Related Research Articles

In cell biology, the cytoplasm is all of the material within a cell, enclosed by the cell membrane, except for the cell nucleus. The material inside the nucleus and contained within the nuclear membrane is termed the nucleoplasm. The main components of the cytoplasm are cytosol, the organelles, and various cytoplasmic inclusions. The cytoplasm is about 80% water and usually colorless.

Cell biology is a branch of biology studying the structure and function of the cell, also known as the basic unit of life. Cell biology encompasses both prokaryotic and eukaryotic cells and can be divided into many sub-topics which may include the study of cell metabolism, cell communication, cell cycle, biochemistry, and cell composition. The study of cells is performed using several techniques such as cell culture, various types of microscopy, and cell fractionation. These have allowed for and are currently being used for discoveries and research pertaining to how cells function, ultimately giving insight into understanding larger organisms. Knowing the components of cells and how cells work is fundamental to all biological sciences while also being essential for research in biomedical fields such as cancer, and other diseases. Research in cell biology is interconnected to other fields such as genetics, molecular genetics, biochemistry, molecular biology, medical microbiology, immunology, and cytochemistry.

Proteins are large biomolecules or macromolecules that are comprised of one or more long chains of amino acid residues. Proteins perform a vast array of functions within organisms, including catalysing metabolic reactions, DNA replication, responding to stimuli, providing structure to cells and organisms, and transporting molecules from one location to another. Proteins differ from one another primarily in their sequence of amino acids, which is dictated by the nucleotide sequence of their genes, and which usually results in protein folding into a specific 3D structure that determines its activity.
Biophysics is an interdisciplinary science that applies approaches and methods traditionally used in physics to study biological phenomena. Biophysics covers all scales of biological organization, from molecular to organismic and populations. Biophysical research shares significant overlap with biochemistry, molecular biology, physical chemistry, physiology, nanotechnology, bioengineering, computational biology, biomechanics, developmental biology and systems biology.

The cytoskeleton is a complex, dynamic network of interlinking protein filaments present in the cytoplasm of all cells, including bacteria and archaea. It extends from the cell nucleus to the cell membrane and is composed of similar proteins in the various organisms. In eukaryotes, it is composed of three main components, microfilaments, intermediate filaments and microtubules, and these are all capable of rapid growth or disassembly dependent on the cell's requirements.
A kinesin is a protein belonging to a class of motor proteins found in eukaryotic cells.

Tight junctions, also known as occluding junctions or zonulae occludentes are multiprotein junctional complexes whose general function is to prevent leakage of transported solutes and water and seals the paracellular pathway. Tight junctions may also serve as leaky pathways by forming selective channels for small cations, anions, or water. Tight junctions are present mostly in vertebrates. The corresponding junctions that occur in invertebrates are septate junctions.
The Max Planck Institute for Biophysical Chemistry in Göttingen is a research institute of the Max Planck Society. Currently, 850 people work at the institute, about half of them are scientists.

Motor proteins are a class of molecular motors that can move along the cytoplasm of animal cells. They convert chemical energy into mechanical work by the hydrolysis of ATP. Flagellar rotation, however, is powered by a proton pump.
Molecular biophysics is a rapidly evolving interdisciplinary area of research that combines concepts in physics, chemistry, engineering, mathematics and biology. It seeks to understand biomolecular systems and explain biological function in terms of molecular structure, structural organization, and dynamic behaviour at various levels of complexity. This discipline covers topics such as the measurement of molecular forces, molecular associations, allosteric interactions, Brownian motion, and cable theory. Additional areas of study can be found on Outline of Biophysics. The discipline has required development of specialized equipment and procedures capable of imaging and manipulating minute living structures, as well as novel experimental approaches.

Jennifer Lippincott-Schwartz is a Senior Group Leader at Howard Hughes Medical Institute's Janelia Research Campus and a founding member of the Neuronal Cell Biology Program at Janelia. Previously, she was the Chief of the Section on Organelle Biology in the Cell Biology and Metabolism Program, in the Division of Intramural Research in the Eunice Kennedy Shriver National Institute of Child Health and Human Development at the National Institutes of Health from 1993 to 2016. Lippincott-Schwartz received her Ph.D. from Johns Hopkins University, and performed post-doctoral training with Dr. Richard Klausner at the NICHD, NIH in Bethesda, Maryland.
The following outline is provided as an overview of and topical guide to biophysics:

The cell membrane is a biological membrane that separates the interior of all cells from the outside environment which protects the cell from its environment. The cell membrane consists of a lipid bilayer, including cholesterols that sit between phospholipids to maintain their fluidity at various temperatures. The membrane also contains membrane proteins, including integral proteins that go across the membrane serving as membrane transporters, and peripheral proteins that loosely attach to the outer (peripheral) side of the cell membrane, acting as enzymes shaping the cell. The cell membrane controls the movement of substances in and out of cells and organelles. In this way, it is selectively permeable to ions and organic molecules. In addition, cell membranes are involved in a variety of cellular processes such as cell adhesion, ion conductivity and cell signalling and serve as the attachment surface for several extracellular structures, including the cell wall, the carbohydrate layer called the glycocalyx, and the intracellular network of protein fibers called the cytoskeleton. In the field of synthetic biology, cell membranes can be artificially reassembled.
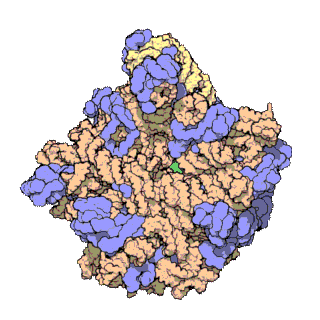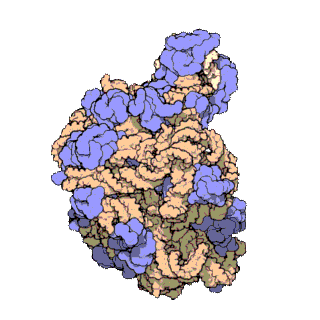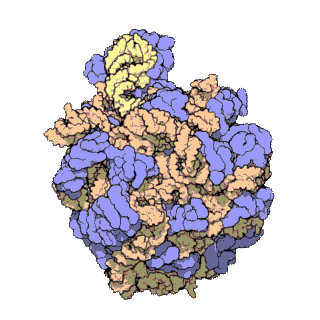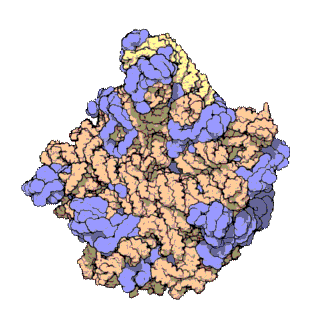
The term macromolecular assembly (MA) refers to massive chemical structures such as viruses and non-biologic nanoparticles, cellular organelles and membranes and ribosomes, etc. that are complex mixtures of polypeptide, polynucleotide, polysaccharide or other polymeric macromolecules. They are generally of more than one of these types, and the mixtures are defined spatially, and with regard to their underlying chemical composition and structure. Macromolecules are found in living and nonliving things, and are composed of many hundreds or thousands of atoms held together by covalent bonds; they are often characterized by repeating units. Assemblies of these can likewise be biologic or non-biologic, though the MA term is more commonly applied in biology, and the term supramolecular assembly is more often applied in non-biologic contexts. MAs of macromolecules are held in their defined forms by non-covalent intermolecular interactions, and can be in either non-repeating structures, or in repeating linear, circular, spiral, or other patterns. The process by which MAs are formed has been termed molecular self-assembly, a term especially applied in non-biologic contexts. A wide variety of physical/biophysical, chemical/biochemical, and computational methods exist for the study of MA; given the scale of MAs, efforts to elaborate their composition and structure and discern mechanisms underlying their functions are at the forefront of modern structure science.
Traction force microscopy (TFM) is an experimental method for determining the tractions on the surface of a biological cell by obtaining measurements of the surrounding displacement field within an in vitro extracellular matrix (ECM).
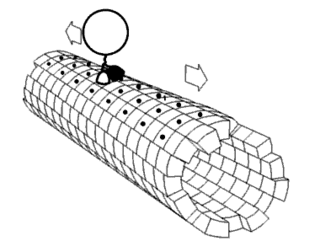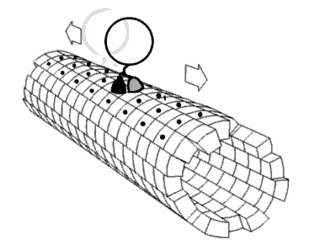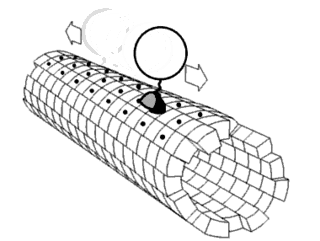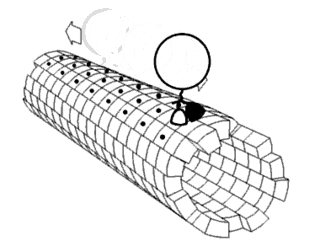
Force Spectrum Microscopy (FSM) is an application of active microrheology developed to measure aggregate random forces in the cytoplasm. Large, inert flow tracers are injected into live cells and become lodged inside the cytoskeletal mesh, wherein it is oscillated by repercussions from active motor proteins. The magnitude of these random forces can be inferred from the frequency of oscillation of tracer particles. Tracking the fluctuations of tracer particles using optical microscopy can isolate the contribution of active random forces to intracellular molecular transport from that of Brownian motion.
Cell mechanics is a sub-field of biophysics that focuses on the mechanical properties and behavior of living cells and how it relates to cell function. It encompasses aspects of cell biophysics, biomechanics, soft matter physics and rheology, mechanobiology and cell biology.

Barbara E. Ehrlich is Professor of Pharmacology and of Cellular and Molecular Physiology at Yale University working on the biophysics of membrane ion channels. Recent research investigates the function of polycystin-2, the inositol trisphosphate receptor, and the ryanodine receptor.

Denis Wirtz is the vice provost for research and Theophilus Halley Smoot Professor of Engineering Science at Johns Hopkins University. He is an expert in the molecular and biophysical mechanisms of cell motility and adhesion and nuclear dynamics in health and disease. Notably, he was the first to establish how a 3-dimensional environment fundamentally affects the way cancer cells migrate, providing more biologically and medically relevant information than 2D studies. He also pioneered the technique of particle-tracking microrheology to probe the rheological properties of complex fluids and living cells and tissues. He is a professor in the Departments of Chemical Engineering & Biomolecular Engineering and Materials Science & Engineering in the Whiting School of Engineering, and in the Departments of Oncology and Pathology in the Johns Hopkins School of Medicine.
Cell-based models are mathematical models that represent biological cells as a discrete entities. They are used in the field of computational biology for simulating the biomechanics of multicellular structures such as tissues. Their main advantage is the easy integration of cell level processes such as cell division, intracellular processes and single-cell variability within a cell population.
References
- ↑ Phillips, Rob; Kondev, Jan; Theriot, Julie (2008). Physical Biology of the Cell. Garland Science, Taylor and Francis group.
- ↑ "Molecular and Cellular Biophysics" . Retrieved April 4, 2014.
- ↑ "Center for the Physics of Living Cells" . Retrieved April 4, 2014.