Natural selection is the differential survival and reproduction of individuals due to differences in phenotype. It is a key mechanism of evolution, the change in the heritable traits characteristic of a population over generations. Charles Darwin popularised the term "natural selection", contrasting it with artificial selection, which is intentional, whereas natural selection is not.
Quantum evolution is a component of George Gaylord Simpson's multi-tempoed theory of evolution proposed to explain the rapid emergence of higher taxonomic groups in the fossil record. According to Simpson, evolutionary rates differ from group to group and even among closely related lineages. These different rates of evolutionary change were designated by Simpson as bradytelic, horotelic, and tachytelic.
Molecular evolution is the process of change in the sequence composition of cellular molecules such as DNA, RNA, and proteins across generations. The field of molecular evolution uses principles of evolutionary biology and population genetics to explain patterns in these changes. Major topics in molecular evolution concern the rates and impacts of single nucleotide changes, neutral evolution vs. natural selection, origins of new genes, the genetic nature of complex traits, the genetic basis of speciation, the evolution of development, and ways that evolutionary forces influence genomic and phenotypic changes.
Population genetics is a subfield of genetics that deals with genetic differences within and among populations, and is a part of evolutionary biology. Studies in this branch of biology examine such phenomena as adaptation, speciation, and population structure.
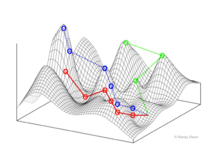
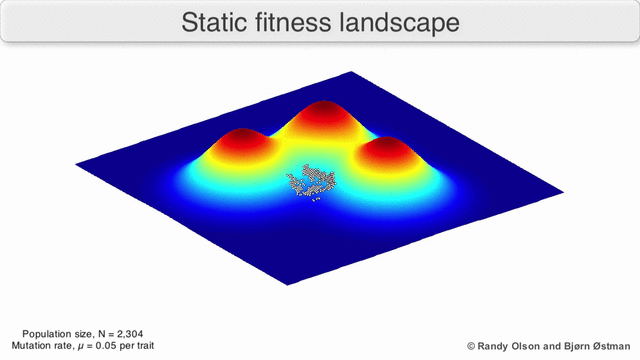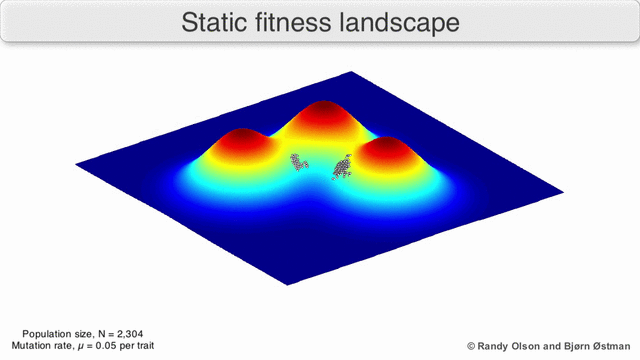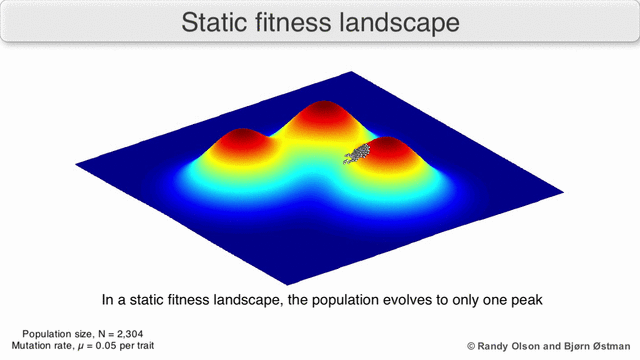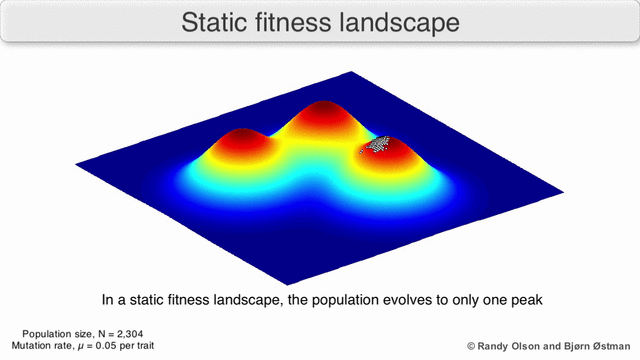
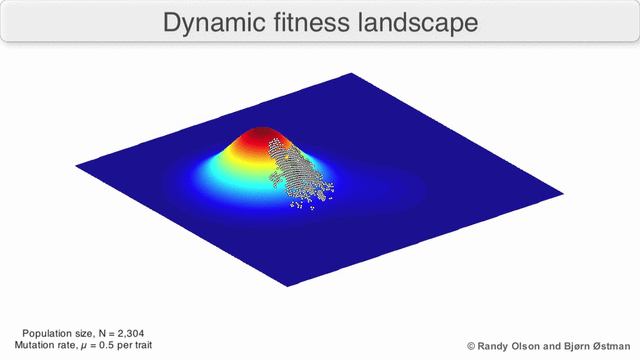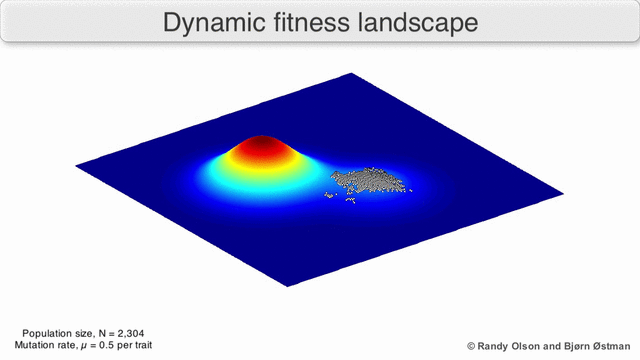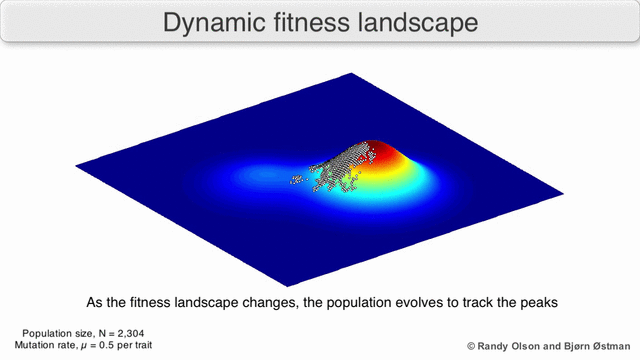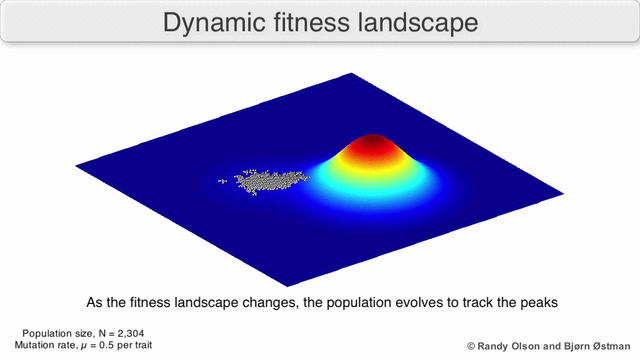
In evolutionary biology, fitness landscapes or adaptive landscapes are used to visualize the relationship between genotypes and reproductive success. It is assumed that every genotype has a well-defined replication rate. This fitness is the "height" of the landscape. Genotypes which are similar are said to be "close" to each other, while those that are very different are "far" from each other. The set of all possible genotypes, their degree of similarity, and their related fitness values is then called a fitness landscape. The idea of a fitness landscape is a metaphor to help explain flawed forms in evolution by natural selection, including exploits and glitches in animals like their reactions to supernormal stimuli.

