRC-II motifs. PhotoRC-I RNAs were detected in the genomes of some cyanobacteria. Although no PhotoRC-II RNA has been detected in cyanobacteria, one is found in the genome of a purified phage that infects cyanobacteria. Both PhotoRC-I and PhotoRC-II RNAs are present in sequences derived from DNA that was extracted from uncultivated marine bacteria.

The psaA RNA motif describes a class of RNAs with a common secondary structure. psaA RNAs are exclusively found in locations that presumably correspond to the 5' untranslated regions of operons formed of psaA and psaB genes. For this reason, it was hypothesized that psaA RNAs function as cis-regulatory elements of these genes. The psaAB genes encode proteins that form subunits in the photosystem I structure used for photosynthesis. psaA RNAs have been detected only in cyanobacteria, which is consistent with their association with photosynthesis.

Yfr2 is a family of non-coding RNAs. Members of the Yrf2 family have been identified in almost all studied species of cyanobacteria. The family was identified through a bioinformatics screen of published cyanobacterial genomes, having previously been grouped in a family of Yfr2–5.
αr35 is a family of bacterial small non-coding RNAs with representatives in a reduced group of α-proteobacteria from the order Hyphomicrobiales. The first member of this family (Smr35B) was found in a Sinorhizobium meliloti 1021 locus located in the symbiotic plasmid B (pSymB). Further homology and structure conservation analysis have identified full-length SmrB35 homologs in other legume symbionts, as well as in the human and plant pathogens Ochrobactrum anthropi and Agrobacterium tumefaciens, respectively. αr35 RNA species are 139-142 nt long and share a common secondary structure consisting of two stem loops and a well conserved rho independent terminator. Most of the αr35 transcripts can be catalogued as trans-acting sRNAs expressed from well-defined promoter regions of independent transcription units within intergenic regions of the α-proteobacterial genomes.
LOC101928193 is a protein which in humans is encoded by the LOC101928193 gene. There are no known aliases for this gene or protein. Similar copies of this gene, called orthologs, are known to exist in several different species across mammals, amphibians, fish, mollusks, cnidarians, fungi, and bacteria. The human LOC101928193 gene is located on the long (q) arm of chromosome 9 with a cytogenic location at 9q34.2. The molecular location of the gene is from base pair 133,189,767 to base pair 133,192,979 on chromosome 9 for an mRNA length of 3213 nucleotides. The gene and protein are not yet well understood by the scientific community, but there is data on its genetic makeup and expression. The LOC101928193 protein is targeted for the cytoplasm and has the highest level of expression in the thyroid, ovary, skin, and testes in humans.

















