
Phospholipids are a class of lipids whose molecule has a hydrophilic "head" containing a phosphate group and two hydrophobic "tails" derived from fatty acids, joined by an alcohol residue. Marine phospholipids typically have omega-3 fatty acids EPA and DHA integrated as part of the phospholipid molecule. The phosphate group can be modified with simple organic molecules such as choline, ethanolamine or serine.
Inositol trisphosphate or inositol 1,4,5-trisphosphate abbreviated InsP3 or Ins3P or IP3 is an inositol phosphate signaling molecule. It is made by hydrolysis of phosphatidylinositol 4,5-bisphosphate (PIP2), a phospholipid that is located in the plasma membrane, by phospholipase C (PLC). Together with diacylglycerol (DAG), IP3 is a second messenger molecule used in signal transduction in biological cells. While DAG stays inside the membrane, IP3 is soluble and diffuses through the cell, where it binds to its receptor, which is a calcium channel located in the endoplasmic reticulum. When IP3 binds its receptor, calcium is released into the cytosol, thereby activating various calcium regulated intracellular signals.

Phosphatidylinositol or inositol phospholipid is a biomolecule. It was initially called "inosite" when it was discovered by Léon Maquenne and Johann Joseph von Scherer in the late 19th century. It was discovered in bacteria but later also found in eukaryotes, and was found to be a signaling molecule.
Phosphatidic acids are anionic phospholipids important to cell signaling and direct activation of lipid-gated ion channels. Hydrolysis of phosphatidic acid gives rise to one molecule each of glycerol and phosphoric acid and two molecules of fatty acids. They constitute about 0.25% of phospholipids in the bilayer.
Second messengers are intracellular signaling molecules released by the cell in response to exposure to extracellular signaling molecules—the first messengers. Second messengers trigger physiological changes at cellular level such as proliferation, differentiation, migration, survival, apoptosis and depolarization.

Inositol phosphates are a group of mono- to hexaphosphorylated inositols. Each form of inositol phosphate is distinguished by the number and position of the phosphate group on the inositol ring.

Phosphoinositide phospholipase C is a family of eukaryotic intracellular enzymes that play an important role in signal transduction processes. These enzymes belong to a larger superfamily of Phospholipase C. Other families of phospholipase C enzymes have been identified in bacteria and trypanosomes. Phospholipases C are phosphodiesterases.
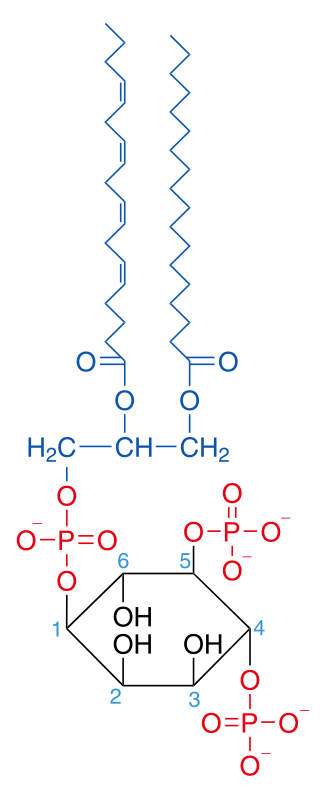
Phosphatidylinositol 4,5-bisphosphate or PtdIns(4,5)P2, also known simply as PIP2 or PI(4,5)P2, is a minor phospholipid component of cell membranes. PtdIns(4,5)P2 is enriched at the plasma membrane where it is a substrate for a number of important signaling proteins. PIP2 also forms lipid clusters that sort proteins.

Lipid signaling, broadly defined, refers to any biological cell signaling event involving a lipid messenger that binds a protein target, such as a receptor, kinase or phosphatase, which in turn mediate the effects of these lipids on specific cellular responses. Lipid signaling is thought to be qualitatively different from other classical signaling paradigms because lipids can freely diffuse through membranes. One consequence of this is that lipid messengers cannot be stored in vesicles prior to release and so are often biosynthesized "on demand" at their intended site of action. As such, many lipid signaling molecules cannot circulate freely in solution but, rather, exist bound to special carrier proteins in serum.
Phospholipase D (EC 3.1.4.4, lipophosphodiesterase II, lecithinase D, choline phosphatase, PLD; systematic name phosphatidylcholine phosphatidohydrolase) is an enzyme of the phospholipase superfamily that catalyses the following reaction