This is a list of topics in evolutionary biology.
Evolvability is defined as the capacity of a system for adaptive evolution. Evolvability is the ability of a population of organisms to not merely generate genetic diversity, but to generate adaptive genetic diversity, and thereby evolve through natural selection.

In evolutionary biology, the Baldwin effect describes an effect of learned behaviour on evolution. James Mark Baldwin and others suggested that an organism's ability to learn new behaviours will affect its reproductive success and will therefore have an effect on the genetic makeup of its species through natural selection. It posits that subsequent selection might reinforce the originally learned behaviors, if adaptive, into more in-born, instinctive ones. Though this process appears similar to Lamarckism, that view proposes that living things inherited their parents' acquired characteristics. The Baldwin effect only posits that learning ability, which is genetically based, is another variable in / contributor to environmental adaptation. First proposed during the Eclipse of Darwinism in the late 19th century, this effect has been independently proposed several times, and today it is generally recognized as part of the modern synthesis.

Pleiotropy occurs when one gene influences two or more seemingly unrelated phenotypic traits. Such a gene that exhibits multiple phenotypic expression is called a pleiotropic gene. Mutation in a pleiotropic gene may have an effect on several traits simultaneously, due to the gene coding for a product used by a myriad of cells or different targets that have the same signaling function.
Evolutionary capacitance is the storage and release of variation, just as electric capacitors store and release charge. Living systems are robust to mutations. This means that living systems accumulate genetic variation without the variation having a phenotypic effect. But when the system is disturbed, robustness breaks down, and the variation has phenotypic effects and is subject to the full force of natural selection. An evolutionary capacitor is a molecular switch mechanism that can "toggle" genetic variation between hidden and revealed states. If some subset of newly revealed variation is adaptive, it becomes fixed by genetic assimilation. After that, the rest of variation, most of which is presumably deleterious, can be switched off, leaving the population with a newly evolved advantageous trait, but no long-term handicap. For evolutionary capacitance to increase evolvability in this way, the switching rate should not be faster than the timescale of genetic assimilation.
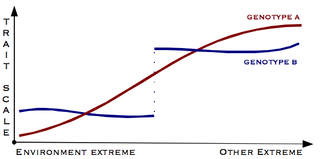
Canalisation is a measure of the ability of a population to produce the same phenotype regardless of variability of its environment or genotype. It is a form of evolutionary robustness. The term was coined in 1942 by C. H. Waddington to capture the fact that "developmental reactions, as they occur in organisms submitted to natural selection...are adjusted so as to bring about one definite end-result regardless of minor variations in conditions during the course of the reaction". He used this word rather than robustness to consider that biological systems are not robust in quite the same way as, for example, engineered systems.

The Neutral Theory of Molecular Evolution is an influential monograph written in 1983 by Japanese evolutionary biologist Motoo Kimura. While the neutral theory of molecular evolution existed since his article in 1968, Kimura felt the need to write a monograph with up-to-date information and evidences showing the importance of his theory in evolution.
Neutral mutations are changes in DNA sequence that are neither beneficial nor detrimental to the ability of an organism to survive and reproduce. In population genetics, mutations in which natural selection does not affect the spread of the mutation in a species are termed neutral mutations. Neutral mutations that are inheritable and not linked to any genes under selection will be lost or will replace all other alleles of the gene. That loss or fixation of the gene proceeds based on random sampling known as genetic drift. A neutral mutation that is in linkage disequilibrium with other alleles that are under selection may proceed to loss or fixation via genetic hitchhiking and/or background selection.
Genetic assimilation is a process described by Conrad H. Waddington by which a phenotype originally produced in response to an environmental condition, such as exposure to a teratogen, later becomes genetically encoded via artificial selection or natural selection. Despite superficial appearances, this does not require the (Lamarckian) inheritance of acquired characters, although epigenetic inheritance could potentially influence the result. Waddington stated that genetic assimilation overcomes the barrier to selection imposed by what he called canalization of developmental pathways; he supposed that the organism's genetics evolved to ensure that development proceeded in a certain way regardless of normal environmental variations.

