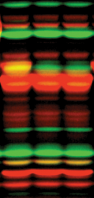
A protease is an enzyme that catalyzes proteolysis, breaking down proteins into smaller polypeptides or single amino acids, and spurring the formation of new protein products. They do this by cleaving the peptide bonds within proteins by hydrolysis, a reaction where water breaks bonds. Proteases are involved in many biological functions, including digestion of ingested proteins, protein catabolism, and cell signaling.
Serine proteases are enzymes that cleave peptide bonds in proteins. Serine serves as the nucleophilic amino acid at the (enzyme's) active site. They are found ubiquitously in both eukaryotes and prokaryotes. Serine proteases fall into two broad categories based on their structure: chymotrypsin-like (trypsin-like) or subtilisin-like.
A metalloproteinase, or metalloprotease, is any protease enzyme whose catalytic mechanism involves a metal. An example is ADAM12 which plays a significant role in the fusion of muscle cells during embryo development, in a process known as myogenesis.
Chemical biology is a scientific discipline spanning the fields of chemistry and biology. The discipline involves the application of chemical techniques, analysis, and often small molecules produced through synthetic chemistry, to the study and manipulation of biological systems. In contrast to biochemistry, which involves the study of the chemistry of biomolecules and regulation of biochemical pathways within and between cells, chemical biology deals with chemistry applied to biology.
In chemical synthesis, "click" chemistry is a class of biocompatible small molecule reactions commonly used in bioconjugation, allowing the joining of substrates of choice with specific biomolecules. Click chemistry is not a single specific reaction, but describes a way of generating products that follow examples in nature, which also generates substances by joining small modular units. In many applications, click reactions join a biomolecule and a reporter molecule. Click chemistry is not limited to biological conditions: the concept of a "click" reaction has been used in chemoproteomic, pharmacological, and various biomimetic applications. However, they have been made notably useful in the detection, localization and qualification of biomolecules.
Affinity labels are a class of enzyme inhibitors that covalently bind to their target causing its inactivation. The hallmark of an affinity label is the use a targeting moiety to specifically and reversibly deliver a weakly reactive group to the enzyme that irreversibly binds to an amino acid residue. The targeting portion of the label often resembles the enzyme's natural substrate so that a similar mode of noncovalent binding is used prior to the covalent linkage. Their usefulness in medicine can be limited by the specificity of the first noncovalent binding step whereas indiscriminate action can be utilized for purposes such as affinity labeling - a technique for the validation of substrate-specific binding of compounds.
Benjamin Franklin Cravatt III is a professor in the Department of Chemistry at The Scripps Research Institute in La Jolla, California. Considered a co-inventor of activity based proteomics and a substantial contributor to research on the endocannabinoid system, he is a prominent figure in the nascent field of chemical biology. Cravatt was elected to the National Academy of Sciences in 2014, and the American Academy of Arts and Sciences in 2016. He is Gilula Chair of Chemical Biology, a Cope Scholar, and a Searle Scholar.

Monoacylglycerol lipase, also known as MAG lipase, acylglycerol lipase, MAGL, MGL or MGLL is an enzyme that, in humans, is encoded by the MGLL gene. MAGL is a 33-kDa, membrane-associated member of the serine hydrolase superfamily and contains the classical GXSXG consensus sequence common to most serine hydrolases. The catalytic triad has been identified as Ser122, His269, and Asp239.

Fatty acid amide hydrolase or FAAH is a member of the serine hydrolase family of enzymes. It was first shown to break down anandamide in 1993. In humans, it is encoded by the gene FAAH.
Serine hydrolases are one of the largest known enzyme classes comprising approximately ~200 enzymes or 1% of the genes in the human proteome. A defining characteristic of these enzymes is the presence of a nucleophilic serine in their active site, which is used for the hydrolysis of substrates. Catalysis proceeds by the formation of an acyl-enzyme intermediate through this serine, followed by water/hydroxide-induced saponification of the intermediate and regeneration of the enzyme. Unlike other non-catalytic serines, the nucleophilic serine of these hydrolases is typically activated by a proton relay involving a catalytic triad consisting of the serine, an acidic residue and a basic residue, although variations on this mechanism exist.
Bioconjugation is a chemical strategy to form a stable covalent link between two molecules, at least one of which is a biomolecule.