Chemokine receptors are cytokine receptors found on the surface of certain cells that interact with a type of cytokine called a chemokine. There have been 20 distinct chemokine receptors discovered in humans. Each has a rhodopsin-like 7-transmembrane (7TM) structure and couples to G-protein for signal transduction within a cell, making them members of a large protein family of G protein-coupled receptors. Following interaction with their specific chemokine ligands, chemokine receptors trigger a flux in intracellular calcium (Ca2+) ions (calcium signaling). This causes cell responses, including the onset of a process known as chemotaxis that traffics the cell to a desired location within the organism. Chemokine receptors are divided into different families, CXC chemokine receptors, CC chemokine receptors, CX3C chemokine receptors and XC chemokine receptors that correspond to the 4 distinct subfamilies of chemokines they bind. The four subfamilies of chemokines differ in the spacing of structurally important cysteine residues near the N-terminal of the chemokine.
Prostaglandin receptors or prostanoid receptors represent a sub-class of cell surface membrane receptors that are regarded as the primary receptors for one or more of the classical, naturally occurring prostanoids viz., prostaglandin D2,, PGE2, PGF2alpha, prostacyclin (PGI2), thromboxane A2 (TXA2), and PGH2. They are named based on the prostanoid to which they preferentially bind and respond, e.g. the receptor responsive to PGI2 at lower concentrations than any other prostanoid is named the Prostacyclin receptor (IP). One exception to this rule is the receptor for thromboxane A2 (TP) which binds and responds to PGH2 and TXA2 equally well.
Gq protein alpha subunit is a family of heterotrimeric G protein alpha subunits. This family is also commonly called the Gq/11 (Gq/G11) family or Gq/11/14/15 family to include closely related family members. G alpha subunits may be referred to as Gq alpha, Gαq, or Gqα. Gq proteins couple to G protein-coupled receptors to activate beta-type phospholipase C (PLC-β) enzymes. PLC-β in turn hydrolyzes phosphatidylinositol 4,5-bisphosphate (PIP2) to diacyl glycerol (DAG) and inositol trisphosphate (IP3). IP3 acts as a second messenger to release stored calcium into the cytoplasm, while DAG acts as a second messenger that activates protein kinase C (PKC).
Phospholipase C epsilon 1 (PLCE1) is an enzyme that in humans is encoded by the PLCE1 gene. This gene encodes a phospholipase enzyme (PLCE1) that catalyzes the hydrolysis of phosphatidylinositol-4,5-bisphosphate to generate two second messengers: inositol 1,4,5-triphosphate (IP3) and diacylglycerol (DAG). Mutations in this gene cause early-onset nephrotic syndrome and have been associated with respiratory chain deficiency with diffuse mesangial sclerosis.

Deleted in Liver Cancer 1 also known as DLC1 and StAR-related lipid transfer protein 12 (STARD12) is a protein which in humans is encoded by the DLC1 gene.

1-Phosphatidylinositol-4,5-bisphosphate phosphodiesterase delta-3 is an enzyme that in humans is encoded by the PLCD3 gene.
Oocyteactivation is a series of processes that occur in the oocyte during fertilization.
The insulin transduction pathway is a biochemical pathway by which insulin increases the uptake of glucose into fat and muscle cells and reduces the synthesis of glucose in the liver and hence is involved in maintaining glucose homeostasis. This pathway is also influenced by fed versus fasting states, stress levels, and a variety of other hormones.

A diglyceride, or diacylglycerol (DAG), is a glyceride consisting of two fatty acid chains covalently bonded to a glycerol molecule through ester linkages. Two possible forms exist, 1,2-diacylglycerols and 1,3-diacylglycerols. Diglycerides are natural components of food fats, though minor in comparison to triglycerides. DAGs can act as surfactants and are commonly used as emulsifiers in processed foods. DAG-enriched oil has been investigated extensively as a fat substitute due to its ability to suppress the accumulation of body fat; with total annual sales of approximately USD 200 million in Japan since its introduction in the late 1990s till 2009.
Substrate presentation is a biological process that activates a protein. The protein is sequestered away from its substrate and then activated by release and exposure of the protein to its substrate. A substrate is typically the substance on which an enzyme acts but can also be a protein surface to which a ligand binds. The substrate is the material acted upon. In the case of an interaction with an enzyme, the protein or organic substrate typically changes chemical form. Substrate presentation differs from allosteric regulation in that the enzyme need not change its conformation to begin catalysis. Substrate presentation is best described for nanoscopic distances (<100 nm).














