Related Research Articles
Hydrolysis is any chemical reaction in which a molecule of water breaks one or more chemical bonds. The term is used broadly for substitution, elimination, and solvation reactions in which water is the nucleophile.

In biology and biochemistry, the active site is the region of an enzyme where substrate molecules bind and undergo a chemical reaction. The active site consists of amino acid residues that form temporary bonds with the substrate and residues that catalyse a reaction of that substrate. Although the active site occupies only ~10–20% of the volume of an enzyme, it is the most important part as it directly catalyzes the chemical reaction. It usually consists of three to four amino acids, while other amino acids within the protein are required to maintain the tertiary structure of the enzymes.
In organic chemistry, transesterification is the process of exchanging the organic group R″ of an ester with the organic group R' of an alcohol. These reactions are often catalyzed by the addition of an acid or base catalyst. The reaction can also be accomplished with the help of other enzymes, particularly lipases.
Isomerases are a general class of enzymes that convert a molecule from one isomer to another. Isomerases facilitate intramolecular rearrangements in which bonds are broken and formed. The general form of such a reaction is as follows:
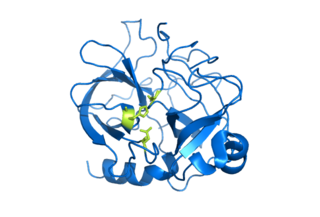
Serine proteases are enzymes that cleave peptide bonds in proteins. Serine serves as the nucleophilic amino acid at the (enzyme's) active site. They are found ubiquitously in both eukaryotes and prokaryotes. Serine proteases fall into two broad categories based on their structure: chymotrypsin-like (trypsin-like) or subtilisin-like.

In acid catalysis and base catalysis, a chemical reaction is catalyzed by an acid or a base. By Brønsted–Lowry acid–base theory, the acid is the proton (hydrogen ion, H+) donor and the base is the proton acceptor. Typical reactions catalyzed by proton transfer are esterifications and aldol reactions. In these reactions, the conjugate acid of the carbonyl group is a better electrophile than the neutral carbonyl group itself. Depending on the chemical species that act as the acid or base, catalytic mechanisms can be classified as either specific catalysis and general catalysis. Many enzymes operate by general catalysis.

A catalytic triad is a set of three coordinated amino acids that can be found in the active site of some enzymes. Catalytic triads are most commonly found in hydrolase and transferase enzymes. An acid-base-nucleophile triad is a common motif for generating a nucleophilic residue for covalent catalysis. The residues form a charge-relay network to polarise and activate the nucleophile, which attacks the substrate, forming a covalent intermediate which is then hydrolysed to release the product and regenerate free enzyme. The nucleophile is most commonly a serine or cysteine amino acid, but occasionally threonine or even selenocysteine. The 3D structure of the enzyme brings together the triad residues in a precise orientation, even though they may be far apart in the sequence.

The enzyme argininosuccinate lyase catalyzes the reversible breakdown of argininosuccinate:
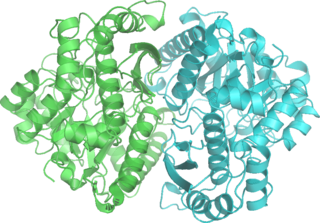
Phosphopyruvate hydratase, usually known as enolase, is a metalloenzyme (EC 4.2.1.11) that catalyses the conversion of 2-phosphoglycerate (2-PG) to phosphoenolpyruvate (PEP), the ninth and penultimate step of glycolysis. The chemical reaction is:

