Related Research Articles

Bioinformatics is an interdisciplinary field that develops methods and software tools for understanding biological data, in particular when the data sets are large and complex. As an interdisciplinary field of science, bioinformatics combines biology, chemistry, physics, computer science, information engineering, mathematics and statistics to analyze and interpret the biological data. Bioinformatics has been used for in silico analyses of biological queries using computational and statistical techniques.

Genomics is an interdisciplinary field of biology focusing on the structure, function, evolution, mapping, and editing of genomes. A genome is an organism's complete set of DNA, including all of its genes as well as its hierarchical, three-dimensional structural configuration. In contrast to genetics, which refers to the study of individual genes and their roles in inheritance, genomics aims at the collective characterization and quantification of all of an organism's genes, their interrelations and influence on the organism. Genes may direct the production of proteins with the assistance of enzymes and messenger molecules. In turn, proteins make up body structures such as organs and tissues as well as control chemical reactions and carry signals between cells. Genomics also involves the sequencing and analysis of genomes through uses of high throughput DNA sequencing and bioinformatics to assemble and analyze the function and structure of entire genomes. Advances in genomics have triggered a revolution in discovery-based research and systems biology to facilitate understanding of even the most complex biological systems such as the brain.

Computational biology refers to the use of data analysis, mathematical modeling and computational simulations to understand biological systems and relationships. An intersection of computer science, biology, and big data, the field also has foundations in applied mathematics, chemistry, and genetics. It differs from biological computing, a subfield of computer engineering which uses bioengineering to build computers.

Leroy "Lee" Edward Hood is an American biologist who has served on the faculties at the California Institute of Technology (Caltech) and the University of Washington. Hood has developed ground-breaking scientific instruments which made possible major advances in the biological sciences and the medical sciences. These include the first gas phase protein sequencer (1982), for determining the sequence of amino acids in a given protein; a DNA synthesizer (1983), to synthesize short sections of DNA; a peptide synthesizer (1984), to combine amino acids into longer peptides and short proteins; the first automated DNA sequencer (1986), to identify the order of nucleotides in DNA; ink-jet oligonucleotide technology for synthesizing DNA and nanostring technology for analyzing single molecules of DNA and RNA.

Molecular genetics is a sub-field of biology that addresses how differences in the structures or expression of DNA molecules manifests as variation among organisms. Molecular genetics often applies an "investigative approach" to determine the structure and/or function of genes in an organism's genome using genetic screens. The field of study is based on the merging of several sub-fields in biology: classical Mendelian inheritance, cellular biology, molecular biology, biochemistry, and biotechnology. Researchers search for mutations in a gene or induce mutations in a gene to link a gene sequence to a specific phenotype. Molecular genetics is a powerful methodology for linking mutations to genetic conditions that may aid the search for treatments/cures for various genetics diseases.
Folding@home is a distributed computing project aimed to help scientists develop new therapeutics for a variety of diseases by the means of simulating protein dynamics. This includes the process of protein folding and the movements of proteins, and is reliant on simulations run on volunteers' personal computers. Folding@home is currently based at the University of Pennsylvania and led by Greg Bowman, a former student of Vijay Pande.

Ehud Shapiro is a multi-disciplinary scientist, artist, entrepreneur and Professor of Computer Science and Biology at the Weizmann Institute of Science. With international reputation, he made fundamental contributions to many scientific disciplines. Shapiro was also an Internet pioneer, a successful Internet entrepreneur, and a pioneer and proponent of E-democracy. Shapiro is the founder of the Ba Rock Band and conceived its original artistic program. He is a winner of two ERC Advanced Grants.

Vijay Satyanand Pande is a Trinidadian-American venture capitalist. Pande is the former director of the biophysics program and is best known for orchestrating the distributed computing disease research project known as Folding@home. His research is focused on distributed computing and computer-modelling of microbiology and on improving computer simulations regarding drug-binding, protein design, and synthetic bio-mimetic polymers. Pande became the ninth general partner at venture capital firm Andreessen Horowitz in November 2015. He is the founding investor of their Bio + Health Fund.
Biomedicine is a branch of medical science that applies biological and physiological principles to clinical practice. Biomedicine stresses standardized, evidence-based treatment validated through biological research, with treatment administered via formally trained doctors, nurses, and other such licensed practitioners.

