Related Research Articles

A protein kinase is a kinase which selectively modifies other proteins by covalently adding phosphates to them (phosphorylation) as opposed to kinases which modify lipids, carbohydrates, or other molecules. Phosphorylation usually results in a functional change of the target protein (substrate) by changing enzyme activity, cellular location, or association with other proteins. The human genome contains about 500 protein kinase genes and they constitute about 2% of all human genes. There are two main types of protein kinase. The great majority are serine/threonine kinases, which phosphorylate the hydroxyl groups of serines and threonines in their targets. Most of the others are tyrosine kinases, although additional types exist. Protein kinases are also found in bacteria and plants. Up to 30% of all human proteins may be modified by kinase activity, and kinases are known to regulate the majority of cellular pathways, especially those involved in signal transduction.

The enzyme-linked immunosorbent assay (ELISA) is a commonly used analytical biochemistry assay, first described by Eva Engvall and Peter Perlmann in 1971. The assay uses a solid-phase type of enzyme immunoassay (EIA) to detect the presence of a ligand in a liquid sample using antibodies directed against the protein to be measured. ELISA has been used as a diagnostic tool in medicine, plant pathology, and biotechnology, as well as a quality control check in various industries.

In biochemistry, allosteric regulation is the regulation of an enzyme by binding an effector molecule at a site other than the enzyme's active site.
A biosensor is an analytical device, used for the detection of a chemical substance, that combines a biological component with a physicochemical detector. The sensitive biological element, e.g. tissue, microorganisms, organelles, cell receptors, enzymes, antibodies, nucleic acids, etc., is a biologically derived material or biomimetic component that interacts with, binds with, or recognizes the analyte under study. The biologically sensitive elements can also be created by biological engineering. The transducer or the detector element, which transforms one signal into another one, works in a physicochemical way: optical, piezoelectric, electrochemical, electrochemiluminescence etc., resulting from the interaction of the analyte with the biological element, to easily measure and quantify. The biosensor reader device connects with the associated electronics or signal processors that are primarily responsible for the display of the results in a user-friendly way. This sometimes accounts for the most expensive part of the sensor device, however it is possible to generate a user friendly display that includes transducer and sensitive element. The readers are usually custom-designed and manufactured to suit the different working principles of biosensors.

In biochemistry and molecular biology, a binding site is a region on a macromolecule such as a protein that binds to another molecule with specificity. The binding partner of the macromolecule is often referred to as a ligand. Ligands may include other proteins, enzyme substrates, second messengers, hormones, or allosteric modulators. The binding event is often, but not always, accompanied by a conformational change that alters the protein's function. Binding to protein binding sites is most often reversible, but can also be covalent reversible or irreversible.

In biochemistry and pharmacology, receptors are chemical structures, composed of protein, that receive and transduce signals that may be integrated into biological systems. These signals are typically chemical messengers which bind to a receptor and produce physiological responses such as change in the electrical activity of a cell. For example, GABA, an inhibitory neurotransmitter, inhibits electrical activity of neurons by binding to GABAA receptors. There are three main ways the action of the receptor can be classified: relay of signal, amplification, or integration. Relaying sends the signal onward, amplification increases the effect of a single ligand, and integration allows the signal to be incorporated into another biochemical pathway.

A receptor antagonist is a type of receptor ligand or drug that blocks or dampens a biological response by binding to and blocking a receptor rather than activating it like an agonist. Antagonist drugs interfere in the natural operation of receptor proteins. They are sometimes called blockers; examples include alpha blockers, beta blockers, and calcium channel blockers. In pharmacology, antagonists have affinity but no efficacy for their cognate receptors, and binding will disrupt the interaction and inhibit the function of an agonist or inverse agonist at receptors. Antagonists mediate their effects by binding to the active site or to the allosteric site on a receptor, or they may interact at unique binding sites not normally involved in the biological regulation of the receptor's activity. Antagonist activity may be reversible or irreversible depending on the longevity of the antagonist–receptor complex, which, in turn, depends on the nature of antagonist–receptor binding. The majority of drug antagonists achieve their potency by competing with endogenous ligands or substrates at structurally defined binding sites on receptors.
In biochemistry, biotinylation is the process of covalently attaching biotin to a protein, nucleic acid or other molecule. Biotinylation is rapid, specific and is unlikely to disturb the natural function of the molecule due to the small size of biotin. Biotin binds to streptavidin and avidin with an extremely high affinity, fast on-rate, and high specificity, and these interactions are exploited in many areas of biotechnology to isolate biotinylated molecules of interest. Biotin-binding to streptavidin and avidin is resistant to extremes of heat, pH and proteolysis, making capture of biotinylated molecules possible in a wide variety of environments. Also, multiple biotin molecules can be conjugated to a protein of interest, which allows binding of multiple streptavidin, avidin or neutravidin protein molecules and increases the sensitivity of detection of the protein of interest. There is a large number of biotinylation reagents available that exploit the wide range of possible labelling methods. Due to the strong affinity between biotin and streptavidin, the purification of biotinylated proteins has been a widely used approach to identify protein-protein interactions and post-translational events such as ubiquitylation in molecular biology.
An immunoassay (IA) is a biochemical test that measures the presence or concentration of a macromolecule or a small molecule in a solution through the use of an antibody (usually) or an antigen (sometimes). The molecule detected by the immunoassay is often referred to as an "analyte" and is in many cases a protein, although it may be other kinds of molecules, of different sizes and types, as long as the proper antibodies that have the required properties for the assay are developed. Analytes in biological liquids such as serum or urine are frequently measured using immunoassays for medical and research purposes.
Affinity chromatography is a method of separating a biomolecule from a mixture, based on a highly specific macromolecular binding interaction between the biomolecule and another substance. The specific type of binding interaction depends on the biomolecule of interest; antigen and antibody, enzyme and substrate, receptor and ligand, or protein and nucleic acid binding interactions are frequently exploited for isolation of various biomolecules. Affinity chromatography is useful for its high selectivity and resolution of separation, compared to other chromatographic methods.