Muconate lactonizing enzymes are involved in the breakdown of lignin-derived aromatics, catechol and protocatechuate, to citric acid cycle intermediates as a part of the β-ketoadipate pathway in soil microbes. Some bacterial species are also capable of dehalogenating chloroaromatic compounds by the action of chloromuconate lactonizing enzymes. MLEs consist of several strands which have variable reaction favorable parts therefore the configuration of the strands affect its ability to accept protons. The bacterial MLEs belong to the enolase superfamily, several structures from which are known. MLEs have an identifying structure made up of two proteins and two Magnesium ions as well as various classes depending on whether it is bacterial or eukaryotic. The reaction mechanism that MLEs undergo are the reverse of beta-elimination in which the enolate alpha-carbon is protonated. MLEs can undergo mutations caused by a deletion of catB structural genes which can cause some bacteria to lose its functions such as the ability to grow. Additional mutations to MLEs can cause its structure and function to alter and could cause the conformation to change therefore making it an inactive enzyme that is unable to bind its substrate. There is another enzyme called Mandelate Racemase that is very similar to MLEs in the structural way as well as them both being a part of the enolase superfamily. They both have the same end product even though they undergo different chemical reactions in order to reach the end product.
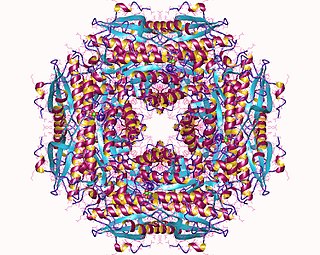
Mandelate racemase is a bacterial enzyme which catalyzes the interconversion of the enantiomers of mandelate via an enol intermediate. This enzyme catalyses the following chemical reaction

Methylmalonyl CoA epimerase is an enzyme involved in fatty acid catabolism that is encoded in human by the "MCEE" gene located on chromosome 2. It is routinely and incorrectly labeled as "methylmalonyl-CoA racemase". It is not a racemase because the CoA moiety has 5 other stereocenters.
The crotonase family comprises mechanistically diverse proteins that share a conserved trimeric quaternary structure, the core of which consists of 4 turns of a (beta/beta/alpha)n superhelix.

In enzymology, an alanine racemase is an enzyme that catalyzes the chemical reaction
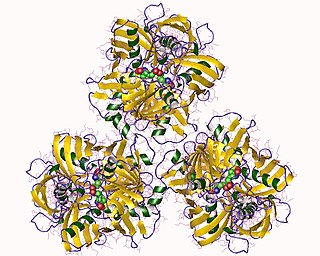
In enzymology, a diaminopimelate epimerase is an enzyme that catalyzes the chemical reaction
In enzymology, a proline racemase is an enzyme that catalyzes the chemical reaction
Fumarate lyase is a substrate of the lyase class of enzymes.

The Enzyme Function Initiative (EFI) is a large-scale collaborative project aiming to develop and disseminate a robust strategy to determine enzyme function through an integrated sequence–structure-based approach. The project was funded in May 2010 by the National Institute of General Medical Sciences as a Glue Grant which supports the research of complex biological problems that cannot be solved by a single research group. The EFI was largely spurred by the need to develop methods to identify the functions of the enormous number proteins discovered through genomic sequencing projects.
Enzyme promiscuity is the ability of an enzyme to catalyse a fortuitous side reaction in addition to its main reaction. Although enzymes are remarkably specific catalysts, they can often perform side reactions in addition to their main, native catalytic activity. These promiscuous activities are usually slow relative to the main activity and are under neutral selection. Despite ordinarily being physiologically irrelevant, under new selective pressures these activities may confer a fitness benefit therefore prompting the evolution of the formerly promiscuous activity to become the new main activity. An example of this is the atrazine chlorohydrolase from Pseudomonas sp. ADP that evolved from melamine deaminase, which has very small promiscuous activity toward atrazine, a man-made chemical.

o-Succinylbenzoate synthase (OSBS) (EC 4.2.1.113) is an enzyme encoded by the menC gene in E.coli, and catalyzes the dehydration of 2-succinyl-6-hydroxy-2,4-cyclohexadiene-1-carboxylate (SHCHC) to form 4-(2'-carboxyphenyl)-4-oxobutyrate, also called o-succinylbenzoate or OSB, hence the name of the enzyme. This reaction is the fourth step in the menaquinone biosynthetic pathway, which is used by bacteria to synthesize menaquinone, also known as vitamin K2.
References
- ↑ Babbitt, Patricia; Hasson, Miriam; Wedekind, Joseph; Palmer, David; Barrett, William; Reed, George; Rayment, Ivan; Ringe, Dagmar; Kenyon, George; Gerlt, John (1996). "The Enolase Superfamily: A General Strategy for Enzyme-Catalyzed Abstraction of the Alpha-Protons of Carboxylic Acids". Biochemistry. 51 (35): 16489–16501. doi:10.1021/bi9616413. PMID 8987982.