Methoxy arachidonyl fluorophosphonate, commonly referred as MAFP, is an irreversible active site-directed enzyme inhibitor that inhibits nearly all serine hydrolases and serine proteases. It inhibits phospholipase A2 and fatty acid amide hydrolase with special potency, displaying IC50 values in the low-nanomolar range. In addition, it binds to the CB1 receptor in rat brain membrane preparations (IC50 = 20 nM), but does not appear to agonize or antagonize the receptor, though some related derivatives do show cannabinoid-like properties.

3-Azidocoumarin is an organic compound that is used in the area of bioconjugation. It is a derivative of coumarin, a natural product and precursor for the widely used Coumadin. Azidocoumarin has emerged as a widely applicable labeling agent in diverse biological systems. In particular, it participates in the aptly named click reaction with alkynes. Bioconjugation involves the labeling of certain cellular components and is applicable to fields such a proteomics and functional genomics with a detachable, fluorescent tag.
The term bioorthogonal chemistry refers to any chemical reaction that can occur inside of living systems without interfering with native biochemical processes. The term was coined by Carolyn R. Bertozzi in 2003. Since its introduction, the concept of the bioorthogonal reaction has enabled the study of biomolecules such as glycans, proteins, and lipids in real time in living systems without cellular toxicity. A number of chemical ligation strategies have been developed that fulfill the requirements of bioorthogonality, including the 1,3-dipolar cycloaddition between azides and cyclooctynes, between nitrones and cyclooctynes, oxime/hydrazone formation from aldehydes and ketones, the tetrazine ligation, the isocyanide-based click reaction, and most recently, the quadricyclane ligation.

Neutral cholesterol ester hydrolase 1 (NCEH) also known as arylacetamide deacetylase-like 1 (AADACL1) or KIAA1363 is an enzyme that in humans is encoded by the NCEH1 gene.
PRIME is a molecular biology research tool developed by Alice Y. Ting and the Ting Lab at MIT for site-specific labeling of proteins in living cells with chemical probes. Probes often have useful biophysical properties, such as fluorescence, and allow imaging of proteins. Ultimately, PRIME enables scientists to study functions of specific proteins of interest.

BCN, also known as bicyclo[6.1.0]nonyne, is a copper-free click chemistry probe that enables highly efficient and completely orthogonal bioconjugation to complex macromolecules including peptides, nucleic acids and proteins, including monoclonal antibodies. The most recent and powerful application of this technology has been in the field of antibody-drug conjugates which results in targeted cancer therapeutics that have an improved therapeutic index, meaning they are more effective and better tolerated. Amongst its most notable features, BCN is ideally suited for aqueous bioconjugations due to its high reactivity with its azide counterpart combined with its high hydrophilicity, relative to all other metal-free click chemistry probes. See also "copper-free click chemistry". Commercial use of BCN is proprietary to Netherlands-based biotechnology company, Synaffix BV.
Chemoproteomics entails a broad array of techniques used to identify and interrogate protein-small molecule interactions. Chemoproteomics complements phenotypic drug discovery, a paradigm that aims to discover lead compounds on the basis of alleviating a disease phenotype, as opposed to target-based drug discovery, in which lead compounds are designed to interact with predetermined disease-driving biological targets. As phenotypic drug discovery assays do not provide confirmation of a compound's mechanism of action, chemoproteomics provides valuable follow-up strategies to narrow down potential targets and eventually validate a molecule's mechanism of action. Chemoproteomics also attempts to address the inherent challenge of drug promiscuity in small molecule drug discovery by analyzing protein-small molecule interactions on a proteome-wide scale. A major goal of chemoproteomics is to characterize the interactome of drug candidates to gain insight into mechanisms of off-target toxicity and polypharmacology.

alpha/beta-Hydrolase domain containing 12 (ABHD12) is a serine hydrolase encoded by the ABHD12 gene that participates in the breakdown of the endocannabinoid neurotransmitter 2-arachidonylglycerol (2-AG) in the central nervous system. It is responsible for about 9% of brain 2-AG hydrolysis. Together, ABHD12 along with two other enzymes, monoacylglycerol lipase (MAGL) and ABHD6, control 99% of 2-AG hydrolysis in the brain. ABHD12 also serves as a lysophospholipase and metabolizes lysophosphatidylserine (LPS).

Degradomics is a sub-discipline of biology encompassing all the genomic and proteomic approaches devoted to the study of proteases, their inhibitors, and their substrates on a system-wide scale. This includes the analysis of the protease and protease-substrate repertoires, also called "protease degradomes". The scope of these degradomes can range from cell, tissue, and organism-wide scales.