The shifting balance theory is a theory of evolution proposed in 1932 by Sewall Wright, suggesting that adaptive evolution may proceed most quickly when a population divides into subpopulations with restricted gene flow. The name of the theory is borrowed from Wright's metaphor of fitness landscapes, attempting to explain how a population may move across an adaptive valley to a higher adaptive peak. According to the theory, this movement occurs in three steps:
- Genetic drift allows a locally adapted subpopulation to move across an adaptive valley to the base of a higher adaptive peak.
- Natural selection will move the subpopulation up the higher peak.
- This new superiorly adapted subpopulation may then expand its range and outcompete or interbreed with other subpopulations, causing the spread of new adaptations and movement of the global population toward the new fitness peak.

Phenotypic Integration is a metric for measuring the correlation of multiple functionally-related traits to each other. Complex phenotypes often require multiple traits working together in order to function properly. Phenotypic integration is significant because it provides an explanation as to how phenotypes are sustained by relationships between traits. Every organism's phenotype is integrated, organized, and a functional whole. Integration is also associated with functional modules. Modules are complex character units that are tightly associated, such as a flower. It is hypothesized that organisms with high correlations between traits in a module have the most efficient functions. The fitness of a particular value for one phenotypic trait frequently depends on the value of the other phenotypic traits, making it important for those traits evolve together. One trait can have a direct effect on fitness, and it has been shown that the correlations among traits can also change fitness, causing these correlations to be adaptive, rather than solely genetic. Integration can be involved in multiple aspects of life, not just at the genetic level, but during development, or simply at a functional level.

Epistasis is a phenomenon in genetics in which the effect of a gene mutation is dependent on the presence or absence of mutations in one or more other genes, respectively termed modifier genes. In other words, the effect of the mutation is dependent on the genetic background in which it appears. Epistatic mutations therefore have different effects on their own than when they occur together. Originally, the term epistasis specifically meant that the effect of a gene variant is masked by that of different gene.
This glossary of genetics and evolutionary biology is a list of definitions of terms and concepts used in the study of genetics and evolutionary biology, as well as sub-disciplines and related fields, with an emphasis on classical genetics, quantitative genetics, population biology, phylogenetics, speciation, and systematics. Overlapping and related terms can be found in Glossary of cellular and molecular biology, Glossary of ecology, and Glossary of biology.
In evolutionary biology, developmental bias refers to the production against or towards certain ontogenetic trajectories which ultimately influence the direction and outcome of evolutionary change by affecting the rates, magnitudes, directions and limits of trait evolution. Historically, the term was synonymous with developmental constraint, however, the latter has been more recently interpreted as referring solely to the negative role of development in evolution.
Bias in the introduction of variation is a theory in the domain of evolutionary biology that asserts biases in the introduction of heritable variation are reflected in the outcome of evolution. It is relevant to topics in molecular evolution, evo-devo, and self-organization. In the context of this theory, "introduction" ("origination") is a technical term for events that shift an allele frequency upward from zero. Formal models demonstrate that when an evolutionary process depends on introduction events, mutational and developmental biases in the generation of variation may influence the course of evolution by a first come, first served effect, so that evolution reflects the arrival of the likelier, not just the survival of the fitter. Whereas mutational explanations for evolutionary patterns are typically assumed to imply or require neutral evolution, the theory of arrival biases distinctively predicts the possibility of mutation-biased adaptation. Direct evidence for the theory comes from laboratory studies showing that adaptive changes are systematically enriched for mutationally likely types of changes. Retrospective analyses of natural cases of adaptation also provide support for the theory. This theory is notable as an example of contemporary structuralist thinking, contrasting with a classical functionalist view in which the course of evolution is determined by natural selection.