Predictor@home was a volunteer computing project that used BOINC software to predict protein structure from protein sequence in the context of the 6th biannual CASP, or Critical Assessment of Techniques for Protein Structure Prediction. A major goal of the project was the testing and evaluating of new algorithms to predict both known and unknown protein structures.

World Community Grid (WCG) is an effort to create the world's largest volunteer computing platform to tackle scientific research that benefits humanity. Launched on November 16, 2004, with proprietary Grid MP client from United Devices and adding support for Berkeley Open Infrastructure for Network Computing (BOINC) in 2005, World Community Grid eventually discontinued the Grid MP client and consolidated on the BOINC platform in 2008. In September 2021, it was announced that IBM transferred ownership to the Krembil Research Institute of University Health Network in Toronto, Ontario.

A designer baby is a baby whose genetic makeup has been selected or altered, often to exclude a particular gene or to remove genes associated with disease. This process usually involves analysing a wide range of human embryos to identify genes associated with particular diseases and characteristics, and selecting embryos that have the desired genetic makeup; a process known as preimplantation genetic diagnosis. Screening for single genes is commonly practiced, and polygenic screening is offered by a few companies. Other methods by which a baby's genetic information can be altered involve directly editing the genome before birth, which is not routinely performed and only one instance of this is known to have occurred as of 2019, where Chinese twins Lulu and Nana were edited as embryos, causing widespread criticism.
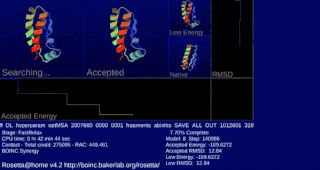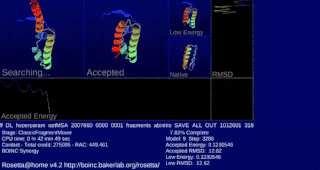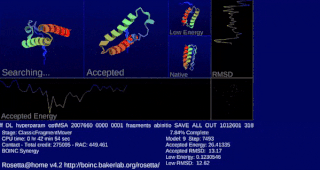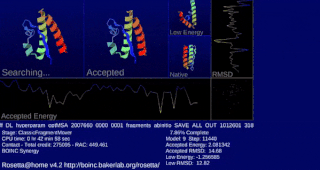
Rosetta@home is a volunteer computing project researching protein structure prediction on the Berkeley Open Infrastructure for Network Computing (BOINC) platform, run by the Baker laboratory at the University of Washington. Rosetta@home aims to predict protein–protein docking and design new proteins with the help of about fifty-five thousand active volunteered computers processing at over 487,946 GigaFLOPS on average as of September 19, 2020. Foldit, a Rosetta@home videogame, aims to reach these goals with a crowdsourcing approach. Though much of the project is oriented toward basic research to improve the accuracy and robustness of proteomics methods, Rosetta@home also does applied research on malaria, Alzheimer's disease, and other pathologies.

George McDonald Church is an American geneticist, molecular engineer, chemist, serial entrepreneur, and pioneer in personal genomics and synthetic biology. He is the Robert Winthrop Professor of Genetics at Harvard Medical School, Professor of Health Sciences and Technology at Harvard University and Massachusetts Institute of Technology, and a founding member of the Wyss Institute for Biologically Inspired Engineering at Harvard. Through his Harvard lab Church has co-founded around 50 biotech companies pushing the boundaries of innovation in the world of life sciences and making his lab as a hotbed of biotech startup activity in Boston. In 2018, the Church lab at Harvard made a record by spinning off 16 biotech companies in one year. The Church lab works on research projects that are distributed in diverse areas of modern biology like developmental biology, neurobiology, info processing, medical genetics, genomics, gene therapy, diagnostics, chemistry & bioengineering, space biology & space genetics, and ecosystem. Research and technology developments at the Church lab have impacted or made direct contributions to nearly all "next-generation sequencing (NGS)" methods and companies. In 2017, Time magazine listed him in Time 100, the list of 100 most influential people in the world. In 2022, he was featured among the most influential people in biopharma by Fierce Pharma, and was listed among the top 8 famous geneticists of all time in human history. As of January 2023, Church serves as a member of the Bulletin of the Atomic Scientists' Board of Sponsors, established by Albert Einstein.

