The urea cycle (also known as the ornithine cycle) is a cycle of biochemical reactions that produces urea (NH2)2CO from ammonia (NH3). Animals that use this cycle, mainly amphibians and mammals, are called ureotelic.

Aromatic L-amino acid decarboxylase, also known as DOPA decarboxylase (DDC), tryptophan decarboxylase, and 5-hydroxytryptophan decarboxylase, is a lyase enzyme, located in region 7p12.2-p12.1.
A metalloproteinase, or metalloprotease, is any protease enzyme whose catalytic mechanism involves a metal. An example is ADAM12 which plays a significant role in the fusion of muscle cells during embryo development, in a process known as myogenesis.

Malate dehydrogenase (EC 1.1.1.37) (MDH) is an enzyme that reversibly catalyzes the oxidation of malate to oxaloacetate using the reduction of NAD+ to NADH. This reaction is part of many metabolic pathways, including the citric acid cycle. Other malate dehydrogenases, which have other EC numbers and catalyze other reactions oxidizing malate, have qualified names like malate dehydrogenase (NADP+).
In molecular biology, biosynthesis is a multi-step, enzyme-catalyzed process where substrates are converted into more complex products in living organisms. In biosynthesis, simple compounds are modified, converted into other compounds, or joined to form macromolecules. This process often consists of metabolic pathways. Some of these biosynthetic pathways are located within a single cellular organelle, while others involve enzymes that are located within multiple cellular organelles. Examples of these biosynthetic pathways include the production of lipid membrane components and nucleotides. Biosynthesis is usually synonymous with anabolism.

A catalytic triad is a set of three coordinated amino acids that can be found in the active site of some enzymes. Catalytic triads are most commonly found in hydrolase and transferase enzymes. An acid-base-nucleophile triad is a common motif for generating a nucleophilic residue for covalent catalysis. The residues form a charge-relay network to polarise and activate the nucleophile, which attacks the substrate, forming a covalent intermediate which is then hydrolysed to release the product and regenerate free enzyme. The nucleophile is most commonly a serine or cysteine amino acid, but occasionally threonine or even selenocysteine. The 3D structure of the enzyme brings together the triad residues in a precise orientation, even though they may be far apart in the sequence.
Glutamine synthetase (GS) is an enzyme that plays an essential role in the metabolism of nitrogen by catalyzing the condensation of glutamate and ammonia to form glutamine:

Methionine synthase also known as MS, MeSe, MTR is responsible for the regeneration of methionine from homocysteine. In humans it is encoded by the MTR gene (5-methyltetrahydrofolate-homocysteine methyltransferase). Methionine synthase forms part of the S-adenosylmethionine (SAMe) biosynthesis and regeneration cycle, and is the enzyme responsible for linking the cycle to one-carbon metabolism via the folate cycle. There are two primary forms of this enzyme, the Vitamin B12 (cobalamin)-dependent (MetH) and independent (MetE) forms, although minimal core methionine synthases that do not fit cleanly into either category have also been described in some anaerobic bacteria. The two dominant forms of the enzymes appear to be evolutionary independent and rely on considerably different chemical mechanisms. Mammals and other higher eukaryotes express only the cobalamin-dependent form. In contrast, the distribution of the two forms in Archaeplastida (plants and algae) is more complex. Plants exclusively possess the cobalamin-independent form, while algae have either one of the two, depending on species. Many different microorganisms express both the cobalamin-dependent and cobalamin-independent forms.