Half maximal inhibitory concentration (IC50) is a measure of the potency of a substance in inhibiting a specific biological or biochemical function. IC50 is a quantitative measure that indicates how much of a particular inhibitory substance (e.g. drug) is needed to inhibit, in vitro, a given biological process or biological component by 50%. The biological component could be an enzyme, cell, cell receptor or microorganism. IC50 values are typically expressed as molar concentration.

Chemogenomics, or chemical genomics, is the systematic screening of targeted chemical libraries of small molecules against individual drug target families with the ultimate goal of identification of novel drugs and drug targets. Typically some members of a target library have been well characterized where both the function has been determined and compounds that modulate the function of those targets have been identified. Other members of the target family may have unknown function with no known ligands and hence are classified as orphan receptors. By identifying screening hits that modulate the activity of the less well characterized members of the target family, the function of these novel targets can be elucidated. Furthermore, the hits for these targets can be used as a starting point for drug discovery. The completion of the human genome project has provided an abundance of potential targets for therapeutic intervention. Chemogenomics strives to study the intersection of all possible drugs on all of these potential targets.
In biochemistry, avidity refers to the accumulated strength of multiple affinities of individual non-covalent binding interactions, such as between a protein receptor and its ligand, and is commonly referred to as functional affinity. Avidity differs from affinity, which describes the strength of a single interaction. However, because individual binding events increase the likelihood of occurrence of other interactions, avidity should not be thought of as the mere sum of its constituent affinities but as the combined effect of all affinities participating in the biomolecular interaction. A particular important aspect relates to the phenomenon of 'avidity entropy'. Biomolecules often form heterogenous complexes or homogeneous oligomers and multimers or polymers. If clustered proteins form an organized matrix, such as the clathrin-coat, the interaction is described as a matricity.

In pharmacology, Schild regression analysis, based upon the Schild equation, both named for Heinz Otto Schild, are tools for studying the effects of agonists and antagonists on the response caused by the receptor or on ligand-receptor binding.
Molecular binding is an attractive interaction between two molecules that results in a stable association in which the molecules are in close proximity to each other. It is formed when atoms or molecules bind together by sharing of electrons. It often, but not always, involves some chemical bonding.
The estrogen receptor test (ERT) is a laboratory test to determine whether cancer cells have estrogen receptors. This information can guide treatment of the cancer.
A ligand binding assay (LBA) is an assay, or an analytic procedure, which relies on the binding of ligand molecules to receptors, antibodies or other macromolecules. A detection method is used to determine the presence and extent of the ligand-receptor complexes formed, and this is usually determined electrochemically or through a fluorescence detection method. This type of analytic test can be used to test for the presence of target molecules in a sample that are known to bind to the receptor.
Lanthanide probes are a non-invasive analytical tool commonly used for biological and chemical applications. Lanthanides are metal ions which have their 4f energy level filled and generally refer to elements cerium to lutetium in the periodic table. The fluorescence of lanthanide salts is weak because the energy absorption of the metallic ion is low; hence chelated complexes of lanthanides are most commonly used. The term chelate derives from the Greek word for “claw,” and is applied to name ligands, which attach to a metal ion with two or more donor atoms through dative bonds. The fluorescence is most intense when the metal ion has the oxidation state of 3+. Not all lanthanide metals can be used and the most common are: Sm(III), Eu(III), Tb(III), and Dy(III).
Chemoproteomics entails a broad array of techniques used to identify and interrogate protein-small molecule interactions. Chemoproteomics complements phenotypic drug discovery, a paradigm that aims to discover lead compounds on the basis of alleviating a disease phenotype, as opposed to target-based drug discovery, in which lead compounds are designed to interact with predetermined disease-driving biological targets. As phenotypic drug discovery assays do not provide confirmation of a compound's mechanism of action, chemoproteomics provides valuable follow-up strategies to narrow down potential targets and eventually validate a molecule's mechanism of action. Chemoproteomics also attempts to address the inherent challenge of drug promiscuity in small molecule drug discovery by analyzing protein-small molecule interactions on a proteome-wide scale. A major goal of chemoproteomics is to characterize the interactome of drug candidates to gain insight into mechanisms of off-target toxicity and polypharmacology.
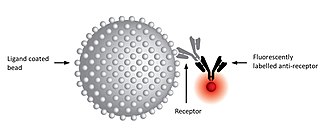
A kinetic exclusion assay (KinExA) is a type of bioassay in which a solution containing receptor, ligand, and receptor-ligand complex is briefly exposed to additional ligand immobilized on a solid phase.
References
- ↑ Tijssen, P. (1 August 1985). Practice and Theory of Enzyme Immunoassays. Elsevier. p. 382. ISBN 9780080858845 . Retrieved 22 January 2015.
In the homogeneous assay, an enzyme modulator (antibody, inhibitor or receptor for the enzyme) is covalently linked to the ligand which competes with free ligand from the test sample for the available antibodies.