The Human Genome Project (HGP) was an international scientific research project with the goal of determining the base pairs that make up human DNA, and of identifying, mapping and sequencing all of the genes of the human genome from both a physical and a functional standpoint. It started in 1990 and was completed in 2003. It remains the world's largest collaborative biological project. Planning for the project started after it was adopted in 1984 by the US government, and it officially launched in 1990. It was declared complete on April 14, 2003, and included about 92% of the genome. Level "complete genome" was achieved in May 2021, with a remaining only 0.3% bases covered by potential issues. The final gapless assembly was finished in January 2022.
The Cancer Genome Project is part of the cancer, aging, and somatic mutation research based at the Wellcome Trust Sanger Institute in the United Kingdom. It aims to identify sequence variants/mutations critical in the development of human cancers. Like The Cancer Genome Atlas project within the United States, the Cancer Genome Project represents an effort in the War on Cancer to improve cancer diagnosis, treatment, and prevention through a better understanding of the molecular basis of the disease. The Cancer Genome Project was launched by Michael Stratton in 2000, and Peter Campbell is now the group leader of the project. The project works to combine knowledge of the human genome sequence with high throughput mutation detection techniques.

Ibercivis was a volunteer computing platform which allows internet users to participate in scientific research by donating unused computer cycles to run scientific simulations and other tasks. The original project, which became operational in 2008, was a scientific collaboration between the Portuguese and Spanish governments, but it is open to the general public and scientific community, both within and beyond the Iberian Peninsula. The project's name is a portmanteau of Iberia and the Latin word civis, meaning 'citizen'.

Molecular cloning is a set of experimental methods in molecular biology that are used to assemble recombinant DNA molecules and to direct their replication within host organisms. The use of the word cloning refers to the fact that the method involves the replication of one molecule to produce a population of cells with identical DNA molecules. Molecular cloning generally uses DNA sequences from two different organisms: the species that is the source of the DNA to be cloned, and the species that will serve as the living host for replication of the recombinant DNA. Molecular cloning methods are central to many contemporary areas of modern biology and medicine.
Immunomics is the study of immune system regulation and response to pathogens using genome-wide approaches. With the rise of genomic and proteomic technologies, scientists have been able to visualize biological networks and infer interrelationships between genes and/or proteins; recently, these technologies have been used to help better understand how the immune system functions and how it is regulated. Two thirds of the genome is active in one or more immune cell types and less than 1% of genes are uniquely expressed in a given type of cell. Therefore, it is critical that the expression patterns of these immune cell types be deciphered in the context of a network, and not as an individual, so that their roles be correctly characterized and related to one another. Defects of the immune system such as autoimmune diseases, immunodeficiency, and malignancies can benefit from genomic insights on pathological processes. For example, analyzing the systematic variation of gene expression can relate these patterns with specific diseases and gene networks important for immune functions.
References
- 1 2 3 4 5 6 Pande lab. "Genome@home FAQ". Stanford University. Archived from the original (FAQ) on 2011-07-27. Retrieved 2011-09-05.
- 1 2 Pande lab. "What is Genome@home?". Stanford University. Archived from the original on 2011-12-04. Retrieved 2011-11-30.
- 1 2 "Genome@home Updates". 2004-03-04. Archived from the original on 2012-10-02. Retrieved 2011-11-30.
- ↑ Pande lab. "Genome@home Scientific Results". Stanford University. Archived from the original on 2011-12-04. Retrieved 2011-11-30.