Glucose-6-phosphate dehydrogenase (G6PD or G6PDH) (EC 1.1.1.49) is a cytosolic enzyme that catalyzes the chemical reaction
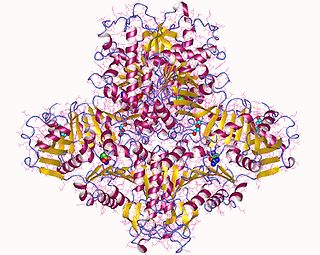
N-Acetylglutamate synthase (NAGS) is an enzyme that catalyses the production of N-acetylglutamate (NAG) from glutamate and acetyl-CoA.
Carbamoyl phosphate synthetase I is a ligase enzyme located in the mitochondria involved in the production of urea. Carbamoyl phosphate synthetase I transfers an ammonia molecule to a molecule of bicarbonate that has been phosphorylated by a molecule of ATP. The resulting carbamate is then phosphorylated with another molecule of ATP. The resulting molecule of carbamoyl phosphate leaves the enzyme.
Acyl-CoA dehydrogenases (ACADs) are a class of enzymes that function to catalyze the initial step in each cycle of fatty acid β-oxidation in the mitochondria of cells. Their action results in the introduction of a trans double-bond between C2 (α) and C3 (β) of the acyl-CoA thioester substrate. Flavin adenine dinucleotide (FAD) is a required co-factor in addition to the presence of an active site glutamate in order for the enzyme to function.

Aspartoacylase is a hydrolytic enzyme that in humans is encoded by the ASPA gene. ASPA catalyzes the deacylation of N-acetyl-l-aspartate (N-acetylaspartate) into aspartate and acetate. It is a zinc-dependent hydrolase that promotes the deprotonation of water to use as a nucleophile in a mechanism analogous to many other zinc-dependent hydrolases. It is most commonly found in the brain, where it controls the levels of N-acetyl-l-aspartate. Mutations that result in loss of aspartoacylase activity are associated with Canavan disease, a rare autosomal recessive neurodegenerative disease.

Dihydrolipoamide dehydrogenase (DLD), also known as dihydrolipoyl dehydrogenase, mitochondrial, is an enzyme that in humans is encoded by the DLD gene. DLD is a flavoprotein enzyme that oxidizes dihydrolipoamide to lipoamide.
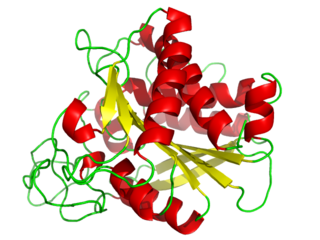
Carboxypeptidase A usually refers to the pancreatic exopeptidase that hydrolyzes peptide bonds of C-terminal residues with aromatic or aliphatic side-chains. Most scientists in the field now refer to this enzyme as CPA1, and to a related pancreatic carboxypeptidase as CPA2.

Carbamoyl phosphate synthetase catalyzes the ATP-dependent synthesis of carbamoyl phosphate from glutamine or ammonia and bicarbonate. This enzyme catalyzes the reaction of ATP and bicarbonate to produce carboxy phosphate and ADP. Carboxy phosphate reacts with ammonia to give carbamic acid. In turn, carbamic acid reacts with a second ATP to give carbamoyl phosphate plus ADP.

Citrin, also known as solute carrier family 25, member 13 (citrin) or SLC25A13, is a protein which in humans is encoded by the SLC25A13 gene.
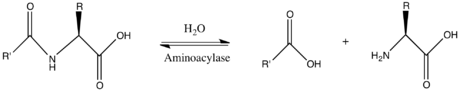
In enzymology, a N-acyl-D-amino-acid deacylase (EC 3.5.1.81) is an enzyme that catalyzes the chemical reaction

The human ETFB gene encodes the Electron-transfer-flavoprotein, beta subunit, also known as ETF-β. Together with Electron-transfer-flavoprotein, alpha subunit, encoded by the 'ETFA' gene, it forms the heterodimeric Electron transfer flavoprotein (ETF). The native ETF protein contains one molecule of FAD and one molecule of AMP, respectively.

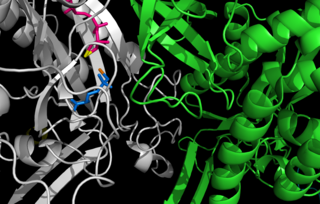
Aminoacylase-1 is an enzyme that in humans is encoded by the ACY1 gene.