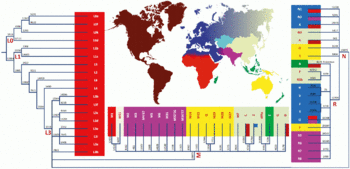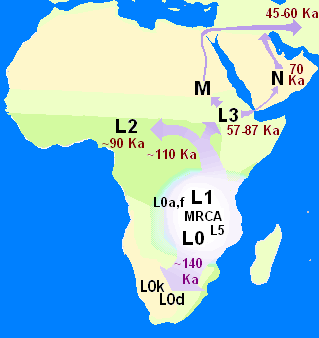
In human genetics, the Mitochondrial Eve is the matrilineal most recent common ancestor (MRCA) of all living humans. In other words, she is defined as the most recent woman from whom all living humans descend in an unbroken line purely through their mothers and through the mothers of those mothers, back until all lines converge on one woman.

A haplotype is a group of alleles in an organism that are inherited together from a single parent, and a haplogroup is a group of similar haplotypes that share a common ancestor with a single-nucleotide polymorphism mutation. More specifically, a haplotype is a combination of alleles at different chromosomal regions that are closely linked and that tend to be inherited together. As a haplogroup consists of similar haplotypes, it is usually possible to predict a haplogroup from haplotypes. Haplogroups pertain to a single line of descent. As such, membership of a haplogroup, by any individual, relies on a relatively small proportion of the genetic material possessed by that individual.

Haplogroup M is a human mitochondrial DNA (mtDNA) haplogroup. An enormous haplogroup spanning all the continents, the macro-haplogroup M, like its sibling the macro-haplogroup N, is a descendant of the haplogroup L3.
Haplogroup J is a human mitochondrial DNA (mtDNA) haplogroup. The clade derives from the haplogroup JT, which also gave rise to haplogroup T. Within the field of medical genetics, certain polymorphisms specific to haplogroup J have been associated with Leber's hereditary optic neuropathy.
Haplogroup T is a human mitochondrial DNA (mtDNA) haplogroup. It is believed to have originated around 25,100 years ago in the Near East.

Haplogroup N is a human mitochondrial DNA (mtDNA) clade. A macrohaplogroup, its descendant lineages are distributed across many continents. Like its sibling macrohaplogroup M, macrohaplogroup N is a descendant of the haplogroup L3.

Haplogroup L3 is a human mitochondrial DNA (mtDNA) haplogroup. The clade has played a pivotal role in the early dispersal of anatomically modern humans.
Haplogroup L2 is a human mitochondrial DNA (mtDNA) haplogroup with a widespread modern distribution, particularly in Subequatorial Africa. Its L2a subclade is a somewhat frequent and widely distributed mtDNA cluster on the continent, as well as among those in the Americas.

Haplogroup L1 is a human mitochondrial DNA (mtDNA) haplogroup. It is most common in Central Africa and West Africa. It diverged from L1-6 at about 140,000 years ago . Its emergence is associated with the early peopling of Africa by anatomically modern humans during the Eemian, and it is now mostly found in African pygmies.
In human mitochondrial genetics, Haplogroup Z is a human mitochondrial DNA (mtDNA) haplogroup.
Haplogroup I is a human mitochondrial DNA (mtDNA) haplogroup. It is believed to have originated about 21,000 years ago, during the Last Glacial Maximum (LGM) period in West Asia. The haplogroup is unusual in that it is now widely distributed geographically, but is common in only a few small areas of East Africa, West Asia and Europe. It is especially common among the El Molo and Rendille peoples of Kenya, various regions of Iran, the Lemko people of Slovakia, Poland and Ukraine, the island of Krk in Croatia, the department of Finistère in France and some parts of Scotland and Ireland.
Haplogroup L may refer to:
Haplogroup L0 is a human mitochondrial DNA (mtDNA) haplogroup.

In human mitochondrial genetics, haplogroup Q is a human mitochondrial DNA (mtDNA) haplogroup typical for Oceania. It is a subgroup of haplogroup M29'Q.
In human mitochondrial genetics, Haplogroup G is a human mitochondrial DNA (mtDNA) haplogroup.

Haplogroup L4 is a human mitochondrial DNA (mtDNA) haplogroup. It is a somewhat uncommon maternal clade primarily found in East Africa, and West Asia.
Haplogroup L5 is a human mitochondrial DNA (mtDNA) clade. It was previously known as L1e.
Haplogroup H is a human mitochondrial DNA (mtDNA) haplogroup. The clade is believed to have originated in Southwest Asia, near present day Syria, around 20,000 to 25,000 years ago. Mitochondrial haplogroup H is today predominantly found in Europe, and is believed to have evolved before the Last Glacial Maximum (LGM). It first expanded in the northern Near East and Southern Caucasus, and later migrations from Iberia suggest that the clade reached Europe before the Last Glacial Maximum. The haplogroup has also spread to parts of Africa, Siberia and Inner Asia. Today, around 40% of all maternal lineages in Europe belong to haplogroup H.
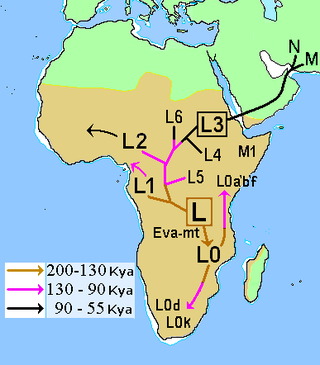
In human mitochondrial genetics, L is the mitochondrial DNA macro-haplogroup that is at the root of the anatomically modern human mtDNA phylogenetic tree. As such, it represents the most ancestral mitochondrial lineage of all currently living modern humans, also dubbed "Mitochondrial Eve".
The human mitochondrial molecular clock is the rate at which mutations have been accumulating in the mitochondrial genome of hominids during the course of human evolution. The archeological record of human activity from early periods in human prehistory is relatively limited and its interpretation has been controversial. Because of the uncertainties from the archeological record, scientists have turned to molecular dating techniques in order to refine the timeline of human evolution. A major goal of scientists in the field is to develop an accurate hominid mitochondrial molecular clock which could then be used to confidently date events that occurred during the course of human